Abstract
Objectives
The broad host range of pseudorabies virus (PRV) and large capacity for foreign DNA make it a promising vector for the development of vaccines and agents of gene therapy.
Results
We show that up to 100 % viral gene disrupting efficiency was achieved by simple co-transfection of the purified PRV genomes with the clustered regularly-interspaced, short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) into cells. Furthermore, CRISPR/Cas9-mediated knock-in of >4-kb-long DNA cassettes into the PRV genome at a positive rate of 50 % by a homology-independent DNA repair mechanism without constructing homology arms. This approach requires only a simple plasmid construction and is applicable to knock-in of other foreign genes.
Conclusion
Our studies offered simple and efficient methods to manipulate PRV.
Similar content being viewed by others
Introduction
Pseudorabies virus (PRV) is a swine alpha-herpesvirus causing substantial economic losses to the pig industry (Knapp and Enquist 1997). Since it has a large and well-characterized double-strand DNA genome with numerous non-essential genes, which can be replaced by foreign genes without affecting virus infectivity or propagation, PRV is a promising vector for vaccine development (Dong et al. 2014). Due to its nonpathogenicity in humans, PRV showed great potential in tumor therapy (Boldogkoi and Nogradi 2003). Traditionally, recombinant PRV is generated by spontaneous homologous recombination in mammalian cells, or through cloning in bacterial artificial chromosome (BAC) in Escherichia coli (Smith and Enquist 2000). However, both methods are laborious.
The CRISPR/Cas9 system has been used for manipulating genomes of large DNA viruses, including type I herpes simplex virus (HIV-1) and adenovirus (ADV) (Bi et al. 2014; Suenaga et al. 2014). The success of viral genome manipulation by CRISPR/Cas9 system requires that viral genome and the CRISPR/Cas9 components co-exist in host cells. Viruses that replicate rapidly in the host cells with poor DNA transfection efficiency may result in the produced progeny viruses being overwhelmingly wild type, making isolation of recombinant ones difficult. Here, we showed that up to 100 % viral gene disrupting efficiency was achieved by cotransfection of the purified PRV genomes with the CRISPR/Cas9 system into PK15 cells with poor transfection efficiency.
Homologous recombination (HR)-mediated gene knock-in enables large DNA viral genome editing using CRISPR/Cas9 system (Bi et al. 2014). However, the labor for constructing vectors containing homology arms and difficulties in inducing HR in some cell types represent technical hurdles for the application of HR-mediated knock-in technology (Nakade et al. 2014). Here, we report an alternative approach for gene knock-in using CRISPR/Cas9 through a homology-independent DNA repair mechanism. Concurrent cleavage of donor plasmid DNA and the selected PRV genome integration site resulted in efficient targeted integration of donor DNA, after cotransfection of a donor plasmid with single guide RNAs (sgRNAs)/Cas9 vectors and viral genomes.
Materials and methods
Viruses and cells
PRV BarthaK61 strain vaccine was purchased from Weike Biotech Co., Harbin, China. The virus has been plaque-purified and adapted to PK15 cells. PK15 cells and Hela cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (Invitrogen, USA) containing 10 % (v/v) fetal bovine sera (FBS) (Gibco, USA), 100 µg streptomycin/ml and 100 IU penicillin/ml.
Plasmid and mutant virus production
All sgRNAs were designed using the online CRISPR Design Tool (http://tools.genome-engineering.org), and target the amino-terminal regions of the PRV Early Protein 0 (EP0) and UL50 open reading frames. The targeting plasmids of the CRISPR/Cas9 system were constructed by introducing synthesized oligo-DNA primers corresponding to each target sequence into the BbsI restriction sites of the pUC19 sgRNA preceding the guide RNA scaffold (Chang et al. 2013) (Fig. 1a). The oligo-DNA primers are listed in Supplementary Table 1. Viral DNA was extracted and purified from infected cells by SDS-proteinase K extraction as described earlier (Smith and Enquist 1999). The red fluorescent protein (RFP) expression cassette was amplified from pmRFP-C1 vector using primers (5′–3′-end) RFP-fwd: AATAGTAATCAATTACGGGGTCATT, RFP-rev: AGATACATTGATGAGTTTGGACAAACCA. The EP0 bait sequence in the donor plasmid, which is the short fragment of EP0 locus encompassing the targeting sequences of sgRNAs, was amplified from the viral DNA using the following primers (5′- to 3′-end): EP0-fwd: GACTGCCCCATCTGTCTG, EP0-rev: CCGTAATTGATTACTATTTCCTCGGTATAGTCTTCACCC. The fusion fragment of RFP expression cassette with the EP0 bait sequence at its 5′-end was generated using High-Fidelity DNA polymerase, and then inserted into the pCloneEZ- Blunt-Amp/HC cloning vector. All constructs were verified by sequencing.
