Abstract
Background:
We investigated the aetiologic role of human papillomavirus (HPV) in 120 penile squamous cell carcinomas (PSCCs) from Vietnam.
Methods:
Human papillomavirus DNA was detected by PCR using SPF10 primers and a primer set targeting HPV-16 E6. The INNO-LiPA HPV genotyping kit was used to determine genotype. Human papillomavirus-16 viral load and physical status were determined by real-time PCR. P16INK4A protein expression was investigated by immunohistochemistry.
Results:
Human papillomavirus DNA was detected in 27 of 120 (23%) PSCCs. The most frequently detected genotype was HPV-16 (24 of 27 cases, 89%). In 16 of 18 (89%) HPV-16-positive cases, the HPV DNA was considered to be integrated into the host genome. The geometric mean of the HPV-16 viral load was 0.4 copies per cell. P16INK4A overexpression was significantly related to PSCCs infected with high-risk HPV (P=0.018) and HPV-16 copy numbers (P<0.001).
Conclusion:
Human papillomavirus-16 DNA integration and p16INK4A overexpression in high-risk HPV detected PSCCs suggested an aetiologic role of high-risk HPV in the development of PSCCs.
Similar content being viewed by others
Main
Penile cancer development is a multi-factorial process involving poor genital hygiene, phimosis, human papillomavirus (HPV), chronic inflammatory and premalignant conditions, and smoking (Misra et al, 2004). According to a systematic review, human papillomavirus (HPV) DNA was detected in 48%, on average, of penile squamous cell carcinomas (PSCCs) and the most frequent HPV-related histotype (66%) was basaloid PSCC (Backes et al, 2009).
The integration of high-risk HPV genome into the host genome is suspected to be an important event for malignant transformation and cancer progression (Wentzensen et al, 2004; Williams et al, 2011). It usually disrupts the E2 gene, a suppressor of the E6/E7 promoter, leading to the overexpression of viral oncogenes E6 and E7. The high viral load, frequently observed in cervical cancer, is also a determinant of cancer development (Wu et al, 2006).
The overexpression of p16INK4A is a useful biomarker for evaluating the aetiologic role of HPV because HPV-E7 disturbs the p16INK4A/cyclin D/Rb pathway, leading to the accumulation of p16INK4A (Narisawa-Saito and Kiyono, 2007). In PSCC, however, the association between p16INK4A overexpression and HPV presence is still unestablished (Ferreux et al, 2003; Poetsch et al, 2011; Stankiewicz et al, 2011a).
To understand the aetiologic role of HPV in the development of PSCCs, we examined the presence, genotype, viral load and physical status of high-risk HPV, and p16INK4A expression in PSCCs in Vietnam.
Materials and Methods
Study subjects
One hundred twenty paraffin-embedded PSCC specimens diagnosed at the National Cancer Hospital (Hanoi, Vietnam) between 2005 and 2010 were examined. Seventeen cervical cancer specimens obtained at the same hospital during the same period were used as positive controls as HPV is a necessary cause of cervical cancers (zur Hausen, 2002). Histological subtypes were confirmed according to the World Health Organization histological classification of PSCCs (Cubilla et al, 2004). On the basis of TNM classification (Pizzocaro et al, 2010), clinical stage was divided into four stages by one of the authors (HD). This study was approved by the Institutional Review Board of Kagoshima University Graduate School of Medical and Dental Sciences.
Human papillomavirus detection and genotyping
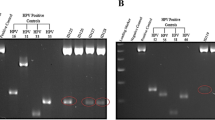
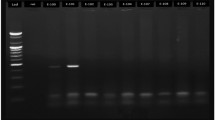
Extracted DNAs from tissue specimens were checked for their qualities and the absence of PCR inhibitors by PCR for β-globin (Khan et al, 2008). Human papillomavirus DNA was detected by PCR using SPF10-biotinylated primers and a HPV-16-E6-specific primer set (Khan et al, 2008). Human papillomavirus typing was performed using the INNO-LiPA HPV Genotyping Extra test (Innogenetics, Ghent, Belgium) (Kleter et al, 1999), which can identify 28 genotypes: HPV-16, -18, -31, -33, -35, -39, -45, -51, -52, -56, -58, -59, -68, -73, -82, -26, -53, -66, -6, -11, -40, -43, -44, -54, -70, -69, -71 and -74.
