Abstract
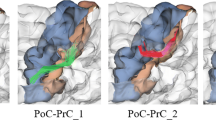
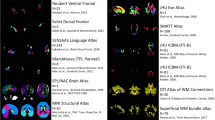
This work presents a framework driven by parcellation of brain gray matter in standard normalized space to classify the neuronal fibers obtained from diffusion tensor imaging (DTI) in entire human brain. Classification of fiber bundles into groups is an important step for the interpretation of DTI data in terms of functional correlates of white matter structures. Connections between anatomically delineated brain regions that are considered to form functional units, such as a short-term memory network, are identified by first clustering fibers based on their terminations in anatomically defined zones of gray matter according to Talairach Atlas, and then refining these groups based on geometric similarity criteria. Fiber groups identified this way can then be interpreted in terms of their functional properties using knowledge of functional neuroanatomy of individual brain regions specified in standard anatomical space, as provided by functional neuroimaging and brain lesion studies.
Chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
References
Mori, S., Crain, B.J., Chacko, V.P., van Zijl, P.C.M.: Three-dimensional tracking of axonal projections in the brain by magnetic resonance imaging. Ann. Neurol. 45, 265–269 (1999)
O’Sullivan, M., Jones, D.K., Summers, P.E., Morris, R.G., Williams, S.C.R., Markus, A.S.: Evidence for cortical disconnection as a mechanism of age-related cognitive decline. Neurology 57, 632–638 (2001)
Ding, Z., Gore, J.C., Anderson, A.W.: Classification and quantification of neuronal fiber pathways using diffusion tensor mri. Magnetic Resonance in Medicine 49, 716–721 (2003)
Corouge, I., Gouttard, S., Gerig, G.: Towards a shape model of white matter fiber bundles using diffusion tensor MRI. In: International Symposium on Biomedical Imaging, pp. 344–347 (2004)
Gerig, G., Gouttard, S., Corouge, I.: Analysis of brain white matter via fiber tract modeling. In: EMBS 2004 (2004)
Zhang, S., Laidlaw, D.H.: DTI fiber clustering in the whole brain
Brun, A., Knutsson, H., Park, H.-J., Shenton, M.E., Westin, C.-F.: Clustering fiber traces using normalized cuts. In: Barillot, C., Haynor, D.R., Hellier, P. (eds.) MICCAI 2004. LNCS, vol. 3216, pp. 368–375. Springer, Heidelberg (2004)
Akers, D., Sherbondy, A., Mackenzie, R., Dougherty, R., Wandell, B.: Exploration of the brain’s white matter pathways with dynamic queries. In: Proceedings IEEE Visualization, pp. 77–384 (2004)
Xu, D., Mori, S., Solaiyappan, M., van Zijl, P.C.M., Davatzikos, C.: A framework for callosal fiber distribution analysis. NeuroImage 71, 31–143 (2002)
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2005 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Xia, Y., Turken, A.U., Whitfield-Gabrieli, S.L., Gabrieli, J.D. (2005). Knowledge-Based Classification of Neuronal Fibers in Entire Brain. In: Duncan, J.S., Gerig, G. (eds) Medical Image Computing and Computer-Assisted Intervention – MICCAI 2005. MICCAI 2005. Lecture Notes in Computer Science, vol 3749. Springer, Berlin, Heidelberg. https://doi.org/10.1007/11566465_26
Download citation
DOI: https://doi.org/10.1007/11566465_26
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-29327-9
Online ISBN: 978-3-540-32094-4
eBook Packages: Computer ScienceComputer Science (R0)