Abstract
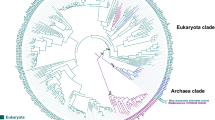
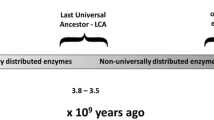
The acyl-CoA dehydrogenases (ACADs) are enzymes that catalyze the α,β-dehydrogenation of acyl-CoA esters in fatty acid and amino acid catabolism. Eleven ACADs are now recognized in the sequenced human genome, and several homologs have been reported from bacteria, fungi, plants, and nematodes. We performed a systematic comparative genomic study, integrating homology searches with methods of phylogenetic reconstruction, to investigate the evolutionary history of this family. Sequence analyses indicate origin of the family in the common ancestor of Archaea, Bacteria, and Eukaryota, illustrating its essential role in the metabolism of early life. At least three ACADs were already present at that time: ancestral glutaryl-CoA dehydrogenase (GCD), isovaleryl-CoA dehydrogenase (IVD), and ACAD10/11. Two gene duplications were unique to the eukaryotic domain: one resulted in the VLCAD and ACAD9 paralogs and another in the ACAD10 and ACAD11 paralogs. The overall patchy distribution of specific ACADs across the tree of life is the result of dynamic evolution that includes numerous rounds of gene duplication and secondary losses, interdomain lateral gene transfer events, alteration of cellular localization, and evolution of novel proteins by domain acquisition. Our finding that eukaryotic ACAD species are more closely related to bacterial ACADs is consistent with endosymbiotic origin of ACADs in eukaryotes and further supported by the localization of all nine previously studied ACADs in mitochondria.








Similar content being viewed by others
Abbreviations
- ACADs:
-
Acyl-CoA dehydrogenases
- ACOX:
-
Acyl-CoA oxidase
- GCD:
-
Glutaryl-CoA dehydrogenase
- IBD:
-
Isobutyryl-CoA dehydrogenase
- IVD:
-
Isovaleryl-CoA dehydrogenase
- VLCAD:
-
Very long-chain acyl-CoA dehydrogenase
- LCAD:
-
Long-chain acyl-CoA dehydrogenase
- SCAD:
-
Short-chain acyl-CoA dehydrogenase
- SBCAD:
-
Short/branched-chain acyl-CoA dehydrogenase, also known as 2-methyl branched chain acyl-CoA dehydrogenase
- MCAD:
-
Medium-chain acyl-CoA dehydrogenase
- LGT:
-
Lateral gene transfer
References
Altschul SF, Madden TL, Schaffer AA, Zhang JH, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Andresen BS, Bross P, Vianey-Saban C, Divry P, Zabot MT, Roe CR, Nada MA, Byskov A, Kruse TA, Neve S, Kristiansen K, Knudsen I, Corydon MJ, Gregersen N (1996) Cloning and characterization of human very-long-chain acyl-CoA dehydrogenase cDNA, chromosomal assignment of the gene and identification in four patients of nine different mutations within the VLCAD gene. Hum Mol Genet 5:461–472
Battaile K, Molin-Case J, Paschke R, Wang M, Bennett D, Vockley J, Kim J-JP (2002) Crystal structure of rat short chain acyl-CoA dehydrogenase complexed with acetoacetyl-CoA; comparison with other acyl-CoA dehydrogenases. J Biol Chem 277:12200–12207
Battaile KP, Nguyen TV, Vockley J, Kim JJ (2004) Structures of isobutyryl-CoA dehydrogenase and enzyme-product complex: comparison with isovaleryl- and short-chain acyl-CoA dehydrogenases. J Biol Chem 279:16526–16534
Bode K, Hooks MA, Couee II (1999) Identification, separation, and characterization of acyl-coenzyme A dehydrogenases involved in mitochondrial beta-oxidation in higher plants. Plant Physiol 119:1305–1314
