Abstract
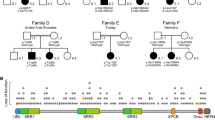
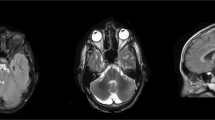
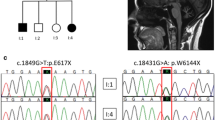
Autosomal recessive spastic ataxia of Charlevoix–Saguenay, more commonly known as ARSACS, is an early-onset cerebellar ataxia with spasticity, amyotrophy, nystagmus, dysarthria, and peripheral neuropathy. SACS is the only gene known to be associated with the ARSACS phenotype. To date, 55 mutations have been reported; of these, only five in Italian patients. We found two novel homozygous nonsense mutations in the giant exon of SACS gene in two unrelated patients with classical ARSACS phenotype. Characterization of the homozygous nature of the mutations through genotyping of the parents, quantitative DNA analysis and indirect STS studies permitted us to confirm in one of the cases that uniparental isodisomy of the paternal chromosome 13 carrying the mutated SACS gene played an etiologic role in the disease.


Similar content being viewed by others
Abbreviations
- STS:
-
Sequence-tagged sites
- UPD:
-
Uniparental disomy
- qPCR:
-
Real-time quantitative PCR
References
Berend SA, Feldman GL, McCaskill C, Czarnecki P, Van Dyke DL, Shaffer LG (1999) Investigation of two cases of paternal disomy 13 suggests timing of isochromosome formation and mechanisms leading to uniparental disomy. Am J Med Genet 82(3):275–281
Criscuolo C, Banfi S, Orio M et al (2004) A novel mutation in SACS gene in a family from southern Italy. Neurology 62(1):100–102
Engert JC, Bérubé P, Mercier J et al (2000) ARSACS, a spastic ataxia common in northeastern Québec, is caused by mutations in a new gene encoding an 11.5-kb ORF. Nat Genet 24(2):120–125
Grieco GS, Malandrini A, Comanducci G et al (2004) Novel SACS mutations in autosomal recessive spastic ataxia of Charlevoix–Saguenay type. Neurology 62(1):103–106
Mercier J, Prévost C, Engert JC, Bouchard JP, Mathieu J, Richter A (2001) Rapid detection of the sacsin mutations causing autosomal recessive spastic ataxia of Charlevoix–Saguenay. Genet Test 5(3):255–259
Parfitt DA, Michael GJ, Vermeulen EG et al (2009) The ataxia protein sacsin is a functional co-chaperone that protects against polyglutamine-expanded ataxin-1. Hum Mol Genet 18(9):1556–1565
Slater H, Shaw JH, Bankier A, Forrest SM, Dawson G (1995) UPD 13: no indication of maternal or paternal imprinting of genes on chromosome 13. J Med Genet 32(6):493
Terracciano A, Casali C, Grieco GS et al (2009) An inherited large-scale rearrangement in SACS associated with spastic ataxia and hearing loss. Neurogenetics 10(2):151–155
Tsai AC, Gibby T, Beischel L, McGavran L, Johnson JP (2004) A child with Angelman syndrome and trisomy 13 findings due to associated paternal UPD 15 and segmental UPD 13. Am J Med Genet A 126A(2):208–212
Acknowledgments
We are grateful to the patients and their families for contributing to this study.
Conflict of Interest
The authors state no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Anesi, L., de Gemmis, P., Pandolfo, M. et al. Two Novel Homozygous SACS Mutations in Unrelated Patients Including the First Reported Case of Paternal UPD as an Etiologic Cause of ARSACS. J Mol Neurosci 43, 346–349 (2011). https://doi.org/10.1007/s12031-010-9448-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12031-010-9448-4