The CRISPR/Cas9 system and the protocol used to disrupt the PRV genome. a Schematic diagrams of the CRISPR/Cas9 system showing the essential elements for the expression of Cas9 protein and sgRNAs respectively. b A diagram depicting the protocol used to disrupt the PRV genome. A mixture of Cas9/sgRNAs and PRV genomes were introduced into PK15 cells. 2–4 days after transfection when CPE was observed, the cells were collected for PCR analysis, and the supernatants were inoculated into the cells grown in 96-well plates after serial dilutions to obtain single viral clones. Subcloned viruses were sequence analyzed
For generation of mutant viruses, mock or sgRNA constructs (0.5 µg), pCDNA3.1 cas9 (0.5 µg) (Chang et al. 2013) expressing Cas9 (Fig. 1a) and viral DNA genome (1 µg) were co-transfected into PK15 cells using Lipofectine 2000 (Invitrogen, USA) according to the manufacturer’s instructions. 2–4 days after transfection, the cells with expected cytopathic effect (CPE) were collected for genotyping analysis by PCR. The supernatants of cells with CPE were serially diluted by 10−1–10−8 fold, and inoculated into newly plated PK15 cells. 3–4 days later, viral genomic DNA was extracted and purified from PK15 cells following a standard protocol. PCR was performed using sequence-specific primer pairs (for EP0: forward, 5′-CGCAGCGCCGCTTTCAGACCCA-3′ and reverse, 5′-GGAGCATGGCCTCGGTCAC-3′; for UL50: forward, 5′-TGGGCTGATCCACCGGGACTC-3′ and reverse, 5′-TGAGGGACGAGCGCCCGAAGA-3′). After the purification of amplified DNA, the EP0 and UL50 fragment were cloned into PMD18-T vector. The colonies containing the inserted genes were sequenced. Mutants were identified by comparison to the wild-type sequence (Fig. 1b).
Endpoint dilution assay
PK15 cells were seeded into 96-well plates at a density of 5 × 103 cells/well. 24 h later, the cells were infected with PRV mutants at a 10-fold serial dilution (10−1–10−8). When CPE occurred, PRV in the highest dilution plate (single colonies) was collected for PCR and western blotting analysis. The number of positive and negative wells was recorded to calculate the efficiency.
Antibodies and western blotting
To examine the expressions of viral proteins, PK15 cells were infected with different mutants or wild-type (WT) PRV at a multiplicity of infection (MOI) of 1 for 24 h. Cells displaying CPE were harvested and lysed in SDS/β-mercaptoethanol protein lysis buffer. Lysates were subjected to SDS-PAGE, and then transferred to nitrocellulose membranes. The membranes were subsequently blocked in 5 % milk-PBS-T (PBS, 0.1 % Tween 20) and incubated with the indicated primary antibodies followed by a species-specific secondary antibody. The antibodies against EP0, US3 and UL50 were raised in mice individually with the N-terminal region of each protein as antigens. The specificity of each raised antibody was verified (Supplementary Fig. 1). Mouse anti-RFP antibody was purchased from Abcam (USA).
One-step growth kinetics of viruses
One-step growth kinetics was conducted as described previously (Smith and Enquist 1999). PK15 cells were infected with the indicated viruses at an MOI of 1, and the supernatants were then collected at the indicated time points for virus titer determination. Virus titers were presented as 50 % tissue culture infectious dose (TCID50). Growth kinetics for each virus was performed in duplicates, and the resulting titers were then averaged.
