Abstract
An effective HIV vaccine must elicit immune responses that recognize genetically diverse viruses1,2. It must generate CD8+ T lymphocytes that control HIV replication and CD4+ T lymphocytes that provide help for the generation and maintenance of both cellular and humoral immune responses against the virus3,4,5. Creating immunogens that can elicit cellular immune responses against the genetically varied circulating isolates of HIV presents a key challenge for creating an HIV vaccine6,7. Polyvalent mosaic immunogens derived by in silico recombination of natural strains of HIV are designed to induce cellular immune responses that recognize genetically diverse circulating virus isolates8. Here we immunized rhesus monkeys by plasmid DNA prime and recombinant vaccinia virus boost with vaccine constructs expressing either consensus or polyvalent mosaic proteins. As compared to consensus immunogens, the mosaic immunogens elicited CD8+ T lymphocyte responses to more epitopes of each viral protein than did the consensus immunogens and to more variant sequences of CD8+ T lymphocyte epitopes. This increased breadth and depth of epitope recognition may contribute both to protection against infection by genetically diverse viruses and to the control of variant viruses that emerge as they mutate away from recognition by cytotoxic T lymphocytes.
Similar content being viewed by others
Main
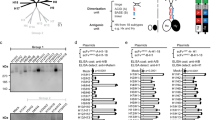
Mosaic immunogen sequences are designed with an algorithm that simulates evolution by recombination, using coverage of potential T lymphocyte epitopes as the selection criterion8. Nine-amino-acid fragments are considered potential epitopes, as nine amino acids is the most common length of major histocompatibility complex (MHC) class I–presented peptides and the most common span between anchor residues of MHC class II–presented peptides9. These sequences form complete proteins and are created with recombination breakpoints that are found in nature. We initiated this study to explore the breadth of CD4+ and CD8+ T lymphocyte responses generated through vaccination with polyvalent mosaic and consensus immunogens.
We selected two HIV genes for use in the immunogens employed in this study8: HIV-1 gag and nef. We separated 30 rhesus monkeys into three cohorts, two experimental groups consisting of 12 monkeys each and a control group consisting of six monkeys. One experimental group received mosaic immunogens, the other experimental group received consensus immunogens, and the control group received empty vectors. We chose a DNA prime–recombinant vaccinia virus boost regimen for this study, as these immunogens elicit both CD4+ and CD8+ T lymphocyte responses10,11,12.
We determined the breadth of the vaccine-elicited cellular immune responses by assessing peripheral blood T lymphocyte recognition of ten natural Gag and Nef sequences with a peptide-stimulated interferon-γ (IFN-γ) enzyme-linked immunospot (ELISPOT) assay and matrix epitope mapping. The Gag and Nef sets of ten indicator proteins included two clade A, two clade B, four clade C and two clade G sequences (a clade of HIV-1 sequences is a group of genetically related, usually geographically discrete, virus isolates). We used the same set of sequences that we had previously used to evaluate vaccine-elicited responses to a consensus Env immunogen13. We selected these isolate sequences to represent recently sampled viruses, diverse geographic origins and genetically distinct subregions of each clade. This strategy allowed us to assess vaccine-induced responses to sequences representative of the diverse HIV-1 strains to which human vaccinees will be exposed. Many nonamers in an HIV-1 strain are rare in the overall population of viruses (44% of Gag nonamers and 67% of Nef nonamers are present in <15% of wild-type sequences, on the basis of the Los Alamos database collection of M group sequences). As rare epitope variants are such a large component of the potential response, it is useful to know how often a vaccine can induce responses to rare nonamers in representative strains. This peptide design strategy allowed us to determine the precise number of vaccine-elicited responses per virus strain that were induced in each monkey and to characterize the cross-reactive potential of each epitope-specific response.
The magnitudes of the vaccine-induced Gag-specific and Nef-specific antibody and ELISPOT responses between the groups of monkeys were comparable (Supplementary Fig. 1a–c); therefore, any detected differences in the cross-reactivity of the cellular responses between these groups of vaccinated monkeys were not simply a manifestation of a more robust general immune response. The median number of epitope-specific total T lymphocyte responses generated by each monkey against Gag and Nef from a single virus strain after vaccination with consensus immunogens was 5.5 (interquartile range 3.0–11.75); the median number per strain after vaccination with mosaic immunogens was 7.75 (interquartile range 6.0–11.25). There was no statistically significant difference between these numbers of responses in the two experimental groups of monkeys (Wilcoxon rank-sum test).
