Abstract
Mammographic density (MD) reflects variations in fat, stromal and epithelial tissues that are thought to be regulated by several genes. High MD is an established risk factor for breast cancer; therefore, genes that regulate MD may indirectly influence breast cancer. These genes might also be fewer in number and easier to identify than those for breast cancer risk outside of inherited predisposition syndromes. In this Perspective, we review the limited genetic studies of MD and propose future directions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Turner, S. T. et al. Genomic loci with pleiotropic effects on coronary artery calcification. Atherosclerosis 185, 340–346 (2006).
Kullo, I. J., Ding, K., Boerwinkle, E., Turner, S. T. & de Andrade, M. Quantitative trait loci influencing low density lipoprotein particle size in African Americans. J. Lipid Res. 47, 1457–1462 (2006).
Carlson, C. S. et al. Selecting a maximally informative set of single-nucleotide polymorphisms for association analyses using linkage disequilibrium. Am. J. Hum. Genet. 74, 106–120 (2004).
Wolfe, J. N. Breast patterns as an index of risk for developing breast cancer. AJR Am. J. Roentgenol. 126, 1130–1137 (1976).
Lehman, C., Holt, S., Peacock, S., White, E. & Urban, N. Use of the American College of Radiology BI-RADS guidelines by community radiologists: concordance of assessments and recommendations assigned to screening mammograms. AJR Am. J. Roentgenol. 179, 15–20 (2002).
Wolfe, J. N., Saftlas, A. F. & Salane, M. Mammographic parenchymal patterns and quantitative evaluation of mammographic densities: a case-control study. AJR Am. J. Roentgenol. 148, 1087–1092 (1987).
Boyd, N. F. et al. Quantitative classification of mammographic densities and breast cancer risk: results from the Canadian National Breast Screening Study. J. Natl Cancer Inst. 87, 670–675 (1995).
Greendale, G. A. et al. Postmenopausal hormone therapy and change in mammographic density. J. Natl Cancer Inst. 95, 30–37 (2003).
Boyd, N. F. et al. Mammographic breast density as an intermediate phenotype for breast cancer. Lancet Oncol. 6, 798–808 (2005).
McCormack, V. A. & Dos Santos Silva, I. Breast density and parenchymal patterns as markers of breast cancer risk: a meta-analysis. Cancer Epidemiol Biomarkers Prev. 15, 1159–1169 (2006).
Byrne, C. et al. Mammographic features and breast cancer risk: effects with time, age, and menopause status. J. Natl Cancer Inst. 87, 1622–1629 (1995).
Tice, J. A., Cummings, S. R., Ziv, E. & Kerlikowske, K. Mammographic breast density and the Gail model for breast cancer risk prediction in a screening population. Breast Cancer Res. Treat. 94, 115–122 (2005).
Chen, J. et al. Projecting absolute invasive breast cancer risk in white women with a model that includes mammographic density. J. Natl Cancer Inst. 98, 1215–1226 (2006).
Barlow, W. E. et al. Prospective breast cancer risk prediction model for women undergoing screening mammography. J. Natl Cancer Inst. 98, 1204–1214 (2006).
Wilkinson, E. et al. Mammographic parenchymal pattern and the risk of breast cancer. J. Natl Cancer Inst. 59, 1397–1400 (1977).
Hainline, S. et al. Mammographic patterns and risk of breast cancer. AJR Am. J. Roentgenol. 130, 1157–1158 (1978).
Saftlas, A. F. et al. Mammographic parenchymal patterns as indicators of breast cancer risk. Am. J. Epidemiol. 129, 518–526 (1989).
Saftlas, A. F. et al. Mammographic densities and risk of breast cancer. Cancer 67, 2833–2838 (1991).
Kaufman, Z., Garstin, W. I., Hayes, R., Michell, M. J. & Baum, M. The mammographic parenchymal patterns of nulliparous women and women with a family history of breast cancer. Clin. Radiol. 43, 385–388 (1991).
Brisson, J. et al. The relation of mammographic features of the breast to breast cancer risk factors. Am. J. Epidemiol. 115, 438–443 (1982).
