Abstract
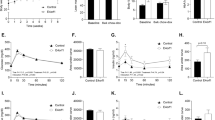
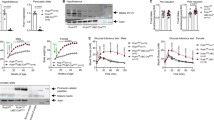
Mice homozygous for the fat mutation develop obesity and hyperglycaemia that can be suppressed by treatment with exogenous insulin. The fat mutation maps to mouse chromosome 8, very close to the gene for carboxypeptidase E (Cpe), which encodes an enzyme (CPE) that processes prohormone intermediates such as proinsulfn. We now demonstrate a defect in proinsulin processing associated with the virtual absence of CPE activity in extracts of fat/fat pancreatic islets and pituitaries. A single Ser202Pro mutation distinguishes the mutant Cpe allele, and abolishes enzymatic activity in vitro. Thus, the fat mutation represents the first demonstration of an obesity–diabetes syndrome elicited by a genetic defect in a prohormone processing pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Leiter, E. Obesity genes and diabetes induction in the mouse. Crit. Rev. FoodSci. Nutr. 33, 333–338 (1993).
Bray, G.A., Fisler, J. & York, D.A. Neuroendocrine control of the development of obesity: understanding gained from studies of experimental animal models. Front Obesity. 11, 128–181 (1990).
Michaud, E.J. et al. Differential expression of a new dominant agouti allele (Aiapy is correlated with methylation state and is influenced by parental lineage. Genes Devel. 8, 1463–1472. (1994).
Lu, D. et al. Agouti protein is an antagonist of the melanocyte-stimulating-hormone receptor. Nature 371, 799–802 (1994).
Zhang, Y. et al. Positional cloning of the mouse obese gene and its human homologue. Nature 372, 425–432 (1994).
Coleman, D.L. & Eicher, E.M. Fat (fat) and tubby (tub), two autosomal recessive mutations causing obesity syndromes in the mouse. J. Hered. 81, 424–427 (1990).
Coleman, D.L. Obese and diabetes: two mutant genes causing diabetes-obesity syndromes in mice. Diabetologia 14, 141–148 (1978).
Leiter, E. & Chapman, H. Obesity-induced diabetes (diabesity) in C57BL/ KsJ mice produces aberrant trans-regulation of sex steroid sulfotransferase genes. J. Clin. Invest. 93, 2007–2013 (1994).
Leiter, E., Chapman, H. & Falany, C. Synergism of obesity genes with hepatic steroid sulfotransferases to mediate diabetes in mice. Diabetes 40, 1360–1363 (1991).
Paigen, B.J. & Coleman, D.L. Linkage of fat to esterase-1. Mouse Genome 86, 240 (1990).
Fricker, L.D., in Peptide Biosynthesis and Processing (ed Fricker, L. D.) 199–228 (CRC Press, Boca Baton, 1991).
Dietrich, W.F. et al. A genetic map of the mouse with 4,006 simple sequence length polymorphisms. Nature Genet. 7, 220–245 (1994).
Steiner, D.F., Smeekens, S.P., Ohagi, S. & Chan, S.J. The new enzymology of precursor processing endopeptidases. J. biol. Chem. 267, 23435–23438 (1992).
Rhodes, C.J. & Alarcón, C. What β-cell defect could lead to hyperproinsulinemia in NIDDM?. Diabetes. 43, 511–517 (1994).
Davidson, H.W. & Mutton, J.C. The insulin secretory granule carboxypeptidase H. Purification and demonstration of involvement in proinsulin processing. Biochem. J. 245, 575–582 (1987).
Orci, L. et al. Direct identification of prohormone conversion site in insulin secreting cells. Cell 42, 671–681 (1985).
Manser, E. et al. Human carboxypeptidase E: isolation and characterization of the cDNA, sequence conservation, expression, and processing in vitro. Biochem. J. 267, 517–525 (1990).
Fricker, L.D. et al. Isolation and sequence analysis of a cDNA for rat carboxypeptidase E [EC 3.4.17.10], a neuropeptide processing enzyme. Molec. Endocrinol. 3, 666–673 (1989).
Fricker, L.D., Evans, C.J., Each, F.S. & Herbert, E. Cloning and sequence analysis of cDNA for bovine carboxypeptidase E. Nature 323, 461–464 (1986).
Roth, W.W., Mackin, R.B., Spiess, J., Goodman, R.E. & Noe, B.D. Primary structure and tissue distribution of anglerfish carboxypeptidase E. Molec. cell. Endocrinol. 78, 171–178 (1991).
Gidh-Jain, M. et al. Glucokinase mutation associated with non-insulin-dependent (type 2) diabetes mellitus have decreased enzymatic activity: Implications for structure/function relationships. Proc. Natl. Acad. Sci. USA 90, 1932–1936 (1993).