To assess whether one of the vaccine strategies induced cellular immune responses that recognized a greater diversity of epitope variants, we determined the average number of variants recognized per epitope-specific response by dividing the total number of variant peptides recognized by the number of epitope-specific responses in each vaccinated monkey. The mosaic-vaccinated monkeys had higher ratios than the consensus-vaccinated monkeys (P = 0.045, Wilcoxon rank-sum test; Supplementary Table 1).
The vectors used in the present immunizations generated higher levels of CD4+ than of CD8+ T lymphocyte immune responses12, and any benefit for CD8+ T lymphocyte responses derived from using mosaic immunogens might be obscured in the overall response by the CD4+ T lymphocyte responses14. As CD4+ T lymphocytes recognize peptide antigens associated with MHC class II molecules, and peptide–MHC class II binding is more promiscuous than peptide–MHC class I binding, benefits associated with mosaic immunogens might be more apparent in CD8+ than CD4+ T lymphocyte responses15,16.
To determine the relative impact of mosaic and consensus immunogens on CD8+ and CD4+ T lymphocyte responses separately, we evaluated the breadth of responses to HIV Gag and Nef that were generated by mosaic and consensus gene-based immunizations using unfractionated and CD8+ T lymphocyte–depleted peripheral blood lymphocyte (PBL) populations. Sufficient lymphocytes were available from seven monkeys in each of the vaccinated groups to carry out this evaluation.
We determined the total number of epitope-specific responses to all tested strains of Gag and Nef by PBLs of each monkey and compared the groups with the Wilcoxon rank-sum test (Fig. 1a). The median number of epitope-specific CD4+ T lymphocyte responses per strain induced by vaccination of the monkeys with the mosaic immunogens was 4.5 (interquartile range 2.0–7.0), and it was 2.0 (interquartile range 1.25–4.0) in monkeys vaccinated with the consensus immunogens. The median number of epitope-specific CD8+ T lymphocyte responses per strain induced by vaccination of the monkeys with the mosaic immunogens was 2.0 (interquartile range 1.0–3.0), and it was 1.0 with the consensus immunogens (interquartile range 0.0–1.0) (Supplementary Fig. 2 contains a full breakdown of responses by protein and by CD4+ and CD8+ T lymphocytes). There were more total T lymphocyte responses (CD4+ plus CD8+) per strain induced by Gag mosaic immunogens than by consensus immunogens by a factor of two (P = 0.023, Poisson regression), but vaccine-induced Nef responses did not differ significantly between the two groups.
(a) CD4+ and CD8+ T lymphocyte responses from seven monkeys per group were assessed for their recognition of specific epitopes of each of the ten indicator Gag and Nef proteins in IFN-γ ELISPOT assays. Unfractionated and CD8+ lymphocyte-depleted PBLs were assessed for their recognition of specific epitope peptides. (b) The breadth of CD4+ and CD8+ T lymphocyte responses by individual monkeys was determined as follows: if a 15-mer peptide from one of the ten sets of indicator proteins was recognized by a T lymphocyte population, it was scored as one positive; if multiple variant forms of the same peptide from more than one set of indicator proteins were recognized by a T lymphocyte population, they were also scored as one positive response. (c) The depth of CD4+ and CD8+ T lymphocyte responses by individual monkeys, as determined by counting the responses made to all variant peptides from each epitopic region of either the Gag or the Nef protein.
We then compared the number of epitope-specific CD4+ and CD8+ T lymphocyte responses per monkey, counting overlapping peptides or sequence variants as one event (Fig. 1b). There was no significant difference in the number of total (Gag-specific plus Nef-specific) (Wilcoxon rank-sum statistics; P = 0.28), Gag-specific (P = 0.18) or Nef-specific (P = 0.45) CD4+ T lymphocyte responses elicited by consensus or mosaic immunogens (Fig. 1b). However, mosaic immunogens induced a significantly larger number of total (P = 0.001) and Gag-specific (P = 0.006) and a trend toward more Nef-specific (P = 0.158) CD8+ T lymphocyte responses.
