Abstract
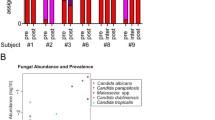
Given the complexity of the airway microbiota in the respiratory tract of cystic fibrosis (CF) patients, it seems crucial to compile the most exhaustive and exact list of the microbial communities inhabiting CF airways. The aim of the present study was to compare the bacterial and fungal diversity of sputa from adult CF patients during non-exacerbation period by culture-based and molecular methods, and ultra-deep-sequencing (UDS). Sputum samples from four CF patients were cultured and analysed by DNA extractions followed by terminal restriction fragment length polymorphism analysis through resolution of bacterial ribosomal gene (rDNA) fragments, and cloning plus sequencing of part of fungal rRNA genes. These approaches were compared with UDS method targeting 16S rDNA gene and the internal transcribed spacer (ITS) 2 region of rDNA. A total of 27 bacterial and 18 fungal genera were detected from the four patients. Five (18%) and 3 (16%) genera were detected by culture for bacteria and fungi, respectively, 9 (33%) and 3 (16%) by first generation sequencing (FGS) methods, and 26 (96%) and 18 (100%) by UDS. The mean number of genera detected by UDS per patient was statistically higher than by culture or FGS methods. Patients with severe airway disease as assessed by standard spirometry exhibited a reduced fungal and bacterial diversity. UDS approach evaluates more extensively the diversity of fungal and bacterial flora compared with cultures. However, it currently remains difficult to routinely use UDS mainly because of the lack of standardization, and the current cost of this method.




Similar content being viewed by others
References
Vaincre la Mucoviscidose. Registre français de la mucoviscidose - Bilan des données 2015. [Internet]. 2015 [cited 2014 Apr 11]. Available from: http://www.vaincrelamuco.org/sites/default/files/registre_francais_de_la_mucoviscidose_-_bilan_2015_v4.pdf.
Waters V, Ratjen F. Antibiotic treatment for nontuberculous mycobacteria lung infection in people with cystic fibrosis. Cochrane Database Syst Rev. 2016;12:CD010004.
Salvatore D, Buzzetti R, Mastella G. An overview of international literature from cystic fibrosis registries. Part 5: Update 2012-2015 on lung disease. Pediatr Pulmonol. 2016;51:1251–63.
European Cystic Fibrosis Society. ECFS Patient Registry Annual Data Report 2014 [Internet]. 2014 [cited 2017 Apr 11]. Available from: https://www.ecfs.eu/projects/ecfs-patient-registry/annual-reports.
LiPuma JJ. The changing microbial epidemiology in cystic fibrosis. Clin Microbiol Rev. 2010;23:299–323.
Huang YJ, LiPuma JJ. The microbiome in cystic fibrosis. Clin Chest Med. 2016;37:59–67.
Parkins MD, Floto RA. Emerging bacterial pathogens and changing concepts of bacterial pathogenesis in cystic fibrosis. J Cyst Fibros. 2015;14:293–304.
Tunney MM, Field TR, Moriarty TF, et al. Detection of anaerobic bacteria in high numbers in sputum from patients with cystic fibrosis. Am J Respir Crit Care Med. 2008;177:995–1001.
Borman AM, Palmer MD, Delhaes L, et al. Lack of standardization in the procedures for mycological examination of sputum samples from CF patients: a possible cause for variations in the prevalence of filamentous fungi. Med Mycol. 2010;48:S88–97.
Williams C, Ranjendran R, Ramage G. Pathogenesis of fungal infections in cystic fibrosis. Curr Fungal Infect Rep. 2016;10:163–9.
Sudfeld CR, Dasenbrook EC, Merz WG, Carroll KC, Boyle MP. Prevalence and risk factors for recovery of filamentous fungi in individuals with cystic fibrosis. J Cyst Fibros. 2010;9:110–6.
