Abstract
Genome-wide DNA sequencing was used to decrypt the phylogeny of multiple samples from distinct areas of cancer and morphologically normal tissue taken from the prostates of three men. Mutations were present at high levels in morphologically normal tissue distant from the cancer, reflecting clonal expansions, and the underlying mutational processes at work in morphologically normal tissue were also at work in cancer. Our observations demonstrate the existence of ongoing abnormal mutational processes, consistent with field effects, underlying carcinogenesis. This mechanism gives rise to extensive branching evolution and cancer clone mixing, as exemplified by the coexistence of multiple cancer lineages harboring distinct ERG fusions within a single cancer nodule. Subsets of mutations were shared either by morphologically normal and malignant tissues or between different ERG lineages, indicating earlier or separate clonal cell expansions. Our observations inform on the origin of multifocal disease and have implications for prostate cancer therapy in individual cases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
05 May 2015
In the version of this article initially published, author Manasa Ramakrishna was omitted from the author list. The error has been corrected in the PDF and HTML versions of this article.
References
Andreoiu, M. & Cheng, L. Multifocal prostate cancer: biologic, prognostic, and therapeutic implications. Hum. Pathol. 41, 781–793 (2010).
Cheng, L. et al. Evidence of independent origin of multiple tumors from patients with prostate cancer. J. Natl. Cancer Inst. 90, 233–237 (1998).
Kobayashi, M. et al. Molecular analysis of multifocal prostate cancer by comparative genomic hybridization. Prostate 68, 1715–1724 (2008).
Boyd, L.K. et al. High-resolution genome-wide copy-number analysis suggests a monoclonal origin of multifocal prostate cancer. Genes Chromosom. Cancer 51, 579–589 (2012).
Lindberg, J. et al. Exome sequencing of prostate cancer supports the hypothesis of independent tumour origins. Eur. Urol. 63, 347–353 (2013).
Clark, J. et al. Complex patterns of ETS gene alteration arise during cancer development in the human prostate. Oncogene 27, 1993–2003 (2008).
Attard, G. et al. Duplication of the fusion of TMPRSS2 to ERG sequences identifies fatal human prostate cancer. Oncogene 27, 253–263 (2008).
Gaisa, N.T. et al. Clonal architecture of human prostatic epithelium in benign and malignant conditions. J. Pathol. 225, 172–180 (2011).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Svensson, M.A. et al. Testing mutual exclusivity of ETS rearranged prostate cancer. Lab. Invest. 91, 404–412 (2011).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Alexandrov, L.B., Nik-Zainal, S., Wedge, D.C., Campbell, P.J. & Stratton, M.R. Deciphering signatures of mutational processes operative in human cancer. Cell Rep. 3, 246–259 (2013).
Nik-Zainal, S. et al. Mutational processes molding the genomes of 21 breast cancers. Cell 149, 979–993 (2012).
Baca, S.C. et al. Punctuated evolution of prostate cancer genomes. Cell 153, 666–677 (2013).
Berger, M.F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
Grasso, C.S. et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature 487, 239–243 (2012).
Barbieri, C.E. et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat. Genet. 44, 685–689 (2012).
Weischenfeldt, J. et al. Integrative genomic analyses reveal an androgen-driven somatic alteration landscape in early-onset prostate cancer. Cancer Cell 23, 159–170 (2013).
Holstege, H. et al. Somatic mutations found in the healthy blood compartment of a 115-yr-old woman demonstrate oligoclonal hematopoiesis. Genome Res. 24, 733–742 (2014).
Mucci, N.R. et al. Expression of nuclear antigen Ki-67 in prostate cancer needle biopsy and radical prostatectomy specimens. J. Natl. Cancer Inst. 92, 1941–1942 (2000).
Slaughter, D.P., Southwick, H.W. & Smejkal, W. Field cancerization in oral stratified squamous epithelium; clinical implications of multicentric origin. Cancer 6, 963–968 (1953).