We also assessed whether mosaic-immunized monkeys developed CD4+ and CD8+ T lymphocyte responses with greater depth than consensus-immunized monkeys by quantifying the number of responses to all of the variant forms of a given epitope (Fig.1c). There were more Gag-specific (P = 0.05) but not Nef-specific (P = 0.40) CD4+ T lymphocyte responses to variant peptides per response in the mosaic-immunized group; there was a median number of 1 (range 1–3) Gag epitope–specific CD4+ T lymphocyte response in the consensus-immunized monkeys and a median number of 3 (range 2–8) of such responses in the mosaic-immunized monkeys (Fig. 1c). For CD8+ T lymphocyte responses, there were more total (Gag-specific plus Nef-specific) responses (P = 0.005), Gag-specific responses (P = 0.001) and a trend toward more Nef-specific responses (P = 0.16) in the mosaic-immunized monkeys than in the consensus-immunized monkeys (Fig. 1c). These findings are comparable to those reported by Barouch et al.17 in this issue (a median of 1 (range 0–2) in the consensus-immunized monkeys and a median of 3 (range 2–4) in the mosaic-immunized monkeys). Together, these data suggest that the benefit of the mosaic immunogens was more apparent for CD8+ T lymphocyte than for CD4+ T lymphocyte responses.
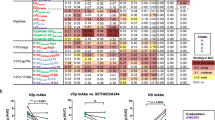
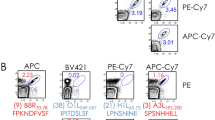
Because we evaluated the reactivity of the PBLs of each vaccinated monkey to peptides spanning the entire Gag and Nef proteins of ten genetically disparate HIV-1 isolates, we were able to define the immune recognition of actual proteins from distinct circulating virus strains in the immunized monkeys. Examples of typical responses and our approach to illustrating them are shown in Figure 2; the complete alignment of all peptides to which at least a single CD8+ T lymphocyte response was made is shown in Supplementary Figure 3.
Consensus and mosaic vaccine-induced CD8+ T lymphocyte responses were assessed for recognition of variant forms of the same region of a viral protein. For each example, the variant HIV-1 sequences are shown aligned to the M group consensus of that sequence. Amino acid identity to the consensus sequence is indicated by a dash. For some sequences, a blank space is inserted to maintain the alignment. The peptides recognized by PBLs of a vaccinated monkey are shown in black at the top and are preceded by the number of the responding monkey. The variant peptides in the same region that are not recognized are shown in red. Every unique peptide sequence recognized by PBLs is shown in a different shade of green, and white boxes represent peptides that are not recognized. A large box represents an exact match of a number of sequences. (a) A highly restricted CD8+ T lymphocyte response. CD8+ T lymphocytes from monkey 58 recognize only the peptide sequence that matches the vaccine sequence. (b) A cross-reactive CD8+ T lymphocyte response. Three different variants of peptide 15 and 16 sequences exist in ten indicator Gag proteins, and CD8+ T lymphocytes from monkey 228 recognize all three variants. (c) A highly restricted CD8+ T lymphocyte response. CD8+ T lymphocytes of monkey 65 recognize only the variant peptide that matches one of the four mosaic sequences used in the mosaic immunogen cocktail. (d) A cross-reactive CD8+ T lymphocyte response. CD8+ T lymphocytes from monkey 65 recognize four different variant forms of the peptide, all of which differ in sequence from the vaccine immunogens.
Some vaccine-induced CD8+ T lymphocyte responses were highly strain specific, capable of recognizing peptide variants from only one or two of the ten test strains (Figs. 2a,c and 3). We observed broadly cross-reactive responses to individual epitopes, but they were very rare, with all ten peptide variants recognized only six times in all vaccinated monkeys (Figs. 2b and 3). There were some responses to two overlapping peptides, probably representing the recognition of a single epitope (Fig. 2b). CD8+ T lymphocyte responses were usually highly specific. Extending the breadth of responses to more than two variant peptides was rare in both groups of vaccinated monkeys. This type of potentially valuable response occurred only six times among the consensus vaccine–induced CD8+ T lymphocyte responses and 17 times among responses in mosaic vaccine recipients (binomial test P = 0.03; Figs. 2d and 3).
All of the Gag and Nef peptides recognized by PBLs of individual monkeys were aligned to the HIV-1 M group consensus sequences as described in the legend of Figure 2. Lymphocyte recognition of any peptide sequence is shown in rectangles of shades of green, orange or yellow, and nonrecognition of a peptide is shown in unshaded rectangles.