Rogers GB, Carroll MP, Serisier DJ, et al. Characterization of bacterial community diversity in cystic fibrosis lung infections by use of 16S ribosomal DNA terminal restriction fragment length polymorphism profiling. J Clin Microbiol. 2004;42:5176–83.
Rogers GB, Skelton S, Serisier DJ, van der Gast CJ, Bruce KD. Determining cystic fibrosis-affected lung microbiology: comparison of spontaneous and serially induced sputum samples by use of terminal restriction fragment length polymorphism profiling. J Clin Microbiol. 2010;48:78–86.
Suhr MJ, Banjara N, Hallen-Adams HE. Sequence-based methods for detecting and evaluating the human gut mycobiome. Lett Appl Microbiol. 2016;62:209–15.
Armougom F, Bittar F, Stremler N, et al. Microbial diversity in the sputum of a cystic fibrosis patient studied with 16S rDNA pyrosequencing. Eur J Clin Microbiol Infect Dis. 2009;28:1151–4.
Guss AM, Roeselers G, Newton ILG, et al. Phylogenetic and metabolic diversity of bacteria associated with cystic fibrosis. ISME J. 2011;5:20–9.
Delhaes L, Monchy S, Fréalle E, et al. The airway microbiota in cystic fibrosis: a complex fungal and bacterial community–implications for therapeutic management. PLoS ONE. 2012;7(4):e36313.
Kramer R, Sauer-Heilborn A, Welte T, et al. Cohort study of airway mycobiome in adult cystic fibrosis patients: differences in community structure between fungi and bacteria reveal predominance of transient fungal elements. J Clin Microbiol. 2015;53:2900–7.
Willger SD, Grim SL, Dolben EL, et al. Characterization and quantification of the fungal microbiome in serial samples from individuals with cystic fibrosis. Microbiome. 2014;2:40.
Mahboubi MA, Carmody LA, Foster BK, et al. Culture-based and culture-independent bacteriologic analysis of cystic fibrosis respiratory specimens. J Clin Microbiol. 2016;54:613–9.
Daniels TWV, Rogers GB, Stressmann FA, et al. Impact of antibiotic treatment for pulmonary exacerbations on bacterial diversity in cystic fibrosis. J Cyst Fibros. 2013;12:22–8.
White TJ, Bruns L, Lee S, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: A Guide to Methods and Applications. New York: Academic Press; 1990. p. 315– 322.
Sitterlé E, Rodriguez C, Mounier R, et al. Contribution of ultra deep sequencing in the clinical diagnosis of a new fungal pathogen species: Basidiobolus meristosporus. Front Microbiol. 2017;8:334.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
Wang Q, Garrity GM, Tiedje JM, Cole JR. Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–7.
van der Gast CJ, Walker AW, Stressmann FA, et al. Partitioning core and satellite taxa from within cystic fibrosis lung bacterial communities. ISME J. 2011;5:780–90.
Coburn B, Wang PW, Diaz Caballero J, et al. Lung microbiota across age and disease stage in cystic fibrosis. Sci Rep. 2015;5.
Blainey PC, Milla CE, Cornfield DN, Quake SR. Quantitative analysis of the human airway microbial ecology reveals a pervasive signature for cystic fibrosis. Sci Transl Med. 2012;4:153ra130.
Huffnagle GB, Noverr MC. The emerging world of the fungal microbiome. Trends Microbiol. 2013;21:334–41.
Nguyen LDN, Viscogliosi E, Delhaes L. The lung mycobiome: an emerging field of the human respiratory microbiome. Front Microbiol. 2015;6:89.
Zakharkina T, Heinzel E, Koczulla RA, et al. Analysis of the airway microbiota of healthy individuals and patients with chronic obstructive pulmonary disease by T-RFLP and clone sequencing. PLoS ONE. 2013;8(7):e68302.
Aas JA, Paster BJ, Stokes LN, Olsen I, Dewhirst FE. Defining the normal bacterial flora of the oral cavity. J Clin Microbiol. 2005;43:5721–32.
