Abstract
Chronic hepatitis C virus (HCV) infection is a global health issue. Although its progression is reported to be closely associated with lipids, the way in which the plasma lipidome changes during the development of chronic HCV infection in humans is currently unknown. Using an improved quantitative high-throughput lipidomic platform, we profiled 284 lipids in human plasma samples obtained from healthy controls (n = 11) and patients with chronic HCV infection (n = 113). The intrahepatic inflammation grade (IG) of liver tissue was determined by biopsy. Two types of mass spectrometers were integrated into a single lipidomic platform with a wide dynamic range. Compared with previous methods, the performance of this method was significantly improved in terms of both the number of target sphingolipids identified and the specificity of the high-resolution mass spectrometer. As a result, 44 sphingolipids, one diacylglycerol, 43 triglycerides, 24 glycerophosphocholines, and 5 glycerophospho-ethanolamines were successfully identified and quantified. The lipid profiles of individuals with chronic HCV infection were significantly different from those of healthy individuals. Several lipids showed significant differences between mild and severe intrahepatic inflammation grades, indicating that they could be utilized as novel noninvasive indicators of intrahepatic IG. Using multivariate analysis, healthy controls could be discriminated from HCV patients based on their plasma lipidome; however, patients with different IGs were not well discriminated. Based on these results, we speculate that variations in lipid composition arise as a result of HCV infection, and are caused by HCV-related digestive system disorders rather than progression of the disease.

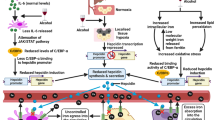
Flowchart of the lipidomic platform



Similar content being viewed by others
Abbreviations
- DG:
-
Diacylglycerol
- HCV:
-
Hepatitis C virus
- HPLC-MS:
-
High-performance liquid chromatography tandem mass spectrometry
- HRMS:
-
High-resolution mass spectrometry
- IG:
-
Inflammation grade
- LDA:
-
Lipid data analyzer
- LTQ-FT:
-
Hybrid ion trap–Fourier transform mass spectrometry
- MS:
-
Mass spectrometry
- MTBE:
-
Methyl-tert-butyl ether
- OPLS-DA:
-
Orthogonal partial least squares discriminant analysis
- PC:
-
Glycerophosphocholine
- PE:
-
Glycerophosphoethanolamine
- QC:
-
Quality control
- QQQ:
-
Triple quadrupole mass spectrometry
- SM:
-
Sphingomyelin
- TG:
-
Triglyceride
References
Alter MJ (2007) Epidemiology of hepatitis C virus infection. World J Gastroenterol WJG 13(17):2436–2441
Sherman M, Shafran S, Burak K, Doucette K, Wong W, Girgrah N, Yoshida E, Renner E, Wong P, Deschenes M (2007) Management of chronic hepatitis C: consensus guidelines. Can J Gastroenterol J Can Gastroenterol 21(Suppl C):25C–34C
Davis GL, Albright JE, Cook SF, Rosenberg DM (2003) Projecting future complications of chronic hepatitis C in the United States. Liver Transplant Off Publ Am Assoc Study Liver Dis Int Liver Transplant Soc 9(4):331–338
Ghany MG, Kleiner DE, Alter H, Doo E, Khokar F, Promrat K, Herion D, Park Y, Liang TJ, Hoofnagle JH (2003) Progression of fibrosis in chronic hepatitis C. Gastroenterology 124(1):97–104
Bravo AA, Sheth SG, Chopra S (2001) Liver biopsy. N Engl J Med 344(7):495–500
Brown HA, Marnett LJ (2011) Introduction to lipid biochemistry, metabolism, and signaling. Chem Rev 111(10):5817–5820