Modeling the depth of the peptide-specific responses as a function of vaccine and T lymphocyte type, we found that mosaic vaccine–elicited CD8+ T lymphocyte responses were more likely than consensus vaccine–elicited CD8+ T lymphocyte responses to recognize variant peptides by a factor of 1.6 (P = 0.0006, binomial regression; Fig. 3 and Supplementary Fig. 4). Extending the breadth of a response to more than two variant peptides occurred 42 times among CD4+ T lymphocyte responses in the mosaic–immunized monkeys and only 23 times in the consensus-vaccinated monkeys (binomial test P = 0.05; Fig. 3 and Supplementary Figs. 5 and 6). CD4+ T lymphocyte responses usually had greater depth (encompass a larger fraction of variants) than CD8+ T lymphocyte responses by a factor of 1.48 (P = 0.0004); this was true for both vaccine prototypes and may explain why the benefit of mosaic immunogens was more pronounced for CD8+ T lymphocytes.
Almost all of the indicator strains of Gag and Nef proteins were recognized by one or two epitope-specific CD4+ T lymphocyte responses in the PBLs of both groups of vaccinated monkeys (Fig. 4a). However, we detected more CD4+ T lymphocyte responses that recognized three epitopes in mosaic-vaccinated monkeys than in consensus-vaccinated monkeys. Almost all of the indicator strains of peptides were recognized by at least one epitope-specific CD8+ T lymphocyte response in the PBLs of the mosaic vaccine recipients, whereas a substantial number of viral strains were not recognized by epitope-specific CD8+ T lymphocyte peptide responses in the consensus-immunized monkeys (P = 0.057, Wilcoxon rank sum test; Fig. 4b). The mosaic immunogens also elicited a significantly larger number of CD8+ T lymphocyte responses that recognized a minimum of two (P = 0.0022) and three epitopes (P = 0.0018) of these natural virus strains as compared to the consensus immunogen.
(a,b) The stacks of ten rectangles represent the ten sets of indicator peptides with A1 and A2 shown in turquoise; B1 and B2 in red; C1, C2, C3 and C4 in purple; and G1 and G2 in dark blue. For each monkey, in the left-hand column (min 1), if at least one peptide from an indicator peptide set was recognized by either CD4+ (a) or CD8+ (b) T lymphocytes, the rectangle is shaded with its representative color. If no PBL responses to a particular set of indicator peptides were detected, the rectangle is shown unshaded. Recognition of one, two or three epitopes per strain of indicator proteins by CD4+ and CD8+ T lymphocytes of each of the seven monkeys in both vaccine groups is shown as min 1, min 2 and min 3, respectively.
In the recent STEP human vaccine trial, an adenovirus-based HIV-1 Gag-Pol-Nef vaccine induced a median of only two epitope-specific responses to the vaccine in each subject (N. Frahm, personal communication), only 40% of the subjects mounted a response to Gag, and only 32% of vaccinees generated both a CD4+ and a CD8+ T lymphocyte response18. It is possible that this vaccine failed to confer protection because most of the vaccine-elicited responses were not able to recognize the circulating strains of HIV-1.
Our study demonstrates that immunization of nonhuman primates with mosaic immunogens induces CD8+ T lymphocytes with greater cross-reactivity than the consensus immunogen. The mosaic immunogens elicited CD8+ T lymphocyte responses to more epitopes of each viral protein than the consensus immunogens and to more variant sequences of the CD8+ T lymphocyte epitopes. This increased breadth and depth of epitope recognition could contribute to protection against infection by genetically diverse viruses and, in some instances, may block the emergence of common variant viruses.
Methods
Experimental groups and vaccination schedule.
We housed the monkeys at the New England Regional Primate Research Center. We maintained the monkeys in accordance with standards set forth by American Association for Accreditation of Laboratory Animal Care and the study protocols were approved by the Harvard University Medical School Institutional Animal Care and Use Committee. We distributed 30 Mamu-A*01–, Mamu-B*01– and Mamu-B*17–negative rhesus monkeys into three groups, two experimental groups each consisting of 12 monkeys and a control group consisting of six monkeys. One of the experimental groups received immunogens containing single consensus gene inserts and the other received immunogens containing a cocktail of four complementary mosaic gene inserts. On weeks 0, 4 and 8, the experimental groups of monkeys received priming immunizations by the intramuscular route with either a total of 5 mg of M consensus gag or mosaic gag and a total of either 5 mg of M consensus nef or mosaic nef plasmid DNA for each immunization. At week 33, we boosted the monkeys by simultaneous intramuscular and intradermal inoculations with a total of 1 × 109 plaque-forming units of a recombinant vaccinia virus expressing the same antigens. We immunized the control monkeys with empty vectors.