Sherrard LJ, Bell SC, Tunney MM. The role of anaerobic bacteria in the cystic fibrosis airway. Curr Opin Pulm Med. 2016;22:637–43.
Kashinskaya EN, Andree KB, Simonov EP, Solovyev MM. DNA extraction protocols may influence biodiversity detected in the intestinal microbiome: a case study from wild Prussian carp, Carassius gibelio. FEMS Microbiol Ecol. 2017;93(2). pii:fiw240.
Vesty A, Biswas K, Taylor MW, Gear K, Douglas RG. Evaluating the impact of DNA extraction method on the representation of human oral bacterial and fungal communities. PLoS ONE. 2017;12(1):e0169877.
Rogers GB, Bruce KD. Next-generation sequencing in the analysis of human microbiota: essential considerations for clinical application. Mol Diagn Ther. 2010;14:343–50.
Doré J, Ehrlich SD, Levenez F, et al. IHMS_SOP 06 V1: Standard operating procedure for fecal samples DNA extraction, Protocol Q. [Internet]. International Human Microbiome Standards; 2015. Available from: http://www.microbiome-standards.org.
Schoch CL, Seifert KA, Huhndorf S, et al. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA. 2012;109:6241c6.
Lagier J-C, Armougom F, Million M, et al. Microbial culturomics: paradigm shift in the human gut microbiome study. Clin Microbiol Infect. 2012;18:1185–93.
Abdallah RA, Beye M, Diop A, et al. The impact of culturomics on taxonomy in clinical microbiology. Antonie Van Leeuwenhoek. 2017;. doi:10.1007/s10482-017-0871-1.
Vallet S. Viruses in cystic fibrosis patients’ airways. Crit Rev Microbiol. 2017;24:1–19.
Stevens DA, Moss RB, Kurup VP, et al. Allergic bronchopulmonary aspergillosis in cystic fibrosis–state of the art: cystic Fibrosis Foundation Consensus Conference. Clin Infect Dis. 2003;37(Suppl 3):S225–64.
Acknowledgements
We would like to thank Damian Rivett and Jean-Philippe Barnier for their technical assistance, and Genoscreen Society for their critical and technical support.
Funding
This work was supported in part by the Assistance Publique-Hôpitaux de Paris (public sector, bourse de mobilité), and Pfizer France Pharmaceutical Division. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. L. Delhaes has received research grants from the French Ministry of Health and Research (PHRC N°2006/1902), Centre Hospitalier Régional Universitaire de Lille, the association “Vaincre la Mucoviscidose” (MucoFong and Mucofong-ATF N8 2006/351), and Pfizer France Pharmaceutical Division (Nu 2006/158).
Author information
Authors and Affiliations
Contributions
Authors’ Contributions
FW, BW and LD collected sputum samples and clinical data. FB, OC, JMC, FS and LD prepared sputum samples for molecular analysis including UDS. CA performed the statistical analyses including that of UDS. FB and LD contributed to study design. FB was in charge of all the molecular approaches. FB and CA wrote the manuscript with significant contributions of co-authors, especially KB and LD. All authors read and gave approval on the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
FB received grants from Astellas, Pfizer, and payment for lectures from Merck and Gilead.
Ethical Approval
All procedures performed in this study involving human participants were in compliance with the ethical standards of the institutional research committee (Comité de Protection des Personnes of Lille University Hospital, CPP 06/84) and with the 1964 Helsinki Declaration and its latest amendments.
Informed Consent
Informed consent forms were obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Botterel, F., Angebault, C., Cabaret, O. et al. Fungal and Bacterial Diversity of Airway Microbiota in Adults with Cystic Fibrosis: Concordance Between Conventional Methods and Ultra-Deep Sequencing, and Their Practical use in the Clinical Laboratory. Mycopathologia 183, 171–183 (2018). https://doi.org/10.1007/s11046-017-0185-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11046-017-0185-x