Key Points
-
Alternative splicing of a pre-mRNA enables the production of multiple functionally distinct proteins from a single gene. This mechanism of gene regulation is widespread, with particular prevalence in the immune system.
-
Sequences in exons and introns can influence the pattern of splicing of a given gene. Therefore, translationally silent mutations can alter protein expression by changing the splicing of a pre-mRNA, with potential disease consequences.
-
The protein tyrosine kinases FYN, SYK and possibly LCK, are differentially spliced in T cells. At least two isoforms are predicted for each protein, with one isoform being more efficient at promoting T-cell signalling.
-
Cytokine signalling in T cells might also be influenced by alternative splicing. Functionally distinct isoforms have been detected for the cytokines interleukin-2 (IL-2), IL-4 and IL-6, the receptors for IL-4 and IL-7 and the cytokine-induced signalling molecules protein tyrosine kinase 2 (PYK2), myeloid differentiation primary-response gene 88 (MyD88) and IL-1 receptor-associated kinase 1 (IRAK1).
-
The expression of cell-surface molecules, such as CD44, intercellular adhesion molecule 1 (ICAM1), platelet/endothelial cell-adhesion molecule 1 (PECAM1), CD45 and cytotoxic T-lymphocyte antigen 4 (CTLA4) is also influenced by alternative splicing to generate either soluble forms of the molecules or molecules with altered protein–protein interactions. Splicing-dependent changes in the expression of these molecules have been shown to influence the threshold of T-cell activation.
-
Regulation of splicing in response to antigen stimulation has been observed or implied for CD44, CD45, FYN, CTLA4, PECAM1 and MyD88. The mechanisms by which activation-induced splicing regulation occurs are beginning to be understood for the CD44 and CD45 genes, and recent data indicate that some of these alternative splicing events might be regulated by overlapping pathways.
Abstract
Alternative splicing is widely recognized to be a ubiquitous and crucial mechanism for generating protein diversity and regulating protein expression. Numerous immunologically relevant genes have been found to undergo alternative splicing; however, there has been little effort to develop a coherent picture of how alternative splicing might be used as a general mechanism to regulate the function of the immune system. In this review, I summarize the mechanisms by which splicing is controlled in T cells, and discuss the role of alternative splicing and alternative isoform expression in the regulation of T-cell activation and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Modrek, B., Resch, A., Grasso, C. & Lee, C. Genome-wide detection of alternative splicing in expressed sequences of human genes. Nucleic Acids Res. 29, 2850–2859 (2001). This study demonstrates the extent of alternative splicing in humans, particularly in the immune and nervous systems.
Johnson, J. M. et al. Genome-wide survey of human alternative pre-mRNA splicing with exon junction microarrays. Science 302, 2141–2144 (2003).
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Smith, C. W. J. & Valcarcel, J. Alternative pre-mRNA splicing: the logic of combinatorial control. Trends Biochem. Sci. 25, 381–388 (2000).
Black, D. L. & Grabowski, P. J. Alternative pre-mRNA splicing and neuronal function. Prog. Mol. Subcell. Biol. 31, 187–216 (2003).
Wu, J. Y., Tang, H. & Havlioglu, N. Alternative pre-mRNA splicing and regulation of programmed cell death. Prog. Mol. Subcell. Biol. 31, 153–185 (2003).
Black, D. Protein diversity from alternative splicing: a challenge for bioinformatics and post-genome biology. Cell 103, 367–370 (2000).
Graveley, B. R. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17, 100–107 (2001).
Wollerton, M. C., Gooding, C., Wagner, E. J., Garcia-Blanco, M. A. & Smith, C. W. Autoregulation of polypyrimidine tract binding protein by alternative splicing leading to nonsense-mediated decay. Mol. Cell 13, 91–100 (2004).
Ishitani, A. & Geraghty, D. E. Alternative splicing of HLA-G transcripts yields proteins with primary structures resembling both class I and class II antigens. Proc. Natl Acad. Sci. USA 89, 3947–3951 (1992).
Yan, G., Shi, L. & Faustman, D. Novel splicing of the human MHC-encoded peptide transporter confers unique properties. J. Immunol. 162, 852–859 (1999).