Pooled peptide and peptide matrix interferon-γ enzyme-linked immunospot assays.
We coated multiscreen 96-well plates overnight with 100 μl per well of 5 μg ml−1 antibody to human IFN-γ (B27; BD Pharmingen) in endotoxin-free Dulbecco's-PBS (D-PBS). We then washed the plates three times with D-PBS containing 0.1% Tween-20 and blocked them for 2 h with RPMI-1640 containing 10% FBS. Then we added peptide pools and 2 × 105 PBLs to each well in 100-μl reaction volumes in triplicate for pooled peptides assays and in duplicate for peptide matrix assays. Each peptide pool was comprised of 15–amino acid peptides overlapping by 11 amino acids. The pools covered the entire HIV-1 Gag and Nef proteins. Each peptide in a pool was present at a 1 μg ml−1 concentration. After an 18-h incubation at 37 °C, we washed the plates nine times with D-PBS containing 0.1% Tween-20 and once with distilled water. We then incubated the plates with 2 μg ml−1 biotinylated rabbit antibody to human IFN-γ (U-Cytech) for 2 h at 25 °C, washed them six times with D-PBS containing 0.1% Tween-20 and incubated them for 2.5 h with a 1 in 500 dilution of streptavidin-AP (Southern Biotechnology Associates). After five washes with D-PBS containing 0.1% Tween-20 and three washes with D-PBS, we developed the plates with NBT/BCIP chromogen (Pierce), stopped the reaction by washing with tap water, air dried the plates and read them with an ELISPOT reader (Cellular Technology Ltd.). We calculated the number of spot-forming cells per 1 × 106 PBLs. The responses to the indicator peptides in the control group of six monkeys were in the range of 5–108 SFCs per million PBLs. These values were always less than four times below the medium-alone values.
CD8+ T lymphocyte depletion.
We incubated PBLs from the monkeys with phycoerythrin-labeled CD8-specific antibody and then with magnetic beads coated in antibody to phycoerythrin (Miltenyi Biotech) following manufacturer's protocol. We then sorted the labeled PBLs with a Miltenyi AutoMACS cell sorter to deplete CD8+ T lymphocytes. The efficiency of CD8+ T cell depletion in this study was 94.3%–99.7%. We measured IFN-γ ELISPOT responses in unfractionated and CD8+ T lymphocyte–depleted PBLs from these monkeys. In the evaluation of the cellular immune response, if greater than one half of a SFC response by unfractionated PBLs was eliminated by CD8+ T lymphocyte depletion, we designated that response as CD8+ T lymphocyte mediated. The responses of CD8+ T lymphocyte–depleted PBLs were designated as CD4+ T lymphocyte responses. These highly stringent criteria may underestimate the magnitude of the vaccine-elicited CD4+ T lymphocyte responses.
Additional methods.
Detailed methodology is described in the Supplementary Methods.
References
Korber, B.T., Letvin, N.L. & Haynes, B.F. T-cell vaccine strategies for human immunodeficiency virus, the virus with a thousand faces. J. Virol. 83, 8300–8314 (2009).
Barouch, D.H. Challenges in the development of an HIV-1 vaccine. Nature 455, 613–619 (2008).
Letvin, N.L. Progress and obstacles in the development of an AIDS vaccine. Nat. Rev. Immunol. 6, 930–939 (2006).
Heeney, J.L. Requirement of diverse T-helper responses elicited by HIV vaccines: induction of highly targeted humoral and CTL responses. Expert Rev. Vaccines 3, S53–S64 (2004).
McMichael, A.J. HIV vaccines. Annu. Rev. Immunol. 24, 227–255 (2006).
Fischer, W., Liao, H.X., Haynes, B.F., Letvin, N.L. & Korber, B. Coping with viral diversity in HIV vaccine design: a response to Nickle et al. PLoS Comput. Biol. 4, e15 (2008).
