Abstract
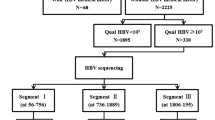
Since 1992, China has promoted hepatitis B vaccination. Concurrently, during this period, increasing use of immunoglobulins and nucleoside analogues might have exerted selective pressure on the hepatitis B virus (HBV) S gene, driving mutations in the HBsAg and changed the subtype. Using the National Center for Biotechnology Information database, we obtained gene sequence information for HBV strains from China and analysed changes in HBsAg subtypes and substitution mutations in HBsAg in 5-year intervals over 25 years to identify potential challenges to the prevention and treatment of hepatitis B. Most HBV sequences from China were genotype C (1996/2833, 70.46%) or B (706/2833, 24.92%). During the implementation of hepatitis B vaccination (recombinant hepatitis B vaccine was subgenotype A2 and HBsAg subtype adw2), the proportion of subtypes ayw1 and adw3 in genotype B and ayw2 in genotype C increased over the programme period. The overall mutation rate in HBsAg tended to decrease for genotype B, whereas, for genotype C, the rate increased gradually and then decreased slightly. Moreover, the mutation rate at some HBsAg amino acid sites (such as sG145 of genotype B and sG130 and sK141 of genotype C) is gradually increasing. HBV strains with internal stop codons of HBsAg (e.g., sC69*) and additional N-glycosylation (e.g., sG130N) mutations should be studied extensively to prevent them from becoming dominant circulating strains. The development of HBV vaccines and antiviral immunoglobulins and use of antiviral drugs may require making corresponding changes.


Similar content being viewed by others
Data availability
The datasets generated and analyzed during this study are available with the corresponding author, on reasonable request.
References
Uz-Zaman MH, Rahman A, Yasmin M (2018) Epidemiology of hepatitis B virus infection in Bangladesh: prevalence among general population, risk groups and genotype distribution. Genes (Basel) 9(11):2–16. https://doi.org/10.3390/genes9110541
Kramvis A (2014) Genotypes and genetic variability of hepatitis B virus. Intervirology 57(3–4):141–150. https://doi.org/10.1159/000360947
Purdy MA, Talekar G, Swenson P et al (2007) A new algorithm for deduction of hepatitis B surface antigen subtype determinants from the amino acid sequence. Intervirology 50(1):45–51. https://doi.org/10.1159/000096312
Gerlich WH, Glebe D, Kramvis A et al (2020) Peculiarities in the designations of hepatitis B virus genes, their products, and their antigenic specificities: a potential source of misunderstandings. Virus Genes 56(2):109–119. https://doi.org/10.1007/s11262-020-01733-9
Norder H, Courouce AM, Coursaget P et al (2004) Genetic diversity of hepatitis B virus strains derived worldwide: genotypes, subgenotypes, and HBsAg subtypes. Intervirology 47(6):289–309. https://doi.org/10.1159/000080872
Ohnuma H, Machida A, Okamoto H et al (1993) Allelic subtypic determinants of hepatitis B surface antigen (i and t) that are distinct from d/y or w/r. J Virol 67(2):927–932
Yu DM, Li XH, Mom V et al (2014) N-glycosylation mutations within hepatitis B virus surface major hydrophilic region contribute mostly to immune escape. J Hepatol 60(3):515–522. https://doi.org/10.1016/j.jhep.2013.11.004
Ho JK, Jeevan-Raj B, Netter HJ (2020) Hepatitis B virus (HBV) subviral particles as protective vaccines and vaccine platforms. Viruses 12(2):1–26. https://doi.org/10.3390/v12020126
Bian T, Yan H, Shen L et al (2013) Change in hepatitis B virus large surface antigen variant prevalence 13 years after implementation of a universal vaccination program in China. J Virol 87(22):12196–12206. https://doi.org/10.1128/JVI.02127-13
Gerlich WH (2015) Prophylactic vaccination against hepatitis B: achievements, challenges and perspectives. Med Microbiol Immunol 204(1):39–55. https://doi.org/10.1007/s00430-014-0373-y
Ni YH, Chang MH, Wu JF et al (2012) Minimization of hepatitis B infection by a 25-year universal vaccination program. J Hepatol 57(4):730–735. https://doi.org/10.1016/j.jhep.2012.05.021
Pancher M, Desire N, Ngo Y et al (2015) Coexistence of circulating HBsAg and anti-HBs antibodies in chronic hepatitis B carriers is not a simple analytical artifact and does not influence HBsAg quantification. J Clin Virol 62:32–37. https://doi.org/10.1016/j.jcv.2014.11.015