Astier, A. et al. Human epidermal Langerhans cells secrete a soluble receptor for IgG (FcγRII/CD32) that inhibits the binding of immune complexes to FcγR+ cells. J. Immunol. 152, 201–212 (1994).
Atkinson, T. P., Hall, C. G., Goldsmith, J. & Kirkham, P. M. Splice variant in TCRζ links T cell receptor signaling to a G-protein-related signaling pathway. Biochem. Biophys. Res. Commun. 310, 761–766 (2003).
Tsytsikov, V. N., Yurovsky, V. V., Atamas, S. P., Alms, W. J. & White, B. Identification and characterization of two alternative splice variants of human interleukin-2. J. Biol. Chem. 271, 23055–23060 (1996).
Alms, W. J., Atamas, S. P., Yurovsky, V. V. & White, B. Generation of a variant of human interleukin-4 by alternative splicing. Mol. Immunol. 33, 361–370 (1996).
Trowbridge, I. S. & Thomas, M. L. CD45: An emerging role as a protein tyrosine phosphatase required for lymphocyte activation and development. Annu. Rev. Immunol. 12, 85–116 (1994).
Oaks, M. K. et al. A native soluble form of CTLA-4. Cell. Immunol. 201, 144–153 (2000).
Arch, R. et al. Participation in normal immune responses of a metastasis-inducing splice variant of CD44. Science 257, 682–685 (1992).
Rothrock, C., Cannon, B., Hahm, B. & Lynch, K. W. A conserved signal-responsive sequence mediates activation-induced alternative splicing of CD45. Mol. Cell 12, 1317–1324 (2003). Identification of the important sequences in the CD45 variable exons that mediate their antigen-induced repression. Evidence is also provided for the existence of common splicing-regulatory pathways in T cells.
Cooke, M. P. & Perlmutter, R. M. Expression of a novel form of the fyn proto-oncogene in hematopoietic cells. New Biol. 1, 66–74 (1989).
Davidson, D., Viallet, J. & Veillette, A. Unique catalytic properties dictate the enhanced function of p59fynT, the hemopoietic cell-specific isoform of the Fyn tyrosine protein kinase, in T cells. Mol. Cell Biol. 14, 4554–4564 (1994). This paper provides a detailed analysis of the functional differences between the FYN isoforms.
Davidson, D., Chow, L. M., Fournel, M. & Veillette, A. Differential regulation of T cell antigen responsiveness by isoforms of the src-related tyrosine protein kinase p59fyn. J. Exp. Med. 175, 1483–1492 (1992).
Weil, R. et al. Altered expression of tyrosine kinases of the Src and Syk families in human T-cell leukemia virus type 1-infected T-cell lines. J. Virol. 73, 3709–3717 (1999).
Germani, A., Malherbe, S. & Rouer, E. The exon 7-spliced Lck isoform in T lymphocytes: a potential regulator of p56lck signaling pathways. Biochem. Biophys. Res. Commun. 301, 680–685 (2003).
Rowley, R. B., Bolen, J. B. & Fargnoli, J. Molecular cloning of rodent p72Syk. Evidence of alternative mRNA splicing. J. Biol. Chem. 270, 12659–12664 (1995).
Yagi, S., Suzuki, K., Hasegawa, A., Okumura, K. & Ra, C. Cloning of the cDNA for the deleted Syk kinase homologous to ZAP-70 from human basophilic leukemia cell line (KU812). Biochem. Biophys. Res. Commun. 200, 28–34 (1994).
Latour, S., Zhang, J., Siraganian, R. P. & Veillette, A. A unique insert in the linker domain of Syk is necessary for its function in immunoreceptor signalling. EMBO J. 17, 2584–2595 (1998). The authors identify the molecular basis of the distinct functions of SYK and SYKB. This work also has implications for the functional difference between SYK and ZAP70 (ζ-chain-associated protein kinase of 70 kDa).
King, P. D. et al. Novel isoforms of murine intercellular adhesion molecule-1 generated by alternative RNA splicing. J. Immunol. 154, 6080–6093 (1995).