Gaschen, B. et al. Diversity considerations in HIV-1 vaccine selection. Science 296, 2354–2360 (2002).
Fischer, W. et al. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nat. Med. 13, 100–106 (2007).
Rammensee, H., Bachmann, J., Emmerich, N.P., Bachor, O.A. & Stevanovic, S. SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics 50, 213–219 (1999).
Santra, S. et al. Recombinant poxvirus boosting of DNA-primed rhesus monkeys augments peak but not memory T lymphocyte responses. Proc. Natl. Acad. Sci. USA 101, 11088–11093 (2004).
Precopio, M.L. et al. Immunization with vaccinia virus induces polyfunctional and phenotypically distinctive CD8+ T cell responses. J. Exp. Med. 204, 1405–1416 (2007).
Harari, A. et al. An HIV-1 clade C DNA prime, NYVAC boost vaccine regimen induces reliable, polyfunctional and long-lasting T cell responses. J. Exp. Med. 205, 63–77 (2008).
Santra, S. et al. A centralized gene-based HIV-1 vaccine elicits broad cross-clade cellular immune responses in rhesus monkeys. Proc. Natl. Acad. Sci. USA 105, 10489–10494 (2008).
Kong, W.P. et al. Expanded breadth of the T-cell response to mosaic human immunodeficiency virus type 1 envelope DNA vaccination. J. Virol. 83, 2201–2215 (2009).
Moss, C.X., Tree, T.I. & Watts, C. Reconstruction of a pathway of antigen processing and class II MHC peptide capture. EMBO J. 26, 2137–2147 (2007).
Sette, A., Adorini, L., Colon, S.M., Buus, S. & Grey, H.M. Capacity of intact proteins to bind to MHC class II molecules. J. Immunol. 143, 1265–1267 (1989).
Barouch, D.H. et al. Mosaic HIV-1 vaccines expand the breadth and depth of cellular immune responses in rhesus monkeys. Nat. Med. advance online publication, doi:10.1038/nm.2089 (21 February 2010).
McElrath, M.J. et al. HIV-1 vaccine-induced immunity in the test-of-concept Step Study: a case-cohort analysis. Lancet 372, 1894–1905 (2008).
Acknowledgements
We thank G. Tomaras for generous advice and assistance with reagents and antibody data. We also thank C. Lord for assistance with reagents. This work was supported by US National Institutes of Health grant AI-0678501 (Center for HIV AIDS Vaccine Immunology), HIV Research and Design (HIVRAD) P01 AI-61734, National Institute of Allergy and Infectious Diseases contract N01 AI-50010 and a Los Alamos National Laboratory–directed research grant to B.T.K., W.F. and S.W.
Author information
Authors and Affiliations
Contributions
B.T.K., W.F., G.N.P. and B.K.F. designed the vaccine gene inserts. H.-X.L., R.Z. and B.F.H. generated the recombinant vaccinia virus constructs. B.T.K., B.F.H., S.S. and N.L.L. designed the study. A.C. and K.G.M. performed all of the immunizations. H.B., A.B., D.Q. and S.S. designed and conducted the cellular immunologic assays. R.J.P., C.-Y.T. and B.F.H. designed and conducted the antibody assays. M.M., S.W., W.F., J.T., J.S. and B.T.K. performed the data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Table 1 and Supplementary Methods (PDF 1300 kb)
Rights and permissions
About this article
Cite this article
Santra, S., Liao, HX., Zhang, R. et al. Mosaic vaccines elicit CD8+ T lymphocyte responses that confer enhanced immune coverage of diverse HIV strains in monkeys. Nat Med 16, 324–328 (2010). https://doi.org/10.1038/nm.2108
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2108
This article is cited by
-
Recombinant MVA-prime elicits neutralizing antibody responses by inducing antigen-specific B cells in the germinal center
npj Vaccines (2021)
-
CD8+ T cells in HIV control, cure and prevention
Nature Reviews Immunology (2020)
-
Mathematical model of broadly reactive plasma cell production
Scientific Reports (2020)
-
Development of an anti-HIV vaccine eliciting broadly neutralizing antibodies
AIDS Research and Therapy (2017)
-
Epigraph: A Vaccine Design Tool Applied to an HIV Therapeutic Vaccine and a Pan-Filovirus Vaccine
Scientific Reports (2016)