Sayiner AA, Agca H, Sengonul A et al (2007) A new hepatitis B virus vaccine escape mutation in a renal transplant recipient. J Clin Virol 38(2):157–160. https://doi.org/10.1016/j.jcv.2006.12.002
Chen J, Liu Y, Zhao J et al (2016) Characterization of novel hepatitis B virus PreS/S-gene mutations in a patient with occult hepatitis B virus infection. PLoS ONE 11(5):e0155654. https://doi.org/10.1371/journal.pone.0155654
Su M, Xiang K, Li Y et al (2016) Higher detection rates of amino acid substitutions in HBV reverse transcriptase/surface protein overlapping sequence is correlated with lower serum HBV DNA and HBsAg levels in HBeAg-positive chronic hepatitis B patients with subgenotype B2. Infect Genet Evol 40:275–281. https://doi.org/10.1016/j.meegid.2016.03.019
Yonghao G, Jin X, Jun L et al (2015) An epidemiological serosurvey of hepatitis B virus shows evidence of declining prevalence due to hepatitis B vaccination in central China. Int J Infect Dis 40:75–80. https://doi.org/10.1016/j.ijid.2015.10.002
Gerlich WH (2017) Do we need better hepatitis B vaccines? Indian J Med Res 145(4):414–419. https://doi.org/10.4103/ijmr.IJMR_1852_16
Hsu HY, Chang MH, Liaw SH et al (1999) Changes of hepatitis B surface antigen variants in carrier children before and after universal vaccination in Taiwan. Hepatology 30(5):1312–1317. https://doi.org/10.1002/hep.510300511
Tallo T, Tefanova V, Priimagi L et al (2008) D2: major subgenotype of hepatitis B virus in Russia and the Baltic region. J Gen Virol 89(Pt 8):1829–1839. https://doi.org/10.1099/vir.0.83660-0
Kramvis A, Kew M, Francois G (2005) Hepatitis B virus genotypes. Vaccine 23(19):2409–2423. https://doi.org/10.1016/j.vaccine.2004.10.045
Bartholomeusz A, Schaefer S (2004) Hepatitis B virus genotypes: comparison of genotyping methods. Rev Med Virol 14(1):3–16. https://doi.org/10.1002/rmv.400
Delport W, Poon AF, Frost SD et al (2010) Datamonkey 2010: a suite of phylogenetic analysis tools for evolutionary biology. Bioinformatics 26(19):2455–2457. https://doi.org/10.1093/bioinformatics/btq429
Pan XF, Griffiths UK, Pennington M et al (2015) Systematic review of economic evaluations of vaccination programs in mainland China: are they sufficient to inform decision making? Vaccine 33(46):6164–6172. https://doi.org/10.1016/j.vaccine.2015.09.081
Ponde RA (2011) The underlying mechanisms for the “simultaneous HBsAg and anti-HBs serological profile”. Eur J Clin Microbiol Infect Dis 30(11):1325–1340. https://doi.org/10.1007/s10096-011-1240-z
Fu X, Chen J, Chen H et al (2017) Mutation in the S gene of hepatitis B virus and anti-HBs subtype-nonspecificity contributed to the coexistence of HBsAg and anti-HBs in patients with chronic hepatitis B virus infection. J Med Virol 89(8):1419–1426. https://doi.org/10.1002/jmv.24782
Cassidy A, Mossman S, Olivieri A et al (2011) Hepatitis B vaccine effectiveness in the face of global HBV genotype diversity. Expert Rev Vaccines 10(12):1709–1715. https://doi.org/10.1586/erv.11.151
Van Den Ende C, Marano C, Van Ahee A et al (2017) The immunogenicity and safety of GSK’s recombinant hepatitis B vaccine in adults: a systematic review of 30 years of experience. Expert Rev Vaccines 16(8):811–832. https://doi.org/10.1080/14760584.2017.1338568
Wen W-H, Chen H-L, Ni Y-H et al (2011) Secular trend of the viral genotype distribution in children with chronic hepatitis B virus infection after universal infant immunization. Hepatology 53(2):429–436. https://doi.org/10.1002/hep.24061
Lee KM, Kim YS, Ko YY et al (2001) Emergence of vaccine-induced escape mutant of hepatitis B virus with multiple surface gene mutations in a Korean child. J Korean Med Sci 16(3):359–362
Verheyen J, Neumann-Fraune M, Berg T et al (2012) The detection of HBsAg mutants expressed in vitro using two different quantitative HBsAg assays. J Clin Virol 54(3):279–281. https://doi.org/10.1016/j.jcv.2012.04.010
Thibault V, Servant-Delmas A, Ly TD et al (2017) Performance of HBsAg quantification assays for detection of Hepatitis B virus genotypes and diagnostic escape-variants in clinical samples. J Clin Virol 89:14–21. https://doi.org/10.1016/j.jcv.2017.02.001
Wang T, Cui D, Chen S et al (2019) Analysis of clinical characteristics and S gene sequences in chronic asymptomatic HBV carriers with low-level HBsAg. Clin Res Hepatol Gastroenterol 43(2):179–189. https://doi.org/10.1016/j.clinre.2018.08.015
Kampfrath T, Jortani SA, Levinson SS (2014) Resolving a case of concurrent hepatitis B virus surface antigen (HBsAg) and surface antibody (HBsAb). Clin Chim Acta 430:171–172. https://doi.org/10.1016/j.cca.2014.01.016