Quantitative real-time PCR
To examine the viral load and the physical status of HPV-16, all HPV-16-positive samples were subjected to quantitative real-time PCR with the ABI Prism 7700 Sequence Detection System (Applied Biosystems, Foster City, CA, USA) and 2x QuantiTect SYBR Green PCR kit (Qiagen, Hilden, Germany). Details of the procedure were reported in the previous study (Khan et al, 2008). The physical status of the HPV-16 was determined by the HPV-16 E2/E6 ratio (Peitsaro et al, 2002). A lack of E2 amplification (E2/E6 ratio=0) represents HPV-16 DNA integration into the host genome. When the E2/E6 ratio was equal to or higher than unity (E2/E6 ratio ⩾1), the HPV-16 genome was considered as an episomal form, and the rest (0
Immunohistochemistry for p16INK4A
The immunohistochemistry was conducted, using the mouse monoclonal antibody against p16INK4A (1 : 150 dilutions, 551153, BD Pharmingen, Tokyo, Japan). Details of the procedure were reported in the previous study (Baba et al, 2010). The p16INK4A expression was classified into the following four groups: <10%, 10–49%, 50–89% and ⩾90%. The cases with ⩾10% carcinoma cells stained positively were classified as positive.
Results
The β-globin was detected in all samples, indicating that DNA was available for molecular analysis. Twenty-seven of 120 (23%) PSCCs, including two of three (67%) basaloid PSCCs, were HPV positive (Table 1). Twenty-three HPV-positive cases were detected by SPF10 primers, and additional four cases by a HPV-16-specific primer set (PC-1, PC-4, PC-7 and PC-8 in Supplementary Table 1). The HPV prevalence did not differ by any clinicopathological parameters. In cervical carcinomas, HPV DNA was detected in 94% (16 out of 17) cases.
Human papillomavirus-16 was detected in 24 of 27 PSCCs. Other HPV genotypes detected included HPV-18, -11, -33 and -58 in one case each. Two cases had multiple infections: HPV-16/-58 and HPV-16/-18 (Supplementary Table 1).
Human papillomavirus-16 E6 DNA was quantified in 18 of 24 HPV-16-positive PSCCs (Supplementary Table 1). The geometric means of E6 copies per cell were 0.4 in PSCCs and 3.0 in cervical cancers. Human papillomavirus DNA was in the integrated form in seven (39%), the mixed form in nine (50%) and the episomal form in two (11%) cases. In 13 HPV-16-positive cervical cancers, HPV integrated, mixed and episomal forms were found in four (31%), three (23%) and six (46%) cases, respectively.
Eighteen of 25 high-risk HPV (16 HPV-16, one HPV-16/-58 and one HPV-33)-positive cases and 26 randomly selected HPV-negative PSCCs were subjected to immunohistochemistry. Seven high-risk HPV-positive cases (PC-4, PC-5, PC-11, PC-14, PC-16, PC-17 and PC-22) were not examined due to tissue shortage.
Although p16INK4A had both nuclear and cytoplasmic immunoreactivity (Figure 1A and D), samples showing cytoplasmic immunoreactivity alone (Figure 1B and E) were not regarded as positive because the functionally activated p16INK4A was translocated into the nucleus. P16INK4A was expressed in 10 of 44 (23%) PSCCs (Supplementary Table 2). Human papillomavirus-positive PSCCs tended to show a frequent p16INK4A nuclear expression (Supplementary Table 2, P=0.018). Strong p16INK4A nuclear expression was more frequently observed in HPV-16-positive PSCCs with a viral load ⩾1 than those with a viral load <1 and HPV-negative PSCCs (Table 2, P for trend <0.001). Although all basaloid and non-keratinising types had high viral loads, 83% of the keratinising type harboured low viral loads (Table 2, P=0.032). P16INK4A expression was not related to other factors, including tumour grade or stage (Supplementary Table 2).