Crest, A. B., Aiello, E. J., Anderson, M. L. & Buist, D. S. Varying levels of family history of breast cancer in relation to mammographic breast density (United States). Cancer Causes Control 17, 843–850 (2006).
Ziv, E., Shepherd, J., Smith-Bindman, R. & Kerlikowske, K. Mammographic breast density and family history of breast cancer. J. Natl Cancer Inst. 95, 556–558 (2003).
Wolfe, J. N., Albert, S., Belle, S. & Salane, M. Familial influences on breast parenchymal patterns. Cancer 46, 2433–2437 (1980).
Kaprio, J., Alanko, A., Kivisaari, L. & Standertskjol d-Nordenstam, C. G. Mammographic patterns in twin pairs discordant for breast cancer. Br. J. Radiol. 60, 459–462 (1987).
Haars, G., van Noord, P. A., van Gils, C. H., Peeters, P. H. & Grobbee, D. E. Heritable aspects of dysplastic breast glandular tissue (DY). Breast Cancer Res. Treat. 87, 149–156 (2004).
Boyd, N. F. et al. Heritability of mammographic density, a risk factor for breast cancer. N. Engl. J. Med 347, 886–894 (2002).
Stone, J. et al. The heritability of mammographically dense and nondense breast tissue. Cancer Epidemiol. Biomarkers Prev. 15, 612–617 (2006).
Ursin, G. et al. A revised heritability estimate of mammographic density. Proc. Am. Assoc. Cancer Res. (2007).
Pankow, J. S. et al. Genetic analysis of mammographic breast density in adult women: evidence of a gene effect. J. Natl Cancer Inst. 89, 549–556 (1997).
Tamimi, R. M. et al. Common genetic variation in IGF1, IGFBP-1, and IGFBP-3 in relation to mammographic density: a cross-sectional study. Breast Cancer Res. 9, R18 (2007).
Olson, J. E. et al. A comprehensive examination of CYP19 variation and breast density. Cancer Epidemiol. Biomarkers Prev. 16, 623–625 (2007).
Verheus, M. et al. Common genetic variation in the IGF-1 gene, serum IGF-I levels and breast density. Breast Cancer Res. Treat. 7 Dec 2007 (doi:10.1007/s10549-007-9827-x).
Helvie, M. A., Roubidoux, M. A., Weber, B. L. & Merajver, S. D. Mammography of breast carcinoma in women who have mutations of the breast cancer gene BRCA1: initial experience. AJR Am. J. Roentgenol. 168, 1599–1602 (1997).
Chang, J., Yang, W. T. & Choo, H. F. Mammography in Asian patients with BRCA1 mutations. Lancet 353, 2070–2071 (1999).
Huo, Z. et al. Computerized analysis of mammographic parenchymal patterns for breast cancer risk assessment: feature selection. Med. Phys. 27, 4–12 (2000).
Tilanus-Linthorst, M. et al. A BRCA1/2 mutation, high breast density and prominent pushing margins of a tumor independently contribute to a frequent false-negative mammography. Int. J. Cancer 102, 91–95 (2002).
Lord, S. J. et al. Polymorphisms in genes involved in estrogen and progesterone metabolism and mammographic density changes in women randomized to postmenopausal hormone therapy: results from a pilot study. Breast Cancer Res. 7, R336–R344 (2005).
Takata, Y., Maskarinec, G. & Le Marchand, L. Breast density and polymorphisms in genes coding for CYP1A2 and COMT: the Multiethnic Cohort. BMC Cancer 7, 30 (2007).
Haiman, C. A. et al. Genetic determinants of mammographic density. Breast Cancer Res. 4, R5 (2002).
Lillie, E. O. et al. Polymorphism in the androgen receptor and mammographic density in women taking and not taking estrogen and progestin therapy. Cancer Res. 64, 1237–1241 (2004).
Haiman, C. A. et al. Polymorphisms in steroid hormone pathway genes and mammographic density. Breast Cancer Res. Treat. 77, 27–36 (2003).