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Daschner K, Thalheim C, Guha C, Brennicke A, Binder S (1999) In plants a putative isovaleryl-CoA-dehydrogenase is located in mitochondria. Plant Mol Biol 39:1275–1282
De Bellis L, Gonzali S, Alpi A, Hayashi H, Hayashi M, Nishimura M (2000) Purification and characterization of a novel pumpkin short-chain acyl-coenzyme A oxidase with structural similarity to acyl-coenzyme A dehydrogenases. Plant Physiol 123:327–334
Dieuaide M, Couee I, Pradet A, Raymond P (1993) Effects of glucose starvation on the oxidation of fatty acids by maize root tip mitochondria and peroxisomes: evidence for mitochondrial fatty acid beta-oxidation and acyl-CoA dehydrogenase activity in a higher plant. Biochem J 296(Pt 1):199–207
Djordjevic S, Pace CP, Stankovich MT, Kim JJP (1995) Three-dimensional structure of butyryl-CoA dehydrogenase from Megasphaera esdenii. Biochemistry 34:2163–2171
Duran E, Walker DJ, Johnson KR, Komuniecki PR, Komuniecki RW (1998) Developmental and tissue-specific expression of 2-methyl branched-chain enoyl CoA reductase isoforms in the parasitic nematode, Ascaris suum. Mol Biochem Parasitol 91:307–318
Ensenauer R, He M, Willard JM, Goetzman ES, Corydon TJ, Vandahl BB, Mohsen AW, Isaya G, Vockley J (2005) Human acyl-CoA dehydrogenase-9 plays a novel role in the mitochondrial beta-oxidation of unsaturated fatty acids. J Biol Chem 280:32309–32316
Faivre-Nitschke SE, Couee I, Vermel M, Grienenberger JM, Gualberto JM (2001) Purification, characterization and cloning of isovaleryl-CoA dehydrogenase from higher plant mitochondria. Eur J Biochem 268:1332–1339
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Finocchiaro G, Ito M, Tanaka K (1987) Purification and properties of short chain acyl-CoA, medium chain acyl-CoA, and isovaleryl-CoA dehydrogenases from human liver. J Biol Chem 262:7982–7989
Foster PG, Hickey DA (1999) Compositional bias may affect both DNA-based and protein-based phylogenetic reconstructions. J Mol Evol 48:284–290
Fu Z, Wang M, Paschke R, Rao KS, Frerman FE, Kim JJ (2004) Crystal structures of human glutaryl-CoA dehydrogenase with and without an alternate substrate: structural bases of dehydrogenation and decarboxylation reactions. Biochemistry 43:9674–9684
Gerhardt B, Fischer K, Maier U (1995) Effect of palmitoylcarnitine on mitochondrial activities. Planta 196:720–726
Goetzman ES, Mohsen AW, Prasad K, Vockley J (2005) Convergent evolution of a 2-methylbutyryl-CoA dehydrogenase from isovaleryl-CoA dehydrogenase in Solanum tuberosum. J Biol Chem 280:4873–4879
Goodman SI, Kratz LE, DiGiulio KA, Biery BJ, Goodman KE, Isaya G, Frerman FE (1995) Cloning of glutaryl-CoA dehydrogenase cDNA, and expression of wild type and mutant enzymes in Escherichia coli. Hum Mol Genet 4:1493–1498
Graham IA, Eastmond PJ (2002) Pathways of straight and branched chain fatty acid catabolism in higher plants. Prog Lipid Res 41:156–181
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52:696–704
Hall CL (1978) Acyl-CoA dehydrogenases and electron-transferring flavoprotein. Methods Enzymol 53:502–518
Hamada K, Ago H, Kuramitsu S, Miyano M (2004) Crystal structure of Thermus thermophilus medium-chain acyl-CoA dehydrogenase. PDB
Hayashi H, De Bellis L, Ciurli A, Kondo M, Hayashi M, Nishimura M (1999) A novel acyl-CoA oxidase that can oxidize short-chain acyl-CoA in plant peroxisomes. J Biol Chem 274:12715–12721
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17:754–755