Representative examples of p16INK4A immunostaining in PSCCs. (A) HPV positive, p16INK4A both nuclear and cytoplasmic positive; (B) HPV positive, p16INK4A cytoplasmic positive only; (C) HPV positive, p16 INK4A both nuclear and cytoplasmic negative; (D) HPV negative, p16 INK4A both nuclear and cytoplasmic positive; (E) HPV negative, p16INK4A cytoplasmic positive only; (F) HPV negative, p16INK4A both nuclear and cytoplasmic negative.
Discussion
In the present study, p16INK4A overexpression was frequently observed in high-risk HPV-positive PSCCs, which is consistent with the findings in cervical cancers (Hwang and Shroyer 2012) and some studies of PSCCs (Ferreux et al, 2003; Stankiewicz et al, 2011a). Regarding HPV-negative PSCCs, p16INK4A expression was frequently suppressed by p16INK4A gene mutation or promoter hypermethylation (Poetsch et al, 2011). However, three HPV-negative PSCCs also showed p16INK4A overexpression (Table 2), which might occur independently of HPV infection such as mutational inactivation of pRB (Marur et al, 2010). A high viral load of HPV-16 was also related to strong p16INK4A nuclear expression and almost none of the cases with viral load <1 copy per cell showed p16INK4A overexpression. To our knowledge, this is the first study reporting this association. P16INK4A overexpression may predict HPV transcription activity as reported in tonsillar cancer (Hoffmann et al, 2010).
The median HPV-16 viral loads (ranges) were 60.1255 (10.7–1239) and 0.0355 (0.002–0.322) copies per cell in the high (⩾1 per cell) and low (<1 per cell) viral load groups, respectively. These ranges were similar to those in HPV-16 E6 mRNA-positive and -negative PSCCs, respectively (Heideman et al, 2007). Thus, approximately one copy per cell is a reasonable threshold to distinguish the viral transcription activity.
Relatively low viral loads in PSCCs might be due to the frequent HPV-16 integration because the viral load decreases after viral integration into the host genome (Berumen et al, 1995). The high frequency (89%) of HPV-16 integration in Vietnamese PSCCs (both integrated and mixed forms) is consistent with the findings by Tornesello et al (1997) and Kalantari et al (2008). Human papillomavirus integration, a marker of HPV-induced neoplasia, is not always found in cervical cancers (Cullen et al, 1991; Vernon et al, 1997). The discrepancy in HPV-16 integration rate between PSCCs and cervical cancers in the current study could be explained by the difference in tumour histological grades or aggressiveness, as HPV integration frequently occurs in a severe dysplastic lesion and invasive cervical carcinomas (Wentzensen et al, 2004). However, our sample size was too small to explore this hypothesis: only two PSCCs were in episomal form.
Demographic features and genetic backgrounds may contribute to the geographical difference of HPV prevalence in PSCCs worldwide. To date, there is no study reporting the HPV presence in PSCC in Vietnam. Although the HPV prevalence in Vietnamese PSCCs was relatively lower than the world average, a high frequency (94%) of HPV in cervical cancers indicated the appropriateness of our HPV DNA detection procedure.
Human papillomavirus presence was not significantly related to other PSCC risk factors including phimosis and smoking. Although clinical information was not obtained from nearly half of the study subjects, the HPV prevalence in these cases (20–25%) differed little from that of the entire group (23%). Thus, our negative finding was unlikely caused by biased information.