Results and discussion
Disruption of PRV EP0 and UL50 genes using CRISPR/Cas9 system
In the previous studies, the CRISPR/Cas9 system was transfected into cells followed by wild-type HIV-1 and ADV infection and progeny virus isolation (Bi et al. 2014). However, for cells with low transfection efficiency, this sequential approach may yield a low proportion of mutant viruses in the progeny as the majority of viral infected cells may not have the CRISPR/Cas9 system transfected into them. This situation would make the task of selecting mutant viruses challenging. We first followed the published procedure and transfected PK15 cells with the sgRNAs and Cas9-expressing plasmid followed by infection of the cells with PRV at different MOIs. We synthesized two sgRNAs specific to EP0 to increase Cas9 efficiency (Fig. 2a). Unfortunately, we failed to isolate recombinant viruses and all of the progeny viruses we checked were wild type (data not shown). The likely reason for this is that the low transfection efficiency of PK15 cells, which is about 30 % at the best scenario, resulted in the majority of viral infected cells not expressing CRISPR/Cas9 system. The rapid replication rate of the viral genomes may exempt many of them from being cleaved by Cas9. As a result, the percentage of viral genomes cleaved by CRISPR/Cas9 system was extremely low making the selection of recombinant viruses difficult.
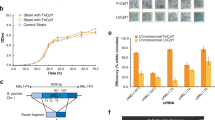
Disruption of the PRV EP0 gene by the CRISPR/Cas9 system in PK15 cells. a Schematic diagram showing the EP0 locus in the PRV genome and the targeting sequences of sgRNA1/2 as well as their respective protospacer adjacent motif (PAM). The expected cleavage sites were indicated by the scissors. b PK15 cells displaying CPE after transfection of Cas9/sgRNAs and PRV genomes in different combinations as indicated were collected for PCR analysis using the primers as indicated in the diagram above. c An example of sequence chromatogram showing the disruption of EP0 via a genomic deletion mediated by two sgRNAs. The red marker points to the junction site repaired. (d and e) PK15 d and Hela e cells were either mock infected or infected with wild-type (WT) or EP0 disrupted mutant (designated as EP0 KO) viruses for 24 h and then lysed for Western analysis with the antibodies against viral proteins EP0 and US3 respectively. f PK15 cells grown in a 24-well plate were inoculated with WT or EP0 KO viruses at an MOI of 1. The culture supernatants were collected at the indicated time points, and used to determine the viral titers. Error bars represent the standard errors of the mean of two replicates
To increase the percentage of viral genomes being cleaved by CRISPR/Cas9 system, we co-transfected viral genomes and CRISPR/Cas9 system components containing two targeting sgRNAs for EP0 (Fig. 2a) into PK15 cells. The subsequent steps were essentially the same as the described in methods (Fig. 1b). Upon PRV-mediated CPE being prominently observed, we collected the cells for genotyping analysis by PCR with the primers specific to EP0 locus. If the CRISPR/Cas9 is effective, shorter PCR products resulted from the targeted deletions are expected. Strikingly, PCR analysis showed that only a shorter band was generated (Fig. 2b), indicating that the great majority of viruses collected were recombinant. We subcloned the viruses by inoculating them in 96-well plates after dilution, followed by PCR analysis. The results confirmed that most of the progeny viruses, if not all, were EP0 gene disrupted (data not shown). Further sequencing analysis revealed that the viral genomes were cleaved and repaired at the exact cleavage sites mediated by EP0 sgRNA1 and sgRNA2 (Fig. 2c). Western analysis of PK15 (Fig. 2d) and a human cell line Hela (Fig. 2e) infected with wild type or mutant viruses further confirmed that EP0 gene was successfully disrupted in the virus. As expected, the EP0 disrupted virus has slower replication rate than wild type in PK15 cells (Fig. 2f).
This approach guaranteed a near 100 % co-presence of viral genomes with the CRISPR/gRNAs despite only in 30 % of PK-15 cells transfected. Strikingly, a close to 100 % of gene disruption efficiency in progeny viruses was achieved, thus dramatically simplified viral isolation, and shortened the time period required. More importantly, the viral gene EP0 that we disrupted is critically involved in PRV replication indicating our approach is applicable to disruption of a wide range of genes, and not limited to certain non-essential genes.
We confirmed the effectiveness of our viral genome editing approach by disrupting another PRV gene, UL50, which encodes a dUTPase. Two targeting sgRNAs for the UL50 locus were used to induce site-specific double cleavages (Fig. 3a) and resulted in the formation of an approx. 140 bp-shortened DNA fragment (Fig. 3b). Further sequencing analysis revealed that individual deletions were generated at each of the targeting sites by the sgRNAs and joined by a short segment of original UL50 gene (Fig. 3c). Western analysis of PK15 (Fig. 3d) and Hela (Fig. 3e) cells infected with wild type or mutant viruses further confirmed that UL50 gene was successfully disrupted in the virus. We also showed that UL50 disruption slightly affected the viral growth (Fig. 3f).