Osiowy C, Kowalec K, Giles E (2016) Discordant diagnostic results due to a hepatitis B virus T123A HBsAg mutant. Diagn Microbiol Infect Dis 85(3):328–333. https://doi.org/10.1016/j.diagmicrobio.2016.04.004
Wang ML, Tang H (2016) Nucleos(t)ide analogues causes HBV S gene mutations and carcinogenesis. Hepatobiliary Pancreat Dis Int 15(6):579–586. https://doi.org/10.1016/s1499-3872(16)60064-4
Colledge D, Soppe S, Yuen L et al (2017) Stop codons in the hepatitis B surface proteins are enriched during antiviral therapy and are associated with host cell apoptosis. Virology 501:70–78. https://doi.org/10.1016/j.virol.2016.11.007
Xiang KH, Michailidis E, Ding H et al (2017) Effects of amino acid substitutions in hepatitis B virus surface protein on virion secretion, antigenicity HBsAg and viral DNA. J Hepatol 66(2):288–296. https://doi.org/10.1016/j.jhep.2016.09.005
Ding H, Liu B, Zhao C et al (2014) Amino acid similarities and divergences in the small surface proteins of genotype C hepatitis B viruses between nucleos(t)ide analogue-naive and lamivudine-treated patients with chronic hepatitis B. Antiviral Res 102:29–34. https://doi.org/10.1016/j.antiviral.2013.11.015
Cento V, Van Hemert F, Neumann-Fraune M et al (2013) Anti-HBV treatment induces novel reverse transcriptase mutations with reflective effect on HBV S antigen. J Infect 67(4):303–312. https://doi.org/10.1016/j.jinf.2013.05.008
Colagrossi L, Hermans LE, Salpini R et al (2018) Immune-escape mutations and stop-codons in HBsAg develop in a large proportion of patients with chronic HBV infection exposed to anti-HBV drugs in Europe. BMC Infect Dis 18(1):251. https://doi.org/10.1186/s12879-018-3161-2
Warner N, Locarnini S (2008) The antiviral drug selected hepatitis B virus rtA181T/sW172* mutant has a dominant negative secretion defect and alters the typical profile of viral rebound. Hepatology 48(1):88–98. https://doi.org/10.1002/hep.22295
Lee SA, Kim K, Kim H et al (2012) Nucleotide change of codon 182 in the surface gene of hepatitis B virus genotype C leading to truncated surface protein is associated with progression of liver diseases. J Hepatol 56(1):63–69. https://doi.org/10.1016/j.jhep.2011.06.028
Hadiji-Abbes N, Mihoubi W, Martin M et al (2015) Characterization of C69R variant HBsAg: effect on binding to anti-HBs and the structure of virus-like particles. Arch Virol 160(10):2427–2433. https://doi.org/10.1007/s00705-015-2515-y
Shirvani-Dastgerdi E, Winer BY, Celia-Terrassa T et al (2017) Selection of the highly replicative and partially multidrug resistant rtS78T HBV polymerase mutation during TDF-ETV combination therapy. J Hepatol 67(2):246–254. https://doi.org/10.1016/j.jhep.2017.03.027
Gerlich WH, Lu X, Heermann KH (1993) Studies on the attachment and penetration of hepatitis B virus. J Hepatol 17(Suppl 3):S10–S14
Qiao Y, Lu SS, Xu ZH et al (2017) Additional N-glycosylation mutation in the major hydrophilic region of hepatitis B virus S gene is a risk indicator for hepatocellular carcinoma occurrence in patients with coexistence of HBsAg/anti-HBs. Oncotarget 8(37):61719–61730. https://doi.org/10.18632/oncotarget.18682
Zhang S, Cao X, Gao Q et al (2017) Protein glycosylation in viral hepatitis-related HCC: characterization of heterogeneity, biological roles, and clinical implications. Cancer Lett 406:64–70. https://doi.org/10.1016/j.canlet.2017.07.026
Acknowledgements
This work was supported by grants from the Fujian Provincial Health and Family Planning Research Talents Training Project (No. 2018-2-67) and Xiamen Municipal Health and Family Planning Commission. Authors have no conflicting interests to declare relevant to this study. We thank Ms. Bihuan Che for her kindly technical support.
Author information
Authors and Affiliations
Contributions
HY and JT conceived of and designed the study; HY, XF, JT, YW, and ZL performed the research; HY, XF, and JT analyzed the data; and XF wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines of ethical approval were followed.
Additional information
Edited by Wolfram H. Gerlich.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ye, H., Teng, J., Lin, Z. et al. Analysis of HBsAg mutations in the 25 years after the implementation of the hepatitis B vaccination plan in China. Virus Genes 56, 546–556 (2020). https://doi.org/10.1007/s11262-020-01773-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-020-01773-1