Abstract
Mediator is an essential, broadly used eukaryotic transcriptional coactivator. How and what Mediator communicates from activators to RNA polymerase II (RNAPII) remains an open question. Here we performed genome-wide location profiling of Saccharomyces cerevisiae Mediator subunits. Mediator is not found at core promoters but rather occupies the upstream activating sequence, upstream of the pre-initiation complex. In the absence of Kin28 (CDK7) kinase activity or in cells in which the RNAPII C-terminal domain is mutated to replace Ser5 with alanine, however, Mediator accumulates at core promoters together with RNAPII. We propose that Mediator is released quickly from promoters after phosphorylation of Ser5 by Kin28 (CDK7), which also allows for RNAPII to escape from the promoter.