Abstract
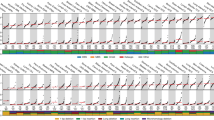
Thousands of somatic mutations accrue in most human cancers, and their causes are largely unknown. We recently showed that the DNA cytidine deaminase APOBEC3B accounts for up to half of the mutational load in breast carcinomas expressing this enzyme. Here we address whether APOBEC3B is broadly responsible for mutagenesis in multiple tumor types. We analyzed gene expression data and mutation patterns, distributions and loads for 19 different cancer types, with over 4,800 exomes and 1,000,000 somatic mutations. Notably, APOBEC3B is upregulated, and its preferred target sequence is frequently mutated and clustered in at least six distinct cancers: bladder, cervix, lung (adenocarcinoma and squamous cell carcinoma), head and neck, and breast. Interpreting these findings in the light of previous genetic, cellular and biochemical studies, the most parsimonious conclusion from these global analyses is that APOBEC3B-catalyzed genomic uracil lesions are responsible for a large proportion of both dispersed and clustered mutations in multiple distinct cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Stephens, P. et al. A screen of the complete protein kinase gene family identifies diverse patterns of somatic mutations in human breast cancer. Nat. Genet. 37, 590–592 (2005).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Jones, S. et al. Frequent mutations of chromatin remodeling gene ARID1A in ovarian clear cell carcinoma. Science 330, 228–231 (2010).
Sjöblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Kumar, A. et al. Exome sequencing identifies a spectrum of mutation frequencies in advanced and lethal prostate cancers. Proc. Natl. Acad. Sci. USA 108, 17087–17092 (2011).
Parsons, D.W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011).
Berger, M.F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
Stransky, N. et al. The mutational landscape of head and neck squamous cell carcinoma. Science 333, 1157–1160 (2011).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Stephens, P.J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Makridakis, N.M. & Reichardt, J.K. Translesion DNA polymerases and cancer. Front. Genet. 3, 174 (2012).
Roberts, S.A. et al. Clustered mutations in yeast and in human cancers can arise from damaged long single-strand DNA regions. Mol. Cell 46, 424–435 (2012).
Drier, Y. et al. Somatic rearrangements across cancer reveal classes of samples with distinct patterns of DNA breakage and rearrangement-induced hypermutability. Genome Res. 23, 228–235 (2013).
Harris, R.S., Petersen-Mahrt, S.K. & Neuberger, M.S. RNA editing enzyme APOBEC1 and some of its homologs can act as DNA mutators. Mol. Cell 10, 1247–1253 (2002).
Di Noia, J.M. & Neuberger, M.S. Molecular mechanisms of antibody somatic hypermutation. Annu. Rev. Biochem. 76, 1–22 (2007).
Longerich, S., Basu, U., Alt, F. & Storb, U. AID in somatic hypermutation and class switch recombination. Curr. Opin. Immunol. 18, 164–174 (2006).
Conticello, S.G. The AID/APOBEC family of nucleic acid mutators. Genome Biol. 9, 229 (2008).
LaRue, R.S. et al. Guidelines for naming nonprimate APOBEC3 genes and proteins. J. Virol. 83, 494–497 (2009).
Malim, M.H. APOBEC proteins and intrinsic resistance to HIV-1 infection. Phil. Trans. R. Soc. Lond. B 364, 675–687 (2009).
Harris, R.S., Hultquist, J.F. & Evans, D.T. The restriction factors of human immunodeficiency virus. J. Biol. Chem. 287, 40875–40883 (2012).
Blanc, V. & Davidson, N.O. C-to-U RNA editing: mechanisms leading to genetic diversity. J. Biol. Chem. 278, 1395–1398 (2003).
Bishop, K.N., Holmes, R.K., Sheehy, A.M. & Malim, M.H. APOBEC-mediated editing of viral RNA. Science 305, 645 (2004).
Petit, V. et al. Murine APOBEC1 is a powerful mutator of retroviral and cellular RNA in vitro and in vivo. J. Mol. Biol. 385, 65–78 (2009).
Ikeda, T. et al. Intrinsic restriction activity by apolipoprotein B mRNA editing enzyme APOBEC1 against the mobility of autonomous retrotransposons. Nucleic Acids Res. 39, 5538–5554 (2011).
Petersen-Mahrt, S.K., Harris, R.S. & Neuberger, M.S. AID mutates E. coli suggesting a DNA deamination mechanism for antibody diversification. Nature 418, 99–103 (2002).
Petersen-Mahrt, S.K. & Neuberger, M.S. In vitro deamination of cytosine to uracil in single-stranded DNA by apolipoprotein B editing complex catalytic subunit 1 (APOBEC1). J. Biol. Chem. 278, 19583–19586 (2003).
Pham, P., Bransteitter, R., Petruska, J. & Goodman, M.F. Processive AID-catalysed cytosine deamination on single-stranded DNA simulates somatic hypermutation. Nature 424, 103–107 (2003).
Chelico, L., Pham, P., Calabrese, P. & Goodman, M.F. APOBEC3G DNA deaminase acts processively 3′ → 5′ on single-stranded DNA. Nat. Struct. Mol. Biol. 13, 392–399 (2006).
Hultquist, J.F. et al. Human and rhesus APOBEC3D, APOBEC3F, APOBEC3G, and APOBEC3H demonstrate a conserved capacity to restrict Vif-deficient HIV-1. J. Virol. 85, 11220–11234 (2011).
