Abstract
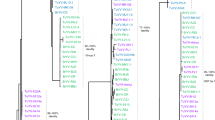
The diversity of ZYMV isolates was analysed by the biological and molecular characterisation of 11 isolates sampled from cucumber, squash and zucchini between 2001 and 2006 in various localities of Slovakia and Czech Republic. Analysis of the molecular variability targeting three separate genomic regions of the ZYMV genome [P1, P3 and (Cter)NIb-(Nter)CP] revealed a remarkable low level of nucleotide variability between isolates, despite their temporal and spatial distinction. Phylogenetic analysis based on the 5′-terminal part of the CP gene highlighted the close relatedness of Slovak, Czech and other central European isolates. Low level of genetic diversity within central European ZYMV isolates is in contrast to the diversity observed for isolates from other geographical regions, in particular Asia. No evidence of recombination in the ZYMV genome was detected. Sequence comparison between aggressive and moderate ZYMV isolates revealed one amino acid difference in the N-terminal part of the P3 protein, potentially involved in the tolerance breaking.



Similar content being viewed by others
References
V. Lisa, G. Boccardo, G. D’Agostino, G. Dellavalle, M. d’Aquilio, Phytopathology 71, 667 (1981)
C. Desbiez, H. Lecoq, Plant Pathol 46, 809 (1997)
J. Chod, M. Jokeš, Ochr Rostlin 27, 111 (1991)
I. Tobias, L. Palkovics, Pest Manag Sci 59, 493 (2003)
F. Revers, O. Le Gall, T. Candresse, A.J. Maule, Mol Plant Microbe Interact 12, 367 (1999)
C. Desbiez, A. Gal-On, M. Girard, C. Wipf-Scheibel, H. Lecoq, Phytopathology 93, 1478 (2003)
G. Labonne, M. Yvon, J.B. Quiot, L. Avinent, G. Llacer, Acta Hortic 386, 207 (1995)
J.D. Thompson, T.J. Gibson, F. Plewniak, F. Jeanmougin, D.G. Higgins, Nucleic Acids Res 24, 4876 (1997)
S. Kumar, K. Tamura, M. Nei, Brief Bioinform 5, 150 (2004)
M. Glasa, S. Pittnerová, J Phytopathol 154, 436 (2006)
C.D. Atreya, B. Raccah, T.P. Pirone, Virology 178, 161 (1990)
M.F. Pfosser, H. Baumann, Arch Virol 147, 1599 (2002)
S.W. Kwon, M.S. Kim, H.S. Choi, K.H. Kim, J Gen Plant Pathol 71, 80 (2005)
S.S. Lin, R.F. Hou, S.D. Yeh, Phytopathology 90, 228 (2000)
M.F. Zhao, J. Chen, H.Y. Zheng, M.J. Adams, J.P. Chen, J Phytopathol 151, 307 (2003)
E.P. Rybicki, D.D. Shukla, Arch Virol (Suppl 5), 139 (1992)
L. Glais, M. Tribodet, C. Kerlan, Arch Virol 147, 363 (2002)
M. Glasa, L. Palkovics, P. Komínek, G. Labonne, S. Pittnerová, O. Kúdela, T. Candresse, Z. Šubr J Gen Virol 85, 2671 (2004)
K. Tomimura, A.J. Gibbs, C.E. Jenner, J.A. Walsh, K. Ohshima, Mol Ecol 12, 2099 (2003)
S. Dallot, L. Quiot-Douine, P. Saenz, M.T. Cervera, J.A. Garcia, J.B. Quiot, Phytopathology 91, 159 (2001)
N. Suehiro, T. Natsuaki, T. Watanabe, S. Okuda, J Gen Virol 85, 2087 (2004)
P. Saenz, M.T. Cervera, S. Dallot, L. Quiot, J.B. Quiot, J.L. Riechmann, J.A. Garcia, J Gen Virol 81, 557 (2000)
C.E. Jenner, X. Wang, K. Tomimura, K. Ohshima, F. Ponz, J.A. Walsh, Mol Plant Microbe Interact 16, 777 (2003)
E. Domingo, J Virol 76, 463 (2002)
A. Ali, H. Li, W.L. Schneider, D.J. Sherman, S. Gray, D. Smith, M. Roossinck, J Virol 80, 8345 (2006)
H. Huet, A. Gal-On, E. Meir, H. Lecoq, B. Raccah, J Gen Virol 75, 1407 (1994)
J. Svoboda, J. Polák, Plant Protect Sci 38, 125 (2002)
Acknowledgements
The authors thank Drs H. Lecoq and C. Desbiez for the generous gift of antibodies used for serological detection of cucurbit viruses and Dr T. Candresse for critical reading of manuscript. This work was supported by the grant APVT-51-001304 from the Slovak Research and Development Agency and partially by the VEGA grant 2/7006/27. J.S. receives a support from the Ministry of Agriculture, Czech Republic (project no. 0002700603).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Glasa, M., Svoboda, J. & Nováková, S. Analysis of the molecular and biological variability of Zucchini yellow mosaic virus isolates from Slovakia and Czech Republic. Virus Genes 35, 415–421 (2007). https://doi.org/10.1007/s11262-007-0101-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-007-0101-4