Disruption of the PRV UL50 gene by the CRISPR/Cas9 system in PK15 cells. a A schematic diagram showing the UL50 locus in the PRV genome and the targeting sequences of sgRNA1/2 and their respective PAMs. The expected cleavage sites were indicated by the scissors. b PK15 Cells displaying CPE after transfection with Cas9/sgRNAs and PRV genomes in different combinations as indicated were collected for PCR analysis using the primers as indicated in the diagram above. c An example of sequence chromatogram showing UL50 disruption via genomic deletion mediated by two sgRNAs. The red markers point to the junction sites repaired. d and e PK15 d and Hela e cells were either mock infected or infected with WT or UL50 disrupted mutant (designated as UL50 KO) viruses for 24 h and then lysed for Western analysis with the antibodies against viral proteins UL50 and US3 respectively. f PK15 cells grown in a 24-well plate were inoculated with WT or UL50 KO viruses at an MOI of 1. The culture supernatants were collected at the indicated time points, and used to determine the viral titers. Error bars represent the standard errors of the mean of two replicates
Targeted knock-in of RFP expression cassette into the PRV genome using CRISPR/Cas9 system
We next explored the approach to knock-in a foreign gene at the target site of PRV genome using the CRISPR/Cas9 system in PK15 cells. For easy detection, we used RFP as a reporter gene and the EP0 gene as the targeting locus. We first exploited HR-mediated DNA repairing mechanism to knock-in RFP by providing a homologous repair donor. However, we failed to generate any RFP positive viruses. Inspired by the published reports showing that a linearized foreign gene was efficiently captured at a DNA double strand break (DSB) in the genomes of mammalian cells and zebrafish through a homology-independent DNA repair mechanism (Auer et al. 2014; Cristea et al. 2013), we therefore examined whether this system was efficient for large DNA virus. We constructed a donor plasmid which carries a segment of EP0 sequences encompassing the two Cas9 targeting sequences of EP0, known as the bait sequence, followed by a RPF reporter gene cassette. We co-transfected the donor plasmid together with Cas9/sgRNAs vectors and PRV genomes into PK15 cells. The concurrent cleavage of donor plasmid DNA and the targeted viral genome by Cas9 may introduce the RFP cassette into EP0 locus as shown in (Fig. 4a). Indeed, we isolated the progeny viruses carrying the knock-in cassette of RFP using an endpoint dilution assay with efficiency close to 50 %. The RFP knock-in virus was verified by PCR amplification (Fig. 4b) using integration site-specific primers and fluorescence microscopy (Fig. 4c). Subsequent analysis of the junction sequences revealed indel events, indicating that the error-prone non-homologous end joining (NHEJ) mediated DNA repair mechanism has taken place (data not shown). Western analysis further confirmed the expression of RFP and disruption of EP0 in PRV-RFP infected PK15 and Hela cells without affecting the expression of the other viral gene, US3 (Fig. 4d, e).
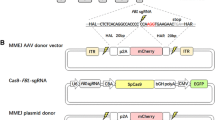
CRISPR/Cas9-mediated knock-in of red fluorescent protein (RFP) cassette into the PRV EP0 locus. a A schematic diagram depicting the likely mechanism by which the RFP cassettes is inserted into the EP0 locus of PRV genome via providing a donor plasmid to induce error-prone non-homologous end joining DNA repair. The donor plasmid contains a bait sequence, which is the short fragment of EP0 locus encompassing the targeting sequences of sgRNAs, followed by the RFP cassette. b PK15 cells infected with WT or the isolated PRV-RFP mutant were collected for PCR analysis using the primers as shown in (a). c PK15 cells were infected with the isolated PRV-RFP mutant, fixed and visualized by immunofluorescence. The nucleus was detected by DAPI straining, Scale bar 30 µm. d and e PK15 d and Hela e cells were either mock infected, or infected with WT or PRV-RFP viruses, or transfected with the expression plasmid pmRFP for 24 h and then lysed for Western analysis with the antibodies against viral proteins EP0 or US3 or RFP
Although homologous recombination mechanism is frequently used to insert a certain gene at the precise genomic site, its efficiency is generally low. It has been demonstrated in mammalian cells and zebrafish that the concurrent cleavage of donor DNA and the targeted genome can effectively integrate the donor DNA in the desired genome through NHEJ (Auer et al. 2014; Cristea et al. 2013). We showed that this approach was also able to efficiently knock-in a foreign gene in PRV genome with relatively high efficiency in PK15 cells. This approach is very convenient, and requires only synthesizing a short bait sequence which contains a pair of Cas9 targeting sequences and then inserting it into a vector preceding the gene expression cassette. In principle, this approach will permit us to knock-in any genes including the rescue mutant of a viral essential gene.