Ikeda Y, Keese S, Fenton WA, Tanaka K (1987) Biosynthesis of four rat liver mitochondrial acyl-CoA dehydrogenases. Import into mitochondria and processing of their precursors in a cell-free system and in cultured cells. Arch Biochem Biophys 252:662–674
Kikuchi M, Hatano N, Yokota S, Shimozawa N, Imanaka T, Taniguchi H (2004) Proteomic analysis of rat liver peroxisome: presence of peroxisome-specific isozyme of Lon protease. J Biol Chem 279:421–428
Kim JJP, Wang M, Paschke R (1993) Crystal structures of medium-chain acyl-CoA dehydrogenase from pig liver mitochondria with and without substrate. Proc Natl Acad Sci USA 90:7523–7527
Kionka C, Kunau WH (1985) Inducible beta-oxidation pathway in Neurospora crassa. J Bacteriol 161:153–157
Klein K (1973) Dehydrogenasen und ETF in Escherichia coli. Studien zum Fettsäureabbau. Doctoral dissertation, University of Cologne, Cologne, Germany
Komuniecki R, Fekete S, Thissen-Parra J (1985) Purification and characterization of the 2-methyl branched-chain Acyl-CoA dehydrogenase, an enzyme involved in NADH-dependent enoyl-CoA reduction in anaerobic mitochondria of the nematode, Ascaris suum. J Biol Chem 260:4770–4777
Krasko A, Schroder HC, Hassanein HM, Batel R, Muller IM, Muller WE (1998) Identification and expression of the SOS response, aidB-like, gene in the marine sponge Geodia cydonium: implication for the phylogenetic relationships of metazoan acyl-CoA dehydrogenases and acyl-CoA oxidases. J Mol Evol 47:343–352
Kunau WH, Dommes V, Schulz H (1995) Beta-oxidation of fatty acids in mitochondria, peroxisomes, and bacteria: a century of continued progress. Prog Lipid Res 34:267–342
Landini P, Hajec LI, Volkert MR (1994) Structure and transcriptional regulation of the Escherichia coli adaptive response gene aidB. J Bacteriol 176:6583–6589
Lea W, Abbas AS, Sprecher H, Vockley J, Schulz H (2000) Long-chain acyl-CoA dehydrogenase is a key enzyme in the mitochondrial beta-oxidation of unsaturated fatty acids. Biochim Biophys Acta 1485:121–128
Lee HJ, Wang M, Paschke R, Nandy A, Ghisla S, Kim JJ (1996) Crystal structures of the wild type and the Glu376Gly/Thr255Glu mutant of human medium-chain acyl-CoA dehydrogenase: influence of the location of the catalytic base on substrate specificity. Biochemistry 35:12412–12420
Lenich AC, Goodman SI (1986) The purification and characterization of glutaryl-coenzyme A dehydrogenase from porcine and human liver. J Biol Chem 261:4090–4096
Masterson C, Wood C (2001) Mitochondrial and peroxisomal beta-oxidation capacities of organs from a non-oilseed plant. Proc Biol Sci 268:1949–1953
Nandy A, Kuchler B, Ghisla S (1996) Molecular evolution and substrate specificity of acyl-CoA dehydrogenases: chimaeric “medium/long’ chain-specific enzyme from medium-chain acyl-CoA dehydrogenase. Biochem Soc Trans 24:105–110
Oey NA, Ruiter JP, Ijlst L, Attie-Bitach T, Vekemans M, Wanders RJ, Wijburg FA (2006) Acyl-CoA dehydrogenase 9 (ACAD 9) is the long-chain acyl-CoA dehydrogenase in human embryonic and fetal brain. Biochem Biophys Res Commun 346:33–37
Page RD (1996) TreeView: an application to display phylogenetic trees on personal computers. Comput Appl Biosci 12:357–358
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Schmidt HA, Strimmer K, Vingron M, von Haeseler A (2002) TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18:502–504