Wang, Y. & Sheibani, N. Expression pattern of alternatively spliced PECAM-1 isoforms in hematopoietic cells and platelets. J. Cell. Biochem. 87, 424–438 (2002).
Robledo, O. et al. ICAM-1 isoforms: specific activity and sensitivity to cleavage by leukocyte elastase and cathepsin G. Eur. J. Immunol. 33, 1351–1360 (2003).
Wang, Y., Su, X., Sorenson, C. M. & Sheibani, N. Modulation of PECAM-1 expression and alternative splicing during differentiation and activation of hematopoietic cells. J. Cell. Biochem. 88, 1012–1024 (2003).
Fox, S. B. et al. Normal human tissues, in addition to some tumors, express multiple different CD44 isoforms. Cancer Res. 54, 4539–4546 (1994).
Ponta, H., Sherman, L. & Herrlich, P. A. CD44: from adhesion molecules to signalling regulators. Nature Rev. Mol. Cell Biol. 4, 33–45 (2003).
Gunthert, U. et al. A new variant of glycoprotein CD44 confers metastatic potential to rat carcinoma cells. Cell 65, 13–24 (1991).
Eisterer, W. et al. CD44 isoforms are differentially regulated in plasma cell dyscrasias and CD44v9 represents a new independent prognostic parameter in multiple myeloma. Leuk. Res. 25, 1051–1057 (2001).
Bartolazzi, A. et al. Regulation of growth and dissemination of a human lymphoma by CD44 splice variants. J. Cell Sci. 108, 1723–1733 (1995).
Konig, H., Ponta, H. & Herrlich, P. Coupling of signal transduction to alternative pre-mRNA splicing by a composite splice regulator. EMBO J. 17, 2904–2913 (1998).
Atamas, S. P. Alternative splice variants of cytokines: making a list. Life Sci. 61, 1105–1112 (1997).
Heaney, M. L. & Golde, D. W. Soluble cytokine receptors. Blood 87, 847–857 (1996).
Rincon, M., Anguita, J., Nakamura, T., Fikrig, E. & Flavell, R. A. Interleukin (IL)-6 directs the differentiation of IL-4-producing CD4+ T cells. J. Exp. Med. 185, 461–469 (1997).
Xu, G. Y. et al. Solution structure of recombinant human interleukin-6. J. Mol. Biol. 268, 468–481 (1997).
Somers, W., Stahl, M. & Seehra, J. S. 1.9 A crystal structure of interleukin 6: implications for a novel mode of receptor dimerization and signaling. EMBO J. 16, 989–997 (1997).
Boulay, J. L. & Paul, W. E. The interleukin-4-related lymphokines and their binding to hematopoietin receptors. J. Biol. Chem. 267, 20525–20528 (1992).
Bihl, M. P. et al. Identification of a novel IL-6 isoform binding to the endogenous IL-6 receptor. Am. J. Respir. Cell. Mol. Biol. 27, 48–56 (2002). An excellent study of the structure–function relationship between IL-6 isoforms.
Paonessa, G. et al. Two distinct and independent sites on IL-6 trigger gp130 dimer formation and signalling. EMBO J. 14, 1942–1951 (1995).
Zav'yalov, V. P. et al. Molecular model of an alternative splice variant of human IL-4, IL-4Δ2, a naturally occurring inhibitor of IL-4-stimulated T cell proliferation. Immunol. Lett. 58, 149–152 (1997).
Denesyuk, A. I., Zav'yalov, V. P., Denessiouk, K. A. & Korpela, T. Molecular models of two competitive inhibitors, IL-2Δ2 and IL-2Δ3, generated by alternative splicing of human interleukin-2. Immunol. Lett. 60, 61–66 (1998).
Kruse, S., Forster, J., Kuehr, J. & Deichmann, K. A. Characterization of the membrane-bound and a soluble form of human IL-4 receptor α produced by alternative splicing. Int. Immunol. 11, 1965–1970 (1999).
Goodwin, R. G. et al. Cloning of the human and murine interleukin-7 receptors: demonstration of a soluble form and homology to a new receptor superfamily. Cell 60, 941–951 (1990).