Leemans, C.R., Braakhuis, B.J.M. & Brakenhoff, R.H. The molecular biology of head and neck cancer. Nat. Rev. Cancer 11, 9–22 (2011).
Dworkin, A.M., Huang, T.H.-M. & Toland, A.E. Epigenetic alterations in the breast: implications for breast cancer detection, prognosis and treatment. Semin. Cancer Biol. 19, 165–171 (2009).
Karavitakis, M., Ahmed, H.U., Abel, P.D., Hazell, S. & Winkler, M.H. Tumor focality in prostate cancer: implications for focal therapy. Nat. Rev. Clin. Oncol. 8, 48–55 (2011).
Tareen, B., Godoy, G. & Taneja, S.S. Focal therapy: a new paradigm for the treatment of prostate cancer. Rev. Urol. 11, 203–212 (2009).
Reva, B., Antipin, Y. & Sander, C. Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Res. 39, e118 (2011).
Chang, X. & Wang, K. wANNOVAR: annotating genetic variants for personal genomes via the web. J. Med. Genet. 49, 433–436 (2012).
Lawrence, M.S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Campbell, P.J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat. Genet. 40, 722–729 (2008).
Warren, A.Y. et al. Method for sampling tissue for research which preserves pathological data in radical prostatectomy. Prostate 73, 194–202 (2013).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 25, 2865–2871 (2009).
Campbell, P.J. et al. Identification of somatically acquired rearrangements in cancer using genome-wide massively parallel paired-end sequencing. Nat. Genet. 40, 722–729 (2008).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl. Acad. Sci. USA 107, 16910–16915 (2010).
Nilsen, G. et al. Copynumber: efficient algorithms for single- and multi-track copy number segmentation. BMC Genomics 13, 591 (2012).
Tsai, M.-F. et al. PrimerZ: streamlined primer design for promoters, exons and human SNPs. Nucleic Acids Res. 35, W63–W65 (2007).
Stephens, P.J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Acknowledgements
This work was funded by Cancer Research UK (grant C5047/A14835), the Dallaglio Foundation and the Wellcome Trust. We also acknowledge support from the Bob Champion Cancer Trust, the Orchid Cancer Appeal, the RoseTrees Trust, the North West Cancer Research Fund, Big C, the King family, the Grand Charity of Freemasons, and the Research Foundation Flanders (FWO). We thank D. Holland from the Infrastructure Management Team and P. Clapham from the Informatics Systems Group at the Wellcome Trust Sanger Institute. We acknowledge the Biomedical Research Centre at the Institute of Cancer Research and the Royal Marsden NHS Foundation Trust, supported by the National Institute for Health Research. We acknowledge support from the National Cancer Research Prostate Cancer: Mechanisms of Progression and Treatment (PROMPT) collaborative (grant G0500966/75466). We thank the National Institute for Health Research, Hutchison Whampoa Limited and the Human Research Tissue Bank (Addenbrooke's Hospital), the Cancer Research UK Cambridge Research Institute Histopathology, the In-situ Hybridisation Core Facility, the Genomics Core Facility Cambridge and the Cambridge University Hospitals Media Studio.