Hong, C. C. et al. Val158Met polymorphism in catechol-O-methyltransferase gene associated with risk factors for breast cancer. Cancer Epidemiol. Biomarkers Prev. 12, 838–847 (2003).
Hong, C. C. et al. Association between the T27C polymorphism in the cytochrome P450 c17α (CYP17) gene and risk factors for breast cancer. Breast Cancer Res. Treat. 88, 217–230 (2004).
Mulhall, C. et al. Pituitary growth hormone and growth hormone-releasing hormone receptor genes and associations with mammographic measures and serum growth hormone. Cancer Epidemiol. Biomarkers Prev. 14, 2648–2654 (2005).
Lai, J. H. et al. A polymorphic locus in the promoter region of the IGFBP3 gene is related to mammographic breast density. Cancer Epidemiol. Biomarkers Prev. 13, 573–582 (2004).
Maskarinec, G., Lurie, G., Williams, A. E. & Le Marchand, L. An investigation of mammographic density and gene variants in healthy women. Int. J. Cancer 112, 683–688 (2004).
van Duijnhoven, F. J. et al. Polymorphisms in the estrogen receptor α gene and mammographic density. Cancer Epidemiol. Biomarkers Prev. 14, 2655–2660 (2005).
dos Santos Silva, I. et al. The insulin-like growth factor system and mammographic features in premenopausal and postmenopausal women. Cancer Epidemiol. Biomarkers Prev. 15, 449–455 (2006).
Mitchell, G. et al. Mammographic density and breast cancer risk in BRCA1 and BRCA2 mutation carriers. Cancer Res. 66, 1866–1872 (2006).
Stone, J. et al. Mammographic density and candidate gene variants: a twins and sisters study. Cancer Epidemiol. Biomarkers Prev. 16, 1479–1484 (2007).
Lee, E. et al. The role of established breast cancer susceptibility loci in mammographic density in young women. Cancer Epidemiol. Biomarkers Prev. 17, 258–260 (2008).
van Duijnhoven, F. J. et al. Influence of estrogen receptor α and progesterone receptor polymorphisms on the effects of hormone therapy on mammographic density. Cancer Epidemiol. Biomarkers Prev. 15, 462–467 (2006).
Warren, R. et al. Associations among mammographic density, circulating sex hormones, and polymorphisms in sex hormone metabolism genes in postmenopausal women. Cancer Epidemiol. Biomarkers Prev. 15, 1502–1508 (2006).
Diorio, C., Brisson, J., Berube, S. & Pollak, M. Genetic polymorphisms involved in insulin-like growth factor (IGF) pathway in relation to mammographic breast density and IGF levels. Cancer Epidemiol. Biomarkers Prev. 17, 880–888 (2008).
Olson, J. E. et al. Mammographic breast density and membrane-bound catechol O-methyltransferase (COMT) genetic polymorphisms. Proc Am. Assoc. Cancer Res. (in the press).
Nedelcheva Kristensen, V. et al. CYP17 and breast cancer risk: the polymorphism in the 5′ flanking area of the gene does not influence binding to Sp-1. Cancer Res. 59, 2825–2828 (1999).
Dunning, A. M. et al. A systematic review of genetic polymorphisms and breast cancer risk. Cancer Epidemiol. Biomarkers Prev. 8, 843–854 (1999).
Lachman, H. M. et al. Human catechol-O-methyltransferase pharmacogenetics: description of a functional polymorphism and its potential application to neuropsychiatric disorders. Pharmacogenetics 6, 243–250 (1996).
Le Marchand, L., Donlon, T., Kolonel, L. N., Henderson, B. E. & Wilkens, L. R. Estrogen metabolism-related genes and breast cancer risk: the multiethnic cohort study. Cancer Epidemiol. Biomarkers Prev. 14, 1998–2003 (2005).
Yaich, L., Dupont, W. D., Cavener, D. R. & Parl, F. F. Analysis of the PvuII restriction fragment-length polymorphism and exon structure of the estrogen receptor gene in breast cancer and peripheral blood. Cancer Res. 52, 77–83 (1992).