Puri R, Duong M, Uno K, Kataoka Y, Nicholls SJ (2012) The emerging role of plasma lipidomics in cardiovascular drug discovery. Expert Opin Drug Discov 7(1):63–72
Zehethofer N, Pinto DM (2008) Recent developments in tandem mass spectrometry for lipidomic analysis. Anal Chim Acta 627(1):62–70
Szymanska E, van Dorsten FA, Troost J, Paliukhovich I, van Velzen EJ, Hendriks MM, Trautwein EA, van Duynhoven JP, Vreeken RJ, Smilde AK (2012) A lipidomic analysis approach to evaluate the response to cholesterol-lowering food intake. Metabolomics Off J Metabolomic Soc 8(5):894–906
Chan RB, Oliveira TG, Cortes EP, Honig LS, Duff KE, Small SA, Wenk MR, Shui G, Di Paolo G (2012) Comparative lipidomic analysis of mouse and human brain with Alzheimer disease. J Biol Chem 287(4):2678–2688
Hu C, Wei H, van den Hoek AM, Wang M, van der Heijden R, Spijksma G, Reijmers TH, Bouwman J, Wopereis S, Havekes LM, Verheij E, Hankemeier T, Xu G, van der Greef J (2011) Plasma and liver lipidomics response to an intervention of rimonabant in ApoE*3Leiden.CETP transgenic mice. PloS One 6(5):e19423
Li M, Zhou Z, Nie H, Bai Y, Liu H (2011) Recent advances of chromatography and mass spectrometry in lipidomics. Anal Bioanal Chem 399(1):243–249
Yamamoto K, Isogai Y, Sato H, Taketomi Y, Murakami M (2011) Secreted phospholipase A2, lipoprotein hydrolysis, and atherosclerosis: integration with lipidomics. Anal Bioanal Chem 400(7):1829–1842
Min HK, Lim S, Chung BC, Moon MH (2011) Shotgun lipidomics for candidate biomarkers of urinary phospholipids in prostate cancer. Anal Bioanal Chem 399(2):823–830
Hart PJ, Francese S, Claude E, Woodroofe MN, Clench MR (2011) MALDI-MS imaging of lipids in ex vivo human skin. Anal Bioanal Chem 401(1):115–125
Fernandez JA, Ochoa B, Fresnedo O, Giralt MT, Rodriguez-Puertas R (2011) Matrix-assisted laser desorption ionization imaging mass spectrometry in lipidomics. Anal Bioanal Chem 401(1):29–51
Dennis EA (2009) Lipidomics joins the omics evolution. Proc Natl Acad Sci USA 106(7):2089–2090
Gao X, Zhang Q, Meng D, Isaac G, Zhao R, Fillmore TL, Chu RK, Zhou J, Tang K, Hu Z, Moore RJ, Smith RD, Katze MG, Metz TO (2012) A reversed-phase capillary ultra-performance liquid chromatography-mass spectrometry (UPLC-MS) method for comprehensive top-down/bottom-up lipid profiling. Anal Bioanal Chem 402(9):2923–2933
Chan SC, Lo SY, Liou JW, Lin MC, Syu CL, Lai MJ, Chen YC, Li HC (2011) Visualization of the structures of the hepatitis C virus replication complex. Biochem Biophys Res Commun 404(1):574–578
Mannova P, Fang R, Wang H, Deng B, McIntosh MW, Hanash SM, Beretta L (2006) Modification of host lipid raft proteome upon hepatitis C virus replication. Mol Cell Proteomics MCP 5(12):2319–2325
Bassendine MF, Sheridan DA, Felmlee DJ, Bridge SH, Toms GL, Neely RD (2011) HCV and the hepatic lipid pathway as a potential treatment target. J Hepatol 55(6):1428–1440
Diamond DL, Syder AJ, Jacobs JM, Sorensen CM, Walters KA, Proll SC, McDermott JE, Gritsenko MA, Zhang Q, Zhao R, Metz TO, Camp DG 2nd, Waters KM, Smith RD, Rice CM, Katze MG (2010) Temporal proteome and lipidome profiles reveal hepatitis C virus-associated reprogramming of hepatocellular metabolism and bioenergetics. PLoS Pathog 6(1):e1000719
Bielawski J, Szulc ZM, Hannun YA, Bielawska A (2006) Simultaneous quantitative analysis of bioactive sphingolipids by high-performance liquid chromatography-tandem mass spectrometry. Methods 39(2):82–91