Author information
Authors and Affiliations
Consortia
Contributions
C.S.C., R.E. and D.E.N. are senior principal investigators who designed and coordinated the study. C.S.F. is a senior principal investigator and histopathology lead. D.S.B. and U.M. are senior principal investigators for this project and bioinformatics project coordinators. D.E., A.F. and M.R.S. are senior principal investigators for this project. D.C.W. and P.V.L. had overall responsibility for data analysis. A.Y.W. is a histopathology lead. G.G. performed chromoplexy analysis. L.B.A. analyzed mutational signatures. H.C.W. was a principal investigator for this particular project who also carried out data analysis and tissue collection. A.B. and S.O'M. are coordinators of the DNA mutation–analysis pipeline. C.E.M. was involved in data analysis and formulation of the manuscript structure. P.C., B.K., J.Z., S.N.-Z. and A.G.L. were involved in data analysis and interpretation. N.D., S.E., L. Matthews and S. Merson completed tissue collection and FISH analysis of DNA preparations. N.C., C.G., M.R. and Z.K.-T. carried out data analysis. D.L. performed data validation. J.K. and H.J.L. collected tissue and performed DNA extractions. S.T. obtained patient consent, collected blood and carried out blood DNA preparations. J.C. and R.H. performed FISH analysis. R.M. and T.V. were involved in data interpretation. R.G.B., P.C.B. and M.F. were involved in determining the overall study design. S.C., K.R., D.J., A.M., L.S., J.H., J.T., S. McLaren, L. Mudie, C.H., E.A., O.J., V. Goody, B.R., M.M. and S.G. ran the data mutational analysis pipeline. C.F., C.C., D.B., N.L. and S.H. completed histopathology and tissue collection. C.O., P.K., A.T., C.W., D.N., E.M., T.D., N.C.S. and V. Gnanapragasam were responsible for tissue collection.
Corresponding authors
Ethics declarations
Competing interests
R.E. has received educational grants from Illumina and GenProbe (formerly Tepnel), Vista Diagnostics and Janssen Pharmaceuticals, as well as honoraria from Succint Communications for talks on prostate cancer genetics.
Additional information
A full list of members and affiliations is provided in the Supplementary Note.
Integrated supplementary information
Supplementary Figure 1 Detection of ERG breakpoints in Case 7.
The top (a) and middle (b) panels are identical to those shown panels b and c of Fig. 1. This figure additionally shows the FISH images (c) that demonstrate the positions of breaks G, H and J (see Fig. 3 for the precise positions of breaks G and H). FISH was carried out exactly as described previously1. “Split” denotes that 5’ and 3’ ERG signals were separated but retained in the cell. “Del” indicates that 5’ ERG signals were lost from the cell, while 3’ ERG signals were retained.
Supplementary Figure 2 Three dimensional reconstructions of prostates from Patient 6, Patient 7, and Patient 8.
Showing the position of the cancer (purple shading) and the locations where samples were selected (labeled black circles) for DNA sequencing. Reconstruction was based on examination of H&E stained sections slices as indicated. Anterior prostate is at the bottom.
Supplementary Figure 3 Mutations and clonal expansions in morphologically normal tissue.
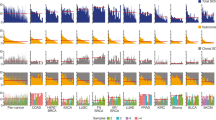
a, Summary of numbers of mutation types. b, Density plots showing the posterior distribution of the fraction of cells bearing a mutation. The fraction of cells is modeled using a previously described Bayesian Dirichlet process2. The median density is indicated by the purple line and 95% confidence intervals by the blue region. The grey histogram shows the observed frequency density of mutations as a function of the fraction of cells bearing the mutation. The y-axis is the probability density. Mutations were present in 10% of cells in Case 8 (not shown).
Supplementary Figure 4 Phylogeny based on copy number alterations alone.
Copy number alterations were detected by the Battenberg algorithm2. Each line is associated with a clone from a particular sample. The length of each line is for ease of visualisation only. The thickness of a line is proportional to its clonal representation.
Supplementary Figure 5 Genome-wide copy number profiles generated by the Battenberg algorithm.
The minor allele copy number is in blue and in purple is the total copy number.
Supplementary Figure 6 Further examples of 2D density plots showing the posterior distribution of the fraction of cells bearing a substitution in two samples.
The fraction of cells is modeled using Bayesian Dirichlet processes. From these plots it can be seen which samples share shared clonal substitutions when there is a peak at (1,1) e.g. 6_T1/6_T2; branched substitutions when there is only peaks along the axes e.g. 7_T2/7_T3 with peaks at (0,1) and (1,0); and samples that contain a sub-clone. An example of samples with a sub clone are 7_T2/7_T5 that has a peak at (0,0.72), which represents subclonal substitutions in 72% of cells in 7_T5 that have occurred only in this sample, after divergence from the other samples. Similarly, 8_T1/8_T3 has a peak at (0.54,0), representing subclonal mutations in 54% of cells in T1 only.