Cai, Q. et al. Genetic polymorphisms in the estrogen receptor α gene and risk of breast cancer: results from the Shanghai Breast Cancer Study. Cancer Epidemiol. Biomarkers Prev. 12, 853–859 (2003).
Onland-Moret, N. C., van Gils, C. H., Roest, M., Grobbee, D. E. & Peeters, P. H. The estrogen receptor α gene and breast cancer risk (The Netherlands). Cancer Causes Control 16, 1195–1202 (2005).
Wedren, S. et al. Oestrogen receptor α gene haplotype and postmenopausal breast cancer risk: a case control study. Breast Cancer Res. 6, R437–449 (2004).
De Vivo, I. et al. A functional polymorphism in the promoter of the progesterone receptor gene associated with endometrial cancer risk. Proc. Natl Acad. Sci. USA 99, 12263–12268 (2002).
Modugno, F. Ovarian cancer and polymorphisms in the androgen and progesterone receptor genes: a HuGE review. Am. J. Epidemiol. 159, 319–335 (2004).
Breast Cancer Association Consortium. Commonly studied single-nucleotide polymorphisms and breast cancer: results from the Breast Cancer Association Consortium. J. Natl Cancer Inst. 98, 1382–1396 (2006).
Deal, C. et al. Novel promoter polymorphism in insulin-like growth factor-binding protein-3: correlation with serum levels and interaction with known regulators. J. Clin. Endocrinol. Metab. 86, 1274–1280 (2001).
Schernhammer, E. S., Hankinson, S. E., Hunter, D. J., Blouin, M. J. & Pollak, M. N. Polymorphic variation at the −202 locus in IGFBP3: Influence on serum levels of insulin-like growth factors, interaction with plasma retinol and vitamin D and breast cancer risk. Int. J. Cancer 107, 60–64 (2003).
Al-Zahrani, A. et al. IGF1 and IGFBP3 tagging polymorphisms are associated with circulating levels of IGF1, IGFBP3 and risk of breast cancer. Hum. Mol. Genet. 15, 1–10 (2006).
Setiawan, V. W. et al. IGF-I genetic variation and breast cancer: the multiethnic cohort. Cancer Epidemiol. Biomarkers Prev. 15, 172–174 (2006).
Canzian, F. et al. Polymorphisms of genes coding for insulin-like growth factor 1 and its major binding proteins, circulating levels of IGF-I and IGFBP-3 and breast cancer risk: results from the EPIC study. Br. J. Cancer 94, 299–307 (2006).
Mitrunen, K. & Hirvonen, A. Molecular epidemiology of sporadic breast cancer. The role of polymorphic genes involved in oestrogen biosynthesis and metabolism. Mutat. Res. 544, 9–41 (2003).
Puranen, T. J., Poutanen, M. H., Peltoketo, H. E., Vihko, P. T. & Vihko, R. K. Site-directed mutagenesis of the putative active site of human 17 β-hydroxysteroid dehydrogenase type 1. Biochem. J. 304 (Pt 1), 289–293 (1994).
Cai, Q. et al. Haplotype analyses of CYP19A1 gene variants and breast cancer risk: results from the Shanghai Breast Cancer Study. Cancer Epidemiol. Biomarkers Prev. 17, 27–32 (2008).
Healey, C. S. et al. Polymorphisms in the human aromatase cytochrome P450 gene (CYP19) and breast cancer risk. Carcinogenesis 21, 189–193 (2000).
Tut, T. G., Ghadessy, F. J., Trifiro, M. A., Pinsky, L. & Yong, E. L. Long polyglutamine tracts in the androgen receptor are associated with reduced trans-activation, impaired sperm production, and male infertility. J. Clin. Endocrinol. Metab. 82, 3777–3782 (1997).
Grierson, A. J., Mootoosamy, R. C. & Miller, C. C. Polyglutamine repeat length influences human androgen receptor/c-Jun mediated transcription. Neurosci. Lett. 277, 9–12 (1999).