In summary, we developed methods which allowed us to successfully manipulate PRV genome by using the CRISPR/Cas9 system in PK15 cells, a cell line with poor transfection efficiency, in a single step, lending supports that the CRISPR/Cas9 system is a powerful tool for viral DNA engineering.
References
Auer TO, Duroure K, De Cian A, Concordet JP, Del Bene F (2014) Highly efficient CRISPR/Cas9-mediated knock-in in zebrafish by homology-independent DNA repair. Genome Res 24:142–153
Bi Y, Sun L, Gao D, Ding C, Li Z, Li Y, Cun W, Li Q (2014) High-efficiency targeted editing of large viral genomes by RNA-guided nucleases. PLoS Pathog 10:e1004090
Boldogkoi Z, Nogradi A (2003) Gene and cancer therapy–pseudorabies virus: a novel research and therapeutic tool? Curr Gene Ther 3:155–182
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X, Xiong JW, Xi JJ (2013) Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res 23:465–472
Cristea S, Freyvert Y, Santiago Y, Holmes MC, Urnov FD, Gregory PD, Cost GJ (2013) In vivo cleavage of transgene donors promotes nuclease-mediated targeted integration. Biotechnol Bioeng 110:871–880
Dong B, Zarlenga DS, Ren X (2014) An overview of live attenuated recombinant pseudorabies viruses for use as novel vaccines. J Immunol Res 2014:824630
Knapp AC, Enquist LW (1997) Pseudorabies virus recombinants expressing functional virulence determinants gE and gI from bovine herpesvirus 1.1. J Virol 71:2731–2739
Nakade S, Tsubota T, Sakane Y, Kume S, Sakamoto N, Obara M, Daimon T, Sezutsu H, Yamamoto T, Sakuma T, Suzuki KT (2014) Microhomology-mediated end-joining-dependent integration of donor DNA in cells and animals using TALENs and CRISPR/Cas9. Nat Commun 5:5560
Smith GA, Enquist LW (1999) Construction and transposon mutagenesis in Escherichia coli of a full-length infectious clone of pseudorabies virus, an alphaherpesvirus. J Virol 73:6405–6414
Smith GA, Enquist LW (2000) A self-recombining bacterial artificial chromosome and its application for analysis of herpesvirus pathogenesis. Proc Natl Acad Sci USA 97:4873–4878
Suenaga T, Kohyama M, Hirayasu K, Arase H (2014) Engineering large viral DNA genomes using the CRISPR-Cas9 system. Microbiol Immunol 58:513–522
Acknowledgments
We thank Dr. Zhengfan Jiang at Peking University and Dr. Zhongde Wang at Utah State University for generously providing reagents and critically reading the manuscript respectively. This work was supported by the research fund from the State Key Laboratory of Agrobiotechnology of China (2015SKLAB6-12) and the Scientific Research Foundation for the Returned Overseas Chinese Scholars, State Education Ministry.
Supporting information
Supplementary Table 1—DNA sequences of sgRNAs used for plasmid construction.
Supplementary Fig. 1—Analysis of the specificity of the antibody produced US3, EP0 and UL50. (A) Western blot analysis of FLAG-US3 expressed in 293T cells. Cell lysates 293T cells transfected with plasmid expressing FLAG-US3 or FLAG-vector were probed with the mouse antibody (left panel) and then re-probed with rabbit anti-FLAG antibody (right panel). (B) Western blot analysis of FLAG-EP0 expressed in 293T cells. Cell lysates 293T cells transfected with plasmid expressing FLAG-EP0 or FLAG-vector were probed with the mouse antibody (left panel) and then re-probed with rabbit anti-FLAG antibody (right panel). (C) Western blot analysis of FLAG-UL50 expressed in293T cells. Cell lysates 293T cells transfected with plasmid expressing FLAG-UL50 or FLAG-vector were probed with the mouse antibody (left panel) and then re-probed with rabbit anti-FLAG antibody (right panel).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xu, A., Qin, C., Lang, Y. et al. A simple and rapid approach to manipulate pseudorabies virus genome by CRISPR/Cas9 system. Biotechnol Lett 37, 1265–1272 (2015). https://doi.org/10.1007/s10529-015-1796-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1796-2