Conclusion
The aetiologic role of high-risk HPV in the development of PSCCs was suggested by its DNA integration into the PSCC genome and its association with p16INK4A overexpression. P16INK4A could be a biomarker for HPV-related PSCCs.
Change history
15 January 2013
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Baba M, Castillo A, Koriyama C, Yanagi M, Matsumoto H, Natsugoe S, Shuyama KY, Khan N, Higashi M, Itoh T, Eizuru Y, Aikou T, Akiba S (2010) Human papillomavirus is frequently detected in gefitinib-responsive lung adenocarcinomas. Oncol Rep 23 (4): 1085–1092
Backes DM, Kurman RJ, Pimenta JM, Smith JS (2009) Systematic review of human papillomavirus prevalence in invasive penile cancer. Cancer Causes Control 20: 449–457
Berumen J, Unger ER, Casas L, Figueroa P (1995) Amplification of human papillomavirus types 16 and 18 in invasive cervical cancer. Hum Pathol 26: 676–681
Cubilla AL, Dillner J, Schellhammer PF, Horenblas S (2004) Tumours of the penis: malignant epithelial tumours. In Eble JN, Sauter G, Epstein JI, Sesterhenn I, (eds) Chap 5. World Health Organization Classification of Tumours Pathology & Genetics of Tumours of the Urinary System and Male Genital Organs pp 281–290. IARC: Lyon
Cullen AP, Reid R, Campion M, Lörincz AT (1991) Analysis of the physical state of different human papillomavirus DNAs in intraepithelial and invasive cervical neoplasm. J Virol 65: 606–612
Ferreux E, Lont AP, Horenblas S, Gallee MP, Raaphorst FM, von Knebel Doeberitz M, Meijer CJ, Snijders PJ (2003) Evidence for at least three alternative mechanisms targeting the p16INK4A/cyclinD/Rb pathway in penile carcinoma, one of which is mediated by high-risk human papillomavirus. J Pathol 201: 109–118
Heideman DA, Waterboer T, Pawlita M, Delis-van Diemen P, Nindl I, Leijte JA, Bonfrer JM, Horenblas S, Meijer CJ, Snijders PJ (2007) Human papillomavirus-16 is the predominant type etiologically involved in penile squamous cell carcinoma. J Clin Oncol 25: 4550–4556
Hoffmann M, Ihloff AS, Görögh T, Weise JB, Fazel A, Krams M, Rittgen W, Schwarz E, Kahn T (2010) P16(INK4a) overexpression predicts translational active human papillomavirus infection in tonsillar cancer. Int J Cancer 127: 1595–1602
Hwang SJ, Shroyer KR (2012) Biomarkers of cervical dysplasia and carcinoma. J Oncol 2012 doi:10.1155/2012/507286
Kalantari M, Villa LL, Calleja-Macias IE, Bernard HU (2008) Human papillomavirus-16 and -18 in penile carcinomas: DNA methylation, chromosomal recombination and genomic variation. Int J Cancer 123: 1832–1840
Khan NA, Castillo A, Koriyama C, Kijima Y, Umekita Y, Ohi Y, Higashi M, Sagara Y, Yoshinaka H, Tsuji T, Natsugoe S, Douchi T, Eizuru Y, Akiba S (2008) Human papillomavirus detected in female breast carcinomas in Japan. Br J Cancer 99: 408–414
Kleter B, Van Doorn LJ, Schrauwen L, Molijin A, Sastrowijoto S, ter Schegget J, Lindeman J, terHarmsel B, Burger M, Quint W (1999) Development and clinical evaluation of a highly sensitive PCR-reverse hybridization line probe assay for detection and identification of anogenital human papillomavirus. J Clin Microbiol 37: 2508–2517
Marur S, D'Souza G, Westra WH, Forastiere AA (2010) HPV-associated head and neck cancer: a virus-related cancer epidemic. Lancet Oncol 11 (8): 781–789
Misra S, Chaturvedi A, Misra NC (2004) Penile carcinoma: a challenge for the developing world. Lancet Oncol 5: 240–247