Lillie, E. O., Bernstein, L. & Ursin, G. The role of androgens and polymorphisms in the androgen receptor in the epidemiology of breast cancer. Breast Cancer Res. 5, 164–173 (2003).
Rebbeck, T. R. et al. Modification of BRCA1-associated breast cancer risk by the polymorphic androgen-receptor CAG repeat. Am. J. Hum. Genet. 64, 1371–1377 (1999).
Antoniou, A. et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case series unselected for family history: a combined analysis of 22 studies. Am. J. Hum. Genet. 72, 1117–1130 (2003).
Brockstedt, U. et al. Analyses of bulky DNA adduct levels in human breast tissue and genetic polymorphisms of cytochromes P450 (CYPs), myeloperoxidase (MPO), quinone oxidoreductase (NQO1), and glutathione S-transferases (GSTs). Mutat. Res. 516, 41–47 (2002).
Long, J. R. et al. Population-based case-control study of AhR (aryl hydrocarbon receptor) and CYP1A2 polymorphisms and breast cancer risk. Pharmacogenet. Genomics 16, 237–243 (2006).
Masson, L. F., Sharp, L., Cotton, S. C. & Little, J. Cytochrome P-450 1A1 gene polymorphisms and risk of breast cancer: a HuGE review. Am. J. Epidemiol. 161, 901–915 (2005).
Shimada, T. et al. Catalytic properties of polymorphic human cytochrome P450 1B1 variants. Carcinogenesis 20, 1607–1613 (1999).
Lee, A. J., Cai, M. X., Thomas, P. E., Conney, A. H. & Zhu, B. T. Characterization of the oxidative metabolites of 17β-estradiol and estrone formed by 15 selectively expressed human cytochrome p450 isoforms. Endocrinology 144, 3382–3398 (2003).
Gaudet, M. M. et al. Genetic variation of Cytochrome P450 1B1 (CYP1B1) and risk of breast cancer among Polish women. Pharmacogenet. Genomics 16, 547–553 (2006).
Wacholder, S., Chanock, S., Garcia-Closas, M., El Ghormli, L. & Rothman, N. Assessing the probability that a positive report is false: an approach for molecular epidemiology studies. J. Natl Cancer Inst. 96, 434–442 (2004).
Vachon, C. M., King, R. A., Atwood, L. D., Kuni, C. C. & Sellers, T. A. Preliminary sibpair linkage analysis of percent mammographic density. J. Natl Cancer Inst. 91, 1778–1779 (1999).
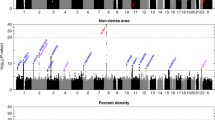
Vachon, C. M. et al. Strong evidence of a genetic determinant for mammographic density, a major risk factor for breast cancer. Cancer Res. 67, 8412–8418 (2007).
Easton, D. F. et al. Genome-wide association study identifies novel breast cancer susceptibility loci. Nature 447, 1087–1093 (2007).
Hunter, D. J. et al. A genome-wide association study identifies alleles in FGFR2 associated with risk of sporadic postmenopausal breast cancer. Nature Genet. 39, 870–874 (2007).
Stacey, S. N. et al. Common variants on chromosomes 2q35 and 16q12 confer susceptibility to estrogen receptor-positive breast cancer. Nature Genet. 39, 865–869 (2007).
Boyd, N. F., Jensen, H. M., Cooke, G. & Han, H. L. Relationship between mammographic and histological risk factors for breast cancer. J. Natl Cancer Inst. 84, 1170–1179 (1992).
Byng, J. W., Boyd, N. F., Fishell, E., Jong, R. A. & Yaffe, M. J. The quantitative analysis of mammographic densities. Phys. Med. Biol. 39, 1629–1638 (1994).
Fletcher, O. et al. Polymorphisms and circulating levels in the insulin-like growth factor system and risk of breast cancer: a systematic review. Cancer Epidemiol. Biomarkers Prev. 14, 2–19 (2005).
