Abstract
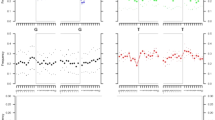
In 1991, nine sets of skeletal remains were excavated from a mass grave near Yekaterinburg, Russia which were believed to include the Russian Tsar Nicholas II, the Tsarina Alexandra, and three of their daughters1. Nuclear DNA testing of the remains verified such a family group, and mito-chondrial DNA (mtDNA) sequences of the presumed Tsarina matched a known maternal relative, Prince Philip2. mtDNA sequences from bone of the presumed Tsar matched two living maternal relatives except at a single position, where the bone sample had a mixture of matching (T) and mismatching (C) bases. Cloning experiments indicated that this mixture was due to heteroplasmy within the Tsar; nevertheless, the ‘mismatch’ fueled a lingering controversy concerning the authenticity of these remains. As a result, the official final report on the fate of the last Russian Royals has been postponed by Russian authorities pending additional, convincing DNA evidence. At the request of the Russian Federation government, we analysed the skeletal remains of the Tsar's brother Georgij Romanov, in order to gain further insight into the occurrence and segregation of heteroplasmic mtDNA variants in the Tsar's maternal lineage. The mtDNA sequence of Georgij Romanov matched that of the putative Tsar, and was heteroplasmic at the same position. This confirms heteroplasmy in the Tsar's lineage, and is powerful evidence supporting the identification of Tsar Nicholas II. The rapid intergenerational shift from heteroplasmy to homoplasmy, and the different heteroplasmic ratios in the brothers, is consistent with a ‘bottleneck’ mechanism of mtDNA segregation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Massie, R.K., Romanovs: the Final Chapter. (Random House, New York, 1995).
Gill, P. et al. Identification of the remains of the Romanov family by DNA analysis. Nature Genet. 6, 130–135 (1994).
Maples, W.R., Affidavit, Circuit Court for the city of Charlottesville, Case No. 8021, November 12, 1993.
Holland, M.M. et al. Mitochondrial DNA sequence analysis of human remains. Crime Lab. Digest 22, 3–8 (1995).
Boom, R. et al. Rapid and simple method for purification of nucleic acids. J. Clin. Microbiol. 28, 495–503 (1990).
Hoss, M. & Pääbo, S. DNA extraction from Pleistocene bones by a silica-based purification method. Nucl. Acids Res. 16, 3913–3914 (1993).
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1991).
Poulton, J. Transmission of mtDNA: cracks in the bottleneck. Am. J. Hum. Genet. 57, 224–226 (1995).
Monnat, R.J. & Loeb, L.A. Nucleotide sequence preservation of human mitochondrial DNA. Proc. Natl. Acad. Sci. USA 82, 2895–2899 (1985).
Laipis, P., Hauswirth, W., O'Brian, I. & Michaels, G. Unequal partitioning of bovine mitochondrial genotypes among siblings. Proc. Natl. Acad. Sci. USA 85, 8107–8110 (1988).
Koehler, C.M. et al. Replacement of bovine mitochondrial DNA by a sequence variant within one generation. Genetics 129, 247–255 (1991).
Bolhuis, P.A. et al. Rapid shift in genotype of human mitochondrial DNA in a family with Leber's hereditary optic neuropathy. Biochem. Biophys. Res. Comm. 170, 994–997 (1990).
Wallace, D.C. Mitochondrial DNA variation in human evolution, degenerative disease, and aging. Am. J. Hum. Genet. 57, 201–223 (1995).
Howell, N. et al. Mitochondrial gene segregation in mammals: is the bottleneck always narrow? Hum. Genet. 90, 117–120 (1992).
Bendall, K.E. & Sykes, B.C. Length heteroplasmy in the first hypervariable segment of the human mtDNA control region. Am. J. Hum. Genet. 57, 248–256 (1995).
Vilkki, J., Savonataus, M. & Singh, G. Segregation of mitochondrial genomes in a heteroplasmic lineage with Leber hereditary optic neuroretinopathy. Am. J. Hum. Genet. 47, 95–100 (1990).
Solignac, M., Monnerot, M. & Mounolou, J.C. Mitochondrial DNA heteroplasmy in Drosophila mauritiana . Proc. Natl. Acad. Sci. USA 80, 6942–6946 (1983).
Case Report of Office of Procurator General of Russian Federation, No. 18/1236 66–93, 1993.
Walsh, P.S., Metzger, D.A. & Higuchi, R. Chelex 100 as a medium for simple extraction of DNA for PCR-based typing from forensic material. BioTechniques 10, 506–513 (1991).
Stoneking, M., Sherry, S.T., Redd, A.J. & Vigilant, L. New approaches to dating suggest a recent age for the human mtDNA ancestor. Phil. Trans. R. Soc. Lond. B 337, 167–175 (1992).
Comas, D., Pääbo, S. & Bertranpetit, J. Heteroplasmy in the control region of human mitochondrial DNA. Genome Res. 5, 89–90 (1995).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ivanov, P., Wadhams, M., Roby, R. et al. Mitochondrial DNA sequence heteroplasmy in the Grand Duke of Russia Georgij Romanov establishes the authenticity of the remains of Tsar Nicholas II. Nat Genet 12, 417–420 (1996). https://doi.org/10.1038/ng0496-417
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0496-417
This article is cited by
-
Comparison of the optimal and suboptimal quantity of mitotype libraries using next-generation sequencing
International Journal of Legal Medicine (2024)
-
Measure quantity of mitochondrial DNA in aged bones or calculate it from nuclear DNA quantitative PCR results?
International Journal of Legal Medicine (2023)
-
Mitochondrial DNA Analysis in Population Isolates: Challenges and Implications for Human Identification
Current Molecular Biology Reports (2023)
-
Exploring statistical weight estimates for mitochondrial DNA matches involving heteroplasmy
International Journal of Legal Medicine (2022)
-
Genetic insights into the social organization of Neanderthals
Nature (2022)