Narisawa-Saito M, Kiyono T (2007) Basic mechanisms of high-risk human papillomavirus-induced carcinogenesis: roles of E6 and E7 proteins. Cancer Sci 98: 1505–1511
Peitsaro P, Johansson B, Syrjanen S (2002) Integrated human papillomavirus type 16 is frequently found in cervical cancer precursors as demonstrated by a novel quantitative real-time PCR technique. J Clin Microbiol 40: 886–891
Pizzocaro G, Algaba F, Horenblas S, Solsona E, Tana S, Van Der Poel H, Watkin NA (2010) EAU Penile cancer guidelines 2009. Eur Urol 57: 1002–1012
Poetsch M, Hemmerich M, Kakies C, Kleist B, Wolf E, vom Dorp F, Hakenberg OW, Protzel C (2011) Alterations in the tumor suppressor gene p16(INK4A) are associated with aggressive behavior of penile carcinomas. Virchows Arch 458 (2): 221–229
Stankiewicz E, Prowse DM, Ktori E, Cuzick J, Ambroisine L, Zhang X, Kudahetti S, Watkin N, Corbishley C, Berney DM (2011a) The retinoblastoma protein/p16 INK4A pathway but not p53 is disrupted by human papillomavirus in penile squamous cell carcinoma. Histopathology 58: 433–439
Tornesello ML, Buonaguro FM, Meglio A, Buonaguro L, Beth-Giraldo E, Giraldo G (1997) Sequence variations and viral genomic state of human papillomavirus type 16 in penile carcinomas from Ugandan patients. J Gen Virol 78: 2199–2208
Vernon SD, Unger ER, Miller DL, Lee DR, Reeves WC (1997) Association of human papillomavirus type 16 integration in the E2 gene with poor disease-free survival from cervical cancer. Int J Cancer 74: 50–56
Wentzensen N, Vinokurova S, von Knebel Doeberitz M (2004) Systematic review of genomic integration sites of human papillomavirus genomes in epithelial dysplasia and invasive cancer of the female lower genital tract. Cancer Res 64 (11): 3878–3884
Williams VM, Filippova M, Soto U, Duerksen-Hughes PJ (2011) HPV-DNA integration and carcinogenesis: putative roles for inflammation and oxidative stress. Future Virol 6: 45–57
Wu Y, Chen Y, Li L, Yu G, Zhang Y, He Y (2006) Associations of high-risk HPV types and viral load with cervical cancer in China. J Clin Virol 35: 264–269
zur Hausen H (2002) Papillomaviruses and cancer: from basic studies to clinical application. Nat Rev Cancer 2: 342–350
Acknowledgements
This work was supported by Grants-in-Aids for Scientific Research on Priority Areas (17015037) of the Ministry of Education, Culture, Sports, Science and Technology, Japan. We would like to express our gratefulness to Dr Ta Van To, Department of Cytology and Pathology, National Cancer Hospital, 43 Quan Su, Hanoi, Vietnam, for kindly giving paraffin block specimens. We thank Joint Research Laboratory, Kagoshima University Graduate School of Medical and Dental Sciences, for the use of their facilities. We also thank Ms Junko Habu for her technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Do, H., Koriyama, C., Khan, N. et al. The etiologic role of human papillomavirus in penile cancers: a study in Vietnam. Br J Cancer 108, 229–233 (2013). https://doi.org/10.1038/bjc.2012.583
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2012.583
Keywords
This article is cited by
-
Low concordance of oral and genital HPV infection among male patients with sexually transmitted infections in Vietnam
BMC Infectious Diseases (2019)
-
Nuclear loss and cytoplasmic expression of androgen receptor in penile carcinomas: role as a driver event and as a prognosis factor
Virchows Archiv (2018)
-
Prevalence of human papillomavirus in penile malignant tumors: viral genotyping and clinical aspects
BMC Urology (2015)