Cheng, I. et al. Haplotype-based association studies of IGFBP1 and IGFBP3 with prostate and breast cancer risk: the multiethnic cohort. Cancer Epidemiol. Biomarkers Prev. 15, 1993–1997 (2006).
Pollak, M. N., Schernhammer, E. S. & Hankinson, S. E. Insulin-like growth factors and neoplasia. Nature Rev. Cancer 4, 505–518 (2004).
Mendez, P., Azcoitia, I. & Garcia-Segura, L. M. Interdependence of oestrogen and insulin-like growth factor-I in the brain: potential for analysing neuroprotective mechanisms. J. Endocrinol. 185, 11–17 (2005).
Sharp, L., Cardy, A. H., Cotton, S. C. & Little, J. CYP17 gene polymorphisms: prevalence and associations with hormone levels and related factors. a HuGE review. Am. J. Epidemiol. 160, 729–740 (2004).
Power, S. G. et al. Molecular analyses of a human sex hormone-binding globulin variant: evidence for an additional carbohydrate chain. J. Clin. Endocrinol. Metab. 75, 1066–1070 (1992).
Cousin, P., Dechaud, H., Grenot, C., Lejeune, H. & Pugeat, M. Human variant sex hormone-binding globulin (SHBG) with an additional carbohydrate chain has a reduced clearance rate in rabbit. J. Clin. Endocrinol. Metab. 83, 235–240 (1998).
Sissung, T. M., Price, D. K., Sparreboom, A. & Figg, W. D. Pharmacogenetics and regulation of human cytochrome P450 1B1: implications in hormone-mediated tumor metabolism and a novel target for therapeutic intervention. Mol. Cancer Res. 4, 135–150 (2006).
Raftogianis, R., Creveling, C., Weinshilboum, R. & Weisz, J. Estrogen metabolism by conjugation. J. Natl Cancer Inst. Monogr., 113–124 (2000).
Kume, T. et al. Characterization of a novel variant (S145C/L311V) of 3α-hydroxysteroid/dihydrodiol dehydrogenase in human liver. Pharmacogenetics 9, 763–771 (1999).
Penning, T. M. et al. Human 3α-hydroxysteroid dehydrogenase isoforms (AKR1C1-AKR1C4) of the aldo-keto reductase superfamily: functional plasticity and tissue distribution reveals roles in the inactivation and formation of male and female sex hormones. Biochem. J. 351, 67–77 (2000).
Yager, J. D. & Davidson, N. E. Estrogen carcinogenesis in breast cancer. N. Engl. J. Med. 354, 270–282 (2006).
Planas-Silva, M. D., Shang, Y., Donaher, J. L., Brown, M. & Weinberg, R. A. AIB1 enhances estrogen-dependent induction of cyclin D1 expression. Cancer Res. 61, 3858–3862 (2001).
Singh, R. R. & Kumar, R. Steroid hormone receptor signaling in tumorigenesis. J. Cell Biochem. 96, 490–505 (2005).
Hayashi, Y. et al. Polymorphism of homopolymeric glutamines in coactivators for nuclear hormone receptors. Endocr. J. 46, 279–284 (1999).
Rosen, C. J. et al. Association between serum insulin growth factor-I (IGF-I) and a simple sequence repeat in IGF-I gene: implications for genetic studies of bone mineral density. J. Clin. Endocrinol. Metab. 83, 2286–2290 (1998).
Rasmussen, S. K. et al. Studies of the variability of the genes encoding the insulin-like growth factor I receptor and its ligand in relation to type 2 diabetes mellitus. J. Clin. Endocrinol. Metab. 85, 1606–1610 (2000).
Almind, K., Inoue, G., Pedersen, O. & Kahn, C. R. A common amino acid polymorphism in insulin receptor substrate-1 causes impaired insulin signaling. Evidence from transfection studies. J. Clin. Invest. 97, 2569–2575 (1996).
Le Stunff, C. et al. Association analysis indicates that a variant GATA-binding site in the PIK3CB promoter is a cis-acting expression quantitative trait locus for this gene and attenuates insulin resistance in obese children. Diabetes 57, 494–502 (2008).