Supplementary Figure 7 Convergent evolution of 8p loss.
A plot showing the B-allele frequency (BAF) of segments of copy number variation detected by the battenberg algorithm on chromosome 8p. Segments showing no variation will have a BAF of 0.5. The majority of samples apart from the adjacent morphologically normal samples show a deletion at 8p. 8p deletions are at different positions and lengths in different tumors samples from the same patient showing convergent evolution. BAF values vary as a result of differing tumor content in each sample, with higher cellularity samples having more divergent BAF values in aberrant regions.
Supplementary Figure 8 Rainfall plot.
We identified localised clusters of hypermutation, a recently phenomenon termed kataegis, using a previously described algorithm. As in previously observed kataegis events, all clusters were constituted of C>T or a mixture of C>T and C>G mutations and appear to have occurred on a single strand of DNA, consistent with the operation of an APOBEC enzyme. All kataegis events were found only in one clone from each patient suggesting that it is not an initiating event. The horizontal axis illustrates the genomic coordinates of the mutations. The vertical axis plots the distance between mutations. The kataegis events in 6_T3 were both within 350bp of a rearrangement breakpoint. In 7_T2 the kataegis mutations occur on 2 chromosome copies indicating that they occurred before a whole genome duplication event, in 6_T3 the kataegis mutations are subclonal, indicating a late event, while the other two kataegis events occur clonally on one chromosome copy.
Supplementary Figure 9 Chromoplexy analysis of rearrangement breakpoints.
It was recently shown that ~40% of somatic rearrangements in prostate cancer were found in a complex series of events, called chromoplexy, that are chained together by virtue of either the proximity of their breakpoints or the existence of a 'deletion bridge' in between them5. In our multifocal prostate cancers, we identified 14 unique chromoplexy events using the ChainFinder algorithm. The oncogenic TMRSS2-ERG fusion occurred as part of a chromoplexy event in some (6_T3, 6_T4, 7_T1/T2 and 8_T1/T2) but not all tumour foci. In patients 6 and 7, we identified multiple chromoplexy events in distinct tumour cell lineages. Circos plots are shown for each sample on the corresponding branch of the phylogenetic tree. The outer rings in the circus plot provides a genomic view of the copy number changes (blue logR<-0.1, red lowR>0.1 and grey otherwise.) Rearrangement events are annotated with non-grey colours if they are found to be in chromoplexy events by the algorithm and grey otherwise.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–3 and Supplementary Note. (PDF 1028 kb)
Supplementary Data Set 1
Copy-number alterations (XLSX 14 kb)
Supplementary Data Set 2
Substitutions detected (XLSX 1241 kb)
Supplementary Data Set 3
Insertions and deletions detected (XLSX 121 kb)
Supplementary Data Set 4
Structural variants detected (XLSX 51 kb)
Rights and permissions
About this article
Cite this article
Cooper, C., Eeles, R., Wedge, D. et al. Analysis of the genetic phylogeny of multifocal prostate cancer identifies multiple independent clonal expansions in neoplastic and morphologically normal prostate tissue. Nat Genet 47, 367–372 (2015). https://doi.org/10.1038/ng.3221
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3221
This article is cited by
-
Multiregion sampling of de novo metastatic prostate cancer reveals complex polyclonality and augments clinical genotyping
Nature Cancer (2024)
-
From molecular mechanisms of prostate cancer to translational applications: based on multi-omics fusion analysis and intelligent medicine
Health Information Science and Systems (2023)
-
Interstitial pneumonia after regression by olaparib for neuroendocrine prostate cancer with BRCA1 mutation: a case report
International Cancer Conference Journal (2023)
-
Cancer origin tracing and timing in two high-risk prostate cancers using multisample whole genome analysis: prospects for personalized medicine
Genome Medicine (2023)
-
Enrichment of “Cribriform” morphologies (intraductal and cribriform adenocarcinoma) and genomic alterations in radiorecurrent prostate cancer
Modern Pathology (2022)