Qu F, Wu CS, Hou JF, Jin Y, Zhang JL (2012) Sphingolipids as new biomarkers for assessment of delayed-type hypersensitivity and response to triptolide. PloS One 7(12):e52454
Hartler J, Trotzmuller M, Chitraju C, Spener F, Kofeler HC, Thallinger GG (2011) Lipid data analyzer: unattended identification and quantitation of lipids in LC-MS data. Bioinformatics 27(4):572–577
Umetrics (2011) Simca −P and multivariate analysis: frequently asked questions, version 1.01. Umetrics, Umeå
Ismail FW, Khan RA, Kamani L, Wadalawala AA, Shah HA, Hamid SS, Jafri W (2012) Nutritional status in patients with hepatitis C. J Coll Phys Surg Pak JCPSP 22(3):139–142
Pohjanen E, Thysell E, Jonsson P, Eklund C, Silfver A, Carlsson IB, Lundgren K, Moritz T, Svensson MB, Antti H (2007) A multivariate screening strategy for investigating metabolic effects of strenuous physical exercise in human serum. J Proteome Res 6(6):2113–2120
Stenlund H, Gorzsas A, Persson P, Sundberg B, Trygg J (2008) Orthogonal projections to latent structures discriminant analysis modeling on in situ FT-IR spectral imaging of liver tissue for identifying sources of variability. Anal Chem 80(18):6898–6906
Bao Y, Zhao T, Wang X, Qiu Y, Su M, Jia W, Jia W (2009) Metabonomic variations in the drug-treated type 2 diabetes mellitus patients and healthy volunteers. J Proteome Res 8(4):1623–1630
Ejsing CS, Duchoslav E, Sampaio J, Simons K, Bonner R, Thiele C, Ekroos K, Shevchenko A (2006) Automated identification and quantification of glycerophospholipid molecular species by multiple precursor ion scanning. Anal Chem 78(17):6202–6214
Katajamaa M, Miettinen J, Oresic M (2006) MZmine: toolbox for processing and visualization of mass spectrometry based molecular profile data. Bioinformatics 22(5):634–636
Quehenberger O, Armando AM, Brown AH, Milne SB, Myers DS, Merrill AH, Bandyopadhyay S, Jones KN, Kelly S, Shaner RL, Sullards CM, Wang E, Murphy RC, Barkley RM, Leiker TJ, Raetz CR, Guan Z, Laird GM, Six DA, Russell DW, McDonald JG, Subramaniam S, Fahy E, Dennis EA (2010) Lipidomics reveals a remarkable diversity of lipids in human plasma. J Lipid Res 51(11):3299–3305
Bird SS, Marur VR, Sniatynski MJ, Greenberg HK, Kristal BS (2011) Lipidomics profiling by high-resolution LC-MS and high-energy collisional dissociation fragmentation: focus on characterization of mitochondrial cardiolipins and monolysocardiolipins. Anal Chem 83(3):940–949
Acknowledgments
The Ministry of Science and Technology of the People’s Republic of China (2012ZX09301002-006), and the National Science and Technology Key Project on “Major Infectious Diseases such as HIV/AIDS, Viral Hepatitis Prevention and Treatment” (2012ZX10002004-006) are acknowledged.
Conflict of interest disclosure
The authors declare no competing financial interest.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Feng Qu and Su-Jun Zheng are co-first author. Jin-Lan Zhang and Zhong-Ping Duan are co-correspondence author.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 703 kb)
Rights and permissions
About this article
Cite this article
Qu, F., Zheng, SJ., Wu, CS. et al. Lipidomic profiling of plasma in patients with chronic hepatitis C infection. Anal Bioanal Chem 406, 555–564 (2014). https://doi.org/10.1007/s00216-013-7479-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-013-7479-8