Canzian, F. et al. Genetic variation in the growth hormone synthesis pathway in relation to circulating insulin-like growth factor-I, insulin-like growth factor binding protein-3, and breast cancer risk: results from the European prospective investigation into cancer and nutrition study. Cancer Epidemiol. Biomarkers Prev. 14, 2316–2325 (2005).
Hasegawa, Y. et al. Identification of novel human GH-1 gene polymorphisms that are associated with growth hormone secretion and height. J. Clin. Endocrinol. Metab. 85, 1290–1295 (2000).
Adams, E. F., Symowski, H., Buchfelder, M. & Poyner, D. R. A polymorphism in the growth hormone (GH)-releasing hormone (GHRH) receptor gene is associated with elevated response to GHRH by human pituitary somatotrophinomas in vitro. Biochem. Biophys. Res. Commun. 275, 33–36 (2000).
Lunn, R. M. et al. XPD polymorphisms: effects on DNA repair proficiency. Carcinogenesis 21, 551–555 (2000).
Savas, S. & Ozcelik, H. Phosphorylation states of cell cycle and DNA repair proteins can be altered by the nsSNPs. BMC Cancer 5, 107 (2005).
Acknowledgements
National Cancer Institute Grants R01 CA97396, R01 CA128931 and P50 CA116201
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Related links
Related links
DATABASES
National Cancer Institute
National Cancer Institute Drug Dictionary
FURTHER INFORMATION
Glossary
- Familial aggregation
-
A tendency for a trait to occur or cluster in multiple family members more often than would be expected by chance.
- Genetic association studies
-
Tests whether a polymorphism in a candidate gene explains the inter-individual variation in a trait under study.
- Haplotype blocks
-
Configurations of alleles within regions on the chromosome that tend to be inherited together (that is, are in high linkage disequilibrium). Little genetic variability is observed within this region among individuals in a population.
- Linkage analysis
-
The process of determining the approximate chromosomal location of a gene associated with the trait being studied by looking for evidence of co-segregation with other marker genes whose locations are already known.
- Linkage disequilibrium
-
Non-random association of alleles at two or more loci on chromosomes in a population, beyond that expected by chance.
- Parent–offspring and twin studies
-
Investigates the proportion of variation in the trait under study that is explained by unmeasured additive genetic factors among respective relative pairs.
- Segregation analysis
-
Tests hypotheses about whether the existence of major genes account for the observed pattern of familial aggregation of the trait and provides evidence for the mode of inheritance of the genes using pedigree data. Statistical models compare Mendelian inheritance patterns of a trait to a model in which there are no restrictions on mode of inheritance or other model parameters.
- Stroma
-
The supportive framework of a biological tissue with an extensive extracellular matrix that serves to support cells, separate tissues and regulate intercellular communication.
- Tag single nucleotide polymorphisms
-
(tagSNPs). A reduced set of single nucleotide polymorphisms that identify or 'tag' other SNPs with which it is in high linkage disequilibrium.
Rights and permissions
About this article
Cite this article
Kelemen, L., Sellers, T. & Vachon, C. Can genes for mammographic density inform cancer aetiology?. Nat Rev Cancer 8, 812–823 (2008). https://doi.org/10.1038/nrc2466
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc2466
This article is cited by
-
Modulation of innate immune response to viruses including SARS-CoV-2 by progesterone
Signal Transduction and Targeted Therapy (2022)
-
Relationships of physical and breast cancer phenotypes with three single-nucleotide polymorphisms (rs2046210, rs3757318, and rs3803662) associated with breast cancer risk in Japanese women
Breast Cancer (2021)
-
Alcohol intake from early adulthood to midlife and mammographic density
Cancer Causes & Control (2016)
-
Associations between breast density and a panel of single nucleotide polymorphisms linked to breast cancer risk: a cohort study with digital mammography
BMC Cancer (2015)
-
Leukocyte telomere length and its association with mammographic density and proliferative diagnosis among women undergoing diagnostic image-guided breast biopsy
BMC Cancer (2015)