Abstract
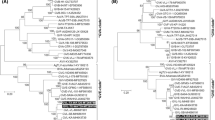
Australian grapevine viroid (AGVd) is found in only three countries in the world. Here, the genetic diversity and phylogenetic relationships of AGVd isolates from three different grape varieties (Thomson Seedless, Jingchuan and Zaoyu) in China were studied. A hundred of independent cDNA clones from each of the three isolates, in total of 300, were sequenced. We identified 29 sequence variants including two predominant ones in Thomson Seedless, and 48 each including a unique predominant one in Jingchuan and Zaoyu. In silico structure analysis revealed that base changes were clustered in the left terminal domain of the predicted secondary structure in all three isolates. Further, these changes were shown to affect their secondary structures to varying degrees. Genetic diversity and phylogenetic analysis of four predominant sequence variants from this study, plus four others from Australia and Tunisia, revealed obvious regional disparity and variety-specificity in AGVd.





Similar content being viewed by others
References
C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball (eds.), Virus Taxonomy. Eighth Report of the International Committee on Taxonomy of Viruses (Elsevier, San Diego, 2005), pp. 1145–1160
M.A. Rezaian, A.M. Koltunow, L.R. Krake, J. Gen. Virol. 69, 413–422 (1988). doi:https://doi.org/10.1099/0022-1317-69-2-413
J.A. Szychowski, A.C. Goheen, J.S. Semancik, J. Am. Enol. 39, 213–216 (1988)
A.M. Koltunow, M.A. Rezaian, Intervirology 30, 194–201 (1989)
M.A. Rezaian, Nucleic Acids Res. 18, 1813–1818 (1990). doi:https://doi.org/10.1093/nar/18.7.1813
A. Elleuch, H. Fakhfakh, M. Pelchat, P. Landry, M. Marrakchi, J.P. Perreault, Eur. J. Plant Pathol. 108, 815–820 (2002). doi:https://doi.org/10.1023/A:1020855405948
S.F. Li, R. Guo, M. Tsuji, T. Sano, Plant. Pathol. 55, 564–564 (2006). doi:https://doi.org/10.1111/j.1365-3059.2006.01424.x
S.F. Li, S. Onodera, T. Sano, K. Yoshida, G.P. Wang, E. Shikata, Ann. Phytopathological Soc. Jpn. 61, 9 (1995)
A. Iijima, Shokubutsu Boeki 44, 130–132 (1990)
E.J.J.H. Domingo, Annu. Rev. Microbiol. 51, 151–178 (1997). doi:https://doi.org/10.1146/annurev.micro.51.1.151
C. Domingo, V. Conejero, P. Vera, Plant Mol. Biol. 24, 725–732 (1994). doi:https://doi.org/10.1007/BF00029854
M. Gandia, L. Rubio, A. Palacio, N. Duran-Vila, Arch. Virol. 150, 1945–1957 (2005). doi:https://doi.org/10.1007/s00705-005-0570-5
S. Ambros, J.C. Desvignes, G. Llacer, R. Flores, J. Gen. Virol. 76, 2625–2629 (1995). doi:https://doi.org/10.1099/0022-1317-76-10-2625
S. Ambros, C. Hernandez, J.C. Desvignes, R. Flores, J. Virol. 72, 7397–7406 (1998)
A. Palacio-Bielsa, J. Romero-Durban, N. Duran-Vila, Arch. Virol. 149, 537–552 (2004). doi:https://doi.org/10.1007/s00705-003-0223-5
X. Foissac, N. Duran-Vila, Arch. Virol. 145, 1975–1983 (2000). doi:https://doi.org/10.1007/s007050070070
H. Polivka, U. Staub, H.J. Gross, J. Gen. Virol. 77, 155–161 (1996). doi:https://doi.org/10.1099/0022-1317-77-1-155
T. Sano, R. Mimura, K. Ohshima, Virus Genes 22, 53 (2001). doi:https://doi.org/10.1023/A:1008182302704
W.X. Xu, N. Hong, G.P. Wang, X. Fan, J. Phytopathol. 156(9), 565–572 (2008). doi:https://doi.org/10.1111/j.1439-0434.2008.01398.x
J.E. Visvader, R.H. Symons, Nucleic Acids Res. 13, 2907–2920 (1985). doi:https://doi.org/10.1093/nar/13.8.2907
M.A. Bracho, E. Barrio, J. Gen. Virol. 79, 2921–2928 (1998)
A.J. Dingley, G. Steger, B. Esters, D. Riesner, S. Grzesiek, J. Mol. Biol. 334, 751–767 (2003). doi:https://doi.org/10.1016/j.jmb.2003.10.015
J.S. Semancik, G. Vidalakis, Arch. Virol. 150, 1059 (2005). doi:https://doi.org/10.1007/s00705-005-0499-8
P. Keese, R.H. Symons, Proc. Natl Acad. Sci. USA 82, 4582–4586 (1985). doi:https://doi.org/10.1073/pnas.82.14.4582
Acknowledgments
This work was supported by grants from the National Basic Research and Development Program of China (973 Program) (No. 2006CB100203 and No. 2009CB119200), the National Natural Science Foundation of China (No.30771403) and the Beijing Natural Science Foundation of China (No. 6072022), and the Opening Project of State Key Laboratory for Biology of Plant Diseases and Insect Pests, Institute of Plant Protection, Chinese Academy of Agricultural Sciences. This work was also supported by NSFC-JSPS Joint Research Project between China and Japan (30811140157). We specially thank Prof. Teruo Sano at Hirosaki University of Japan for his valuable comments and the critical reading of this manuscript. We also thank Mr. Lingxiao Mu, graduate student of Shenyang Agricultural University, for his assistance in artwork preparation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, D., Peng, S., Wu, Z. et al. Genetic diversity and phylogenetic analysis of Australian Grapevine Viroid (AGVd) isolated from different grapevines in China. Virus Genes 38, 178–183 (2009). https://doi.org/10.1007/s11262-008-0306-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-008-0306-1