Hager, M. et al. A missense mutation in the glycogen receptor gene is associated with non-insulin-dependent diabetes mellitus. Nature Genet. 9, 299–304 (1995).
Leonetti, D.L., Prigeon, R.L., Boyko, E.J., Bergstrom, R.W. & Fujimoto, W.Y. Proinsulin as a marker for the development of NIDDM in Japanese-American men. Diabetes 44, 173–179 (1995).
Porte, D.J. & Kahn, S.E. Hyperproinsulinemia and amyloid in NIDDM: clues to the etiology of beta cell dysfunction. Diabetes 38, 1333–1336 (1989).
Porte, D. . β-cells in type II diabetes mellitus. Diabetes 40, 166–180 (1991).
Steiner, D.F. et al. Lessons learned from molecular biology of insulin gene mutations. Diabetes Care 13, 600–609 (1990).
Birkeland, K.I., Torjesen, P.A., Eriksson, J., Vaaler, S. & Groop, L. Hyperproinsulinemia of type II diabetes is not present before the development of hyperglycemia. Diabetes Care. 17, 1307–1310 (1994).
Gadot, M. et al. Hyperproinsulinemia and insulin deficiency in the diabetic Psammomys obesus. Endocrinology. 135, 610–616 (1994).
Poffenbarger, P.L., Chick, W.L., Lavine, R.L., Soeldner, J.S. & Flewelling, J.H. Insulin biosynthesis in experimental hereditary diabetes. Diabetes 20, 677–685 (1971).
Flatt, P.R., Bailey, C.J., Hampton, S.M., Swanston-Flatt, S.K. & Marks, V., C-peptide in spontaneous syndromes of obesity and diabetes in mice. Norm. Metabol. Res. 19, 1–5 (1987).
Ozcelik, T., Suedhof, T.C. & Francke, U. Chromosomal assignment of genes for vacuolar (endomembrane) proton pump subunits VPP1/Vpp-1 (116 KDa) and VPP3/Vpp-3 (58 kd) in human and mouse. Cytogen. Cell Genet. 58, 2008–2009 (1991).
Orci, L. et al. pH-independent and -dependent cleavage of proinsulin in the same secretory vesicle. J. Cell Biol. 126, 1149–1156 (1994).
Jung, Y.-K., Kunczt, C.J., Pearson, R.K., Dixon, J.E. & Fricker, L.D. Structural characterization of the rat carboxypeptidase-E gene. Molec. Endocrinol. 5, 1257–1268 (1991).
Stone, R.L. et al. A mutation in adenylosuccinate lyase associated with mental retardation and autistic features. Nature Genet. 1, 59–63 (1992).
Leiter, E.H. Type C retrovirus production by pancreatic beta cells. Association with accelerated pathogenesis in C3H-db/db (“diabetes”) mice. Am. J. Pathol 119, 22–32 (1985).
Prochazka, M., Serreze, D.V., Frankel, W.N. & Leiter, E.H. NOR/Lt; MHC-matched diabetes-resistant control strain for NOD mice. Diabetes 41, 98–106 (1992).
Lander, E. et al. MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1, 174–181 (1987).
Manley, K. A Macintosh program for storage and analysis of experimental genetic mapping data. Mamm. Genome 4, 301–313 (1993).
Tager, H.S., Rubenstein, A.H. & Steiner, D.F. in Methods in Enzymology (eds O'Malley, B. W. & Hardman, J.G.) 326–345 (Academic Press, New York, 1975).
Fricker, L.D. Methods for studying carboxypeptidase E. Meth. Neurosci. 23, 237–250 (1995).
Fricker, L.D., Das, B. & Angeletti, R.H. Identification of the pH-dependent membrane anchor of carboxypeptidase E (EC 3. 4.17.10). J. biol. Chem. 265, 2476–2482 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Naggert, J., Fricker, L., Varlamov, O. et al. Hyperproinsulinaemia in obese fat/fat mice associated with a carboxypeptidase E mutation which reduces enzyme activity. Nat Genet 10, 135–142 (1995). https://doi.org/10.1038/ng0695-135
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0695-135
This article is cited by
-
Carboxypeptidase E conditional knockout mice exhibit learning and memory deficits and neurodegeneration
Translational Psychiatry (2023)
-
Genomic loci mispositioning in Tmem120a knockout mice yields latent lipodystrophy
Nature Communications (2022)
-
Characterisation of genetic regulatory effects for osteoporosis risk variants in human osteoclasts
Genome Biology (2020)
-
Biochemical and genetic analysis of Ecm14, a conserved fungal pseudopeptidase
BMC Molecular and Cell Biology (2020)
-
Deciphering signature of selection affecting beef quality traits in Angus cattle
Genes & Genomics (2018)