Miyazaki, T. et al. Pyk2 is a downstream mediator of the IL-2 receptor-coupled Jak signaling pathway. Genes Dev. 12, 770–775 (1998).
Benbernou, N., Muegge, K. & Durum, S. K. Interleukin (IL)-7 induces rapid activation of Pyk2, which is bound to Janus kinase 1 and IL-7Rα. J. Biol. Chem. 275, 7060–7065 (2000).
Hazeki, K. et al. Toll-like receptor-mediated tyrosine phosphorylation of paxillin via MyD88-dependent and -independent pathways. Eur. J. Immunol. 33, 740–747 (2003).
Dikic, I. & Schlessinger, J. Identification of a new Pyk2 isoform implicated in chemokine and antigen receptor signaling. J. Biol. Chem. 273, 14301–14308 (1998). Identification and functional characterization of a variant isoform of PYK2.
Cao, Z., Henzel, W. J. & Gao, X. IRAK: a kinase associated with the interleukin-1 receptor. Science 271, 1128–1131 (1996).
Jensen, L. E. & Whitehead, A. S. IRAK1b, a novel alternative splice variant of interleukin-1 receptor-associated kinase (IRAK), mediates interleukin-1 signaling and has prolonged stability. J. Biol. Chem. 276, 29037–29044 (2001).
Wesche, H., Henzel, W. J., Shillinglaw, W., Li, S. & Cao, Z. MyD88: an adapter that recruits IRAK to the IL-1 receptor complex. Immunity 7, 837–847 (1997).
Burns, K. et al. Inhibition of interleukin 1 receptor/Toll-like receptor signaling through the alternatively spliced, short form of MyD88 is due to its failure to recruit IRAK-4. J. Exp. Med. 197, 263–268 (2003). This paper clearly shows the molecular basis for the functional differences between MyD88 isoforms.
Janssens, S., Burns, K., Tschopp, J. & Beyaert, R. Regulation of interleukin-1- and lipopolysaccharide-induced NF-κB activation by alternative splicing of MyD88. Curr. Biol. 12, 467–471 (2002).
Lynch, K. W. & Weiss, A. A model system for the activation-induced alternative-splicing of CD45 implicates protein kinase C and Ras. Mol. Cell Biol. 20, 70–80 (2000).
Xu, Z. & Weiss, A. Negative regulation of CD45 by differential homodimerization of the alternatively spliced isoforms. Nature Immunol. 3, 764–771 (2002). This study demonstrates the functional differences between CD45 isoforms due to the inefficiency of dimerization of the large CD45 isoforms.
Majeti, R., Bilwes, A. M., Noel, J. P., Hunter, T. & Weiss, A. Dimerization-induced inhibition of receptor protein tyrosine phosphatase function through an inhibitory wedge. Science 279, 88–91 (1998).
Hermiston, M. L., Xu, Z., Majeti, R. & Weiss, A. Reciprocal regulation of lymphocyte activation by tyrosine kinases and phosphatases. J. Clin. Invest. 109, 9–14 (2002).
Jacobsen, M. et al. A point mutation in PTPRC is associated with the development of multiple sclerosis. Nature Genet. 26, 495–499 (2000).
Chu, D. H. et al. The Syk protein tyrosine kinase can function independently of CD45 and Lck in T cell antigen receptor signaling. EMBO J. 15, 6251–6261 (1996).
van der Merwe, P. A., Bodian, D. L., Daenke, S., Linsley, P. & Davis, S. J. CD80 (B7-1) binds both CD28 and CTLA-4 with a low affinity and very fast kinetics. J. Exp. Med. 185, 393–403 (1997).
Krummel, M. F. & Allison, J. P. CD28 and CTLA-4 have opposing effects on the response of T cells to stimulation. J. Exp. Med. 182, 459–465 (1995).
Perkins, D. et al. Regulation of CTLA-4 expression during T cell activation. J. Immunol. 156, 4154–4159 (1996).
Magistrelli, G. et al. A soluble form of CTLA-4 generated by alternative splicing is expressed by nonstimulated human T cells. Eur. J. Immunol. 29, 3596–3602 (1999).