Robbiani, D.F. & Nussenzweig, M.C. Chromosome translocation, B cell lymphoma, and activation-induced cytidine deaminase. Annu. Rev. Pathol. 8, 79–103 (2013).
Okazaki, I.M. et al. Constitutive expression of AID leads to tumorigenesis. J. Exp. Med. 197, 1173–1181 (2003).
Yamanaka, S. et al. Apolipoprotein B mRNA-editing protein induces hepatocellular carcinoma and dysplasia in transgenic animals. Proc. Natl. Acad. Sci. USA 92, 8483–8487 (1995).
Burns, M.B. et al. APOBEC3B is an enzymatic source of mutation in breast cancer. Nature 494, 366–370 (2013).
Jarmuz, A. et al. An anthropoid-specific locus of orphan C to U RNA-editing enzymes on chromosome 22. Genomics 79, 285–296 (2002).
Refsland, E.W. et al. Quantitative profiling of the full APOBEC3 mRNA repertoire in lymphocytes and tissues: implications for HIV-1 restriction. Nucleic Acids Res. 38, 4274–4284 (2010).
Koning, F.A. et al. Defining APOBEC3 expression patterns in human tissues and hematopoietic cell subsets. J. Virol. 83, 9474–9485 (2009).
Lackey, L. et al. APOBEC3B and AID have similar nuclear import mechanisms. J. Mol. Biol. 419, 301–314 (2012).
Kohli, R.M. et al. Local sequence targeting in the AID/APOBEC family differentially impacts retroviral restriction and antibody diversification. J. Biol. Chem. 285, 40956–40964 (2010).
Wang, M., Rada, C. & Neuberger, M.S. Altering the spectrum of immunoglobulin V gene somatic hypermutation by modifying the active site of AID. J. Exp. Med. 207, 141–153 (2010).
Albin, J.S. & Harris, R.S. Interactions of host APOBEC3 restriction factors with HIV-1 in vivo: implications for therapeutics. Expert Rev. Mol. Med. 12, e4 (2010).
Palles, C. et al. Germline mutations affecting the proofreading domains of POLE and POLD1 predispose to colorectal adenomas and carcinomas. Nat. Genet. 45, 136–144 (2013).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Berger, M.F. et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 485, 502–506 (2012).
Fujino, T., Navaratnam, N. & Scott, J. Human apolipoprotein B RNA editing deaminase gene (APOBEC1). Genomics 47, 266–275 (1998).
Muramatsu, M. et al. Specific expression of activation-induced cytidine deaminase (AID), a novel member of the RNA-editing deaminase family in germinal center B cells. J. Biol. Chem. 274, 18470–18476 (1999).
Stenglein, M.D., Burns, M.B., Li, M., Lengyel, J. & Harris, R.S. APOBEC3 proteins mediate the clearance of foreign DNA from human cells. Nat. Struct. Mol. Biol. 17, 222–229 (2010).
Sato, Y. et al. Deficiency in APOBEC2 leads to a shift in muscle fiber type, diminished body mass, and myopathy. J. Biol. Chem. 285, 7111–7118 (2010).
Rogozin, I.B., Basu, M.K., Jordan, I.K., Pavlov, Y.I. & Koonin, E.V. APOBEC4, a new member of the AID/APOBEC family of polynucleotide (deoxy)cytidine deaminases predicted by computational analysis. Cell Cycle 4, 1281–1285 (2005).
Rada, C., Jarvis, J.M. & Milstein, C. AID-GFP chimeric protein increases hypermutation of Ig genes with no evidence of nuclear localization. Proc. Natl. Acad. Sci. USA 99, 7003–7008 (2002).
Land, A.M. et al. Endogenous APOBEC3A DNA cytosine deaminase is cytoplasmic and non-genotoxic. J. Biol. Chem. 288, 17253–17260 (2013).
Acknowledgements
We thank The Cancer Genome Atlas (TCGA) Network for generating the RNA-seq and somatic mutation data and for providing open access, and we thank the Harris laboratory members and S. Kaufmann for comments. M.B.B. was supported by a Department of Defense Breast Cancer Research Program Predoctoral Fellowship (BC101124). This work was supported by grants from the Jimmy V Foundation, the Minnesota Ovarian Cancer Alliance and the US National Institutes of Health (R01 AI064046 and P01 GM091743).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study design, data analysis and manuscript preparation. M.B.B. and N.A.T. analyzed data from TCGA. N.A.T. performed mutation and cluster analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–3 (PDF 1276 kb)
Rights and permissions
About this article
Cite this article
Burns, M., Temiz, N. & Harris, R. Evidence for APOBEC3B mutagenesis in multiple human cancers. Nat Genet 45, 977–983 (2013). https://doi.org/10.1038/ng.2701
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2701
This article is cited by
-
Mesoscale DNA features impact APOBEC3A and APOBEC3B deaminase activity and shape tumor mutational landscapes
Nature Communications (2024)
-
Distinguishing preferences of human APOBEC3A and APOBEC3B for cytosines in hairpin loops, and reflection of these preferences in APOBEC-signature cancer genome mutations
Nature Communications (2024)
-
APOBEC3-mediated mutagenesis in cancer: causes, clinical significance and therapeutic potential
Journal of Hematology & Oncology (2023)
-
Acute expression of human APOBEC3B in mice results in RNA editing and lethality
Genome Biology (2023)
-
Proteogenomics decodes the evolution of human ipsilateral breast cancer
Communications Biology (2023)