Souri M, Aoyama T, Hoganson G, Hashimoto T (1998a) Very-long-chain acyl-CoA dehydrogenase subunit assembles to the dimer form on mitochondrial inner membrane. FEBS Lett 426:187–190
Souri M, Aoyama T, Yamaguchi S, Hashimoto T (1998b) Relationship between structure and substrate-chain-length specificity of mitochondrial very-long-chain acyl-coenzyme A dehydrogenase. Eur J Biochem 257:592–598
Strausberg RL, Feingold EA, Grouse LH, Derge JG, Klausner RD, Collins FS, Wagner L, Shenmen CM, Schuler GD, Altschul SF, Zeeberg B, Buetow KH, Schaefer CF, Bhat NK, Hopkins RF, Jordan H, Moore T, Max SI, Wang J, Hsieh F, Diatchenko L, Marusina K, Farmer AA, Rubin GM, Hong L, Stapleton M, Soares MB, Bonaldo MF, Casavant TL, Scheetz TE, Brownstein MJ, Usdin TB, Toshiyuki S, Carninci P, Prange C, Raha SS, Loquellano NA, Peters GJ, Abramson RD, Mullahy SJ, Bosak SA, McEwan PJ, McKernan KJ, Malek JA, Gunaratne PH, Richards S, Worley KC, Hale S, Garcia AM, Gay LJ, Hulyk SW, Villalon DK, Muzny DM, Sodergren EJ, Lu X, Gibbs RA, Fahey J, Helton E, Ketteman M, Madan A, Rodrigues S, Sanchez A, Whiting M, Madan A, Young AC, Shevchenko Y, Bouffard GG, Blakesley RW, Touchman JW, Green ED, Dickson MC, Rodriguez AC, Grimwood J, Schmutz J, Myers RM, Butterfield YS, Krzywinski MI, Skalska U, Smailus DE, Schnerch A, Schein JE, Jones SJ, Marra MA, Mammalian Gene Collection Program T (2002) Generation and initial analysis of more than 15, 000 full-length human and mouse cDNA sequences. Proc Natl Acad Sci USA 99:16899–16903
Swofford DL (2000) PAUP*. Phylogenetic analysis using parsimony (*and other methods). Version 4.0 Beta. Sinauer, Sunderland
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal-W—improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tiffany KA, Roberts DL, Wang M, Paschke R, Mohsen AWA, Vockley J, Kim JJP (1997) Structure of human isovaleryl-coA dehydrogenase at 2.6 angstrom resolution—basis for substrate specificity. Biochemistry 36:8455–8464
Toogood HS, van Thiel A, Basran J, Sutcliffe MJ, Scrutton NS, Leys D (2004) Extensive domain motion and electron transfer in the human electron transferring flavoprotein.medium chain Acyl-CoA dehydrogenase complex. J Biol Chem 279:32904–32912
Valenciano S, Lucas JR, Pedregosa A, Monistrol IF, Laborda F (1996) Induction of beta-oxidation enzymes and microbody proliferation in Aspergillus nidulans. Arch Microbiol 166:336–341
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699
Ye X, Ji C, Zhou C, Zeng L, Gu S, Ying K, Xie Y, Mao Y (2004) Cloning and characterization of a human cDNA ACAD10 mapped to chromosome 12q24.1. Mol Biol Rep 31:191–195
Zhang J, Zhang W, Zou D, Chen G, Wan T, Zhang M, Cao X (2002) Cloning and functional characterization of ACAD-9, a novel member of human acyl-CoA dehydrogenase family. Biochem Biophys Res Commun 297:1033–1042
Acknowledgments
This work was supported by PHS NIH Grant R01-DK54936 and the Pennsylvania Department of Health, Tobacco Formula Funding. We thank Eric Goetzman for discussion on physiological effects of ACAD deficiencies.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below are the links to the electronic supplementary materials.
Rights and permissions
About this article
Cite this article
Swigoňová, Z., Mohsen, AW. & Vockley, J. Acyl-CoA Dehydrogenases: Dynamic History of Protein Family Evolution. J Mol Evol 69, 176–193 (2009). https://doi.org/10.1007/s00239-009-9263-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-009-9263-0