Faustino, N. A. & Cooper, T. A. Pre-mRNA splicing and human disease. Genes Dev. 17, 419–437 (2003).
Cartegni, L., Chew, S. L. & Krainer, A. R. Listening to silence and understanding nonsense: exonic mutations that affect splicing. Nature Rev. Genet. 3, 285–298 (2002).
Pagani, F. & Baralle, F. E. Genomic variants in exons and introns: identifying the splicing spoilers. Nature Rev. Genet. 5, 389–396 (2004).
Oaks, M. K. & Hallett, K. M. Cutting edge: a soluble form of CTLA-4 in patients with autoimmune thyroid disease. J. Immunol. 164, 5015–5018 (2000).
Ueda, H. et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature 423, 506–511 (2003).
Jones, S. A., Horiuchi, S., Topley, N., Yamamoto, N. & Fuller, G. M. The soluble interleukin-6 receptor: mechanisms of production and implications in disease. FASEB J. 15, 43–58 (2001).
Korte, A. et al. Expression analysis and characterization of alternatively spliced transcripts of human IL-7Rα chain encoding two truncated receptor proteins in relapsed childhood all. Cytokine 12, 1597–1608 (2000).
Kotsimbos, A. T. & Hamid, Q. IL-5 and IL-5 receptor in asthma. Mem. Inst. Oswaldo Cruz 92, 75–91 (1997).
Sakkas, L. I., Tourtellotte, C., Berney, S., Myers, A. R. & Platsoucas, C. D. Increased levels of alternatively spliced interleukin 4 (IL-4Δ2) transcripts in peripheral blood mononuclear cells from patients with systemic sclerosis. Clin. Diagn. Lab. Immunol. 6, 660–664 (1999).
Matter, N. et al. Heterogeneous ribonucleoprotein A1 is part of an exon-specific splice-silencing complex controlled by oncogenic signaling pathways. J. Biol. Chem. 275, 35353–35360 (2000).
Weg-Remers, S., Ponta, H., Herrlich, P. & Konig, H. Regulation of alternative pre-mRNA splicing by the ERK MAP-kinase pathway. EMBO J. 20, 4194–4203 (2001).
Matter, N., Herrlich, P. & Konig, H. Signal-dependent regulation of splicing via phosphorylation of Sam68. Nature 420, 691–695 (2002). This is one of the first papers to provide a clear molecular link between activation of the mitogen-activated protein kinase pathway and modulation of a specific splicing regulatory event.
Stickeler, E. et al. The RNA binding protein YB-1 binds A/C-rich exon enhancers and stimulates splicing of the CD44 alternative exon v4. EMBO J. 20, 3821–3830 (2001).
Honig, A., Auboeuf, D., Parker, M. M., O'Malley, B. W. & Berget, S. M. Regulation of alternative splicing by the ATP-dependent DEAD-box RNA helicase p72. Mol. Cell Biol. 22, 5698–5707 (2002).
Lynch, K. W. & Weiss, A. A CD45 polymorphism associated with multiple sclerosis disrupts an exonic splicing silencer. J. Biol. Chem. 276, 24341–24347 (2001).
Zilch, C. F. et al. A point mutation within CD45 exon A is the cause of variant CD45RA splicing in humans. Eur. J. Immunol. 28, 22–29 (1998).
Syken, J. Macian, F., Agarwal, S., Rao, A. & Munger, K. TID1, a mammalian homologue of the drosophila tumor suppressor lethal (2) tumorous imaginal discs, regulates activation-induced cell death in TH2 cells. Oncogene 22, 4636–4641 (2003).
Liu, C., Cheng, J. & Mountz, J. D. Differential expression of human Fas mRNA species upon peripheral blood mononuclear cell activation. Biochemistry J. 310, 957–963 (1995).
Nishimura, H., Washizu, J., Nakamura, N., Enomoto, A. & Yoshikai, Y. Translational efficiency is upregulated by alternative exon in murine IL-15 mRNA. J. Immunol. 160, 936–942 (1998).
Berget, S. M. Exon recognition in vertebrate splicing. J. Biol. Chem. 270, 2411–2414 (1995).
Acknowledgements
I thank B. Graveley, K. Hertel, C. Wuelfing and members of laboratory for critical comments on the manuscript.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Glossary
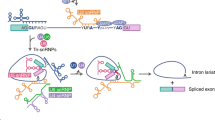
- SPLICING
-
The processing of pre-mRNA such that introns are removed and exons are joined directly to one another.
- CASSETTE EXONS
-
Exons that are included in only a percentage of the final mRNA transcripts that are derived from a given pre-mRNA.
- MUTUALLY EXCLUSIVE EXONS
-
Two or more exons in a single pre-mRNA that are never both included in the final mRNA transcript.
- SPLICE SITES
-
Conserved sequences at the exon–intron boundaries that direct the splicing machinery and determine the precise location of pre-mRNA cleavage and exon joining.
- SPLICEOSOME
-
A ribonucleoprotein complex that is involved in splicing of nuclear pre-mRNA. It is composed of five small nuclear ribonucleoproteins (snRNPs) and more than 50 non-snRNPs that recognize and assemble on exon–intron boundaries to catalyse intron processing of the pre-mRNA.
- SR PROTEIN
-
A group of highly conserved, serine (S)- and arginine (R)-rich splicing-regulatory proteins. In most experimental systems, binding of an SR protein to an exon leads to exon inclusion.
- HETEROGENEOUS NUCLEAR RIBONUCLEOPARTICLE PROTEIN
-
(HnRNP protein). A class of diverse RNA-binding proteins that associate with nascent pre-mRNA and can influence splicing. Several experimental systems have shown that binding of an hnRNP protein to an exon leads to exon skipping.
- EXONIC SPLICING SILENCERS
-
(ESSs). Sequences in an exon that promote exon skipping.
- EXONIC SPLICING ENHANCERS
-
(ESEs). Sequences in an exon that promote exon recognition and inclusion.
- INTRONIC SPLICING SILENCERS
-
(ISSs). Sequences in an intron that promote skipping of a flanking exon.
- INTRONIC SPLICING ENHANCERS
-
(ISEs). Sequences in an intron that promote recognition and inclusion of a flanking exon.
- IMMUNOLOGICAL SYNAPSE
-
A stable region of contact between a T cell and an antigen-presenting cell that forms through cell–cell interaction of adhesion molecules. The mature immunological synapse contains two distinct, membrane domains: a central cluster of T-cell receptors, known as the central supramolecular activation cluster (cSMAC) and a surrounding ring of adhesion molecules known as the peripheral SMAC.
- FOUR-HELICAL BUNDLE
-
A structural motif in proteins in which four α-helices are packed together.
- DOMINANT-NEGATIVE INHIBITOR
-
A protein variant that inhibits the proper function of the co-expressed wild-type protein.
- MIXED LEUKOCYTE REACTION
-
A tissue-culture technique for testing T-cell reactivity. The proliferation of one population of T cells, induced by exposure to inactivated MHC-mismatched stimulator cells, is determined by measuring the incorporation of 3H-thymidine into the DNA of dividing cells.
- EXON SKIPPING
-
Exclusion of an exon from a pre-mRNA into the final mature mRNA transcript.
Rights and permissions
About this article
Cite this article
Lynch, K. Consequences of regulated pre-mRNA splicing in the immune system. Nat Rev Immunol 4, 931–940 (2004). https://doi.org/10.1038/nri1497
Issue Date:
DOI: https://doi.org/10.1038/nri1497
This article is cited by
-
Tnpo3 controls splicing of the pre-mRNA encoding the canonical TCR α chain of iNKT cells
Nature Communications (2023)
-
Genome-wide alternative splicing profile in the posterior kidney of brown trout (Salmo trutta) during proliferative kidney disease
BMC Genomics (2022)
-
Are CD45RO+ and CD45RA- genuine markers for bovine memory T cells?
Animal Diseases (2022)
-
Optimal CD8+ T cell effector function requires costimulation-induced RNA-binding proteins that reprogram the transcript isoform landscape
Nature Communications (2022)
-
Bacterial epigenetics opens door to novel frontier in Infection biology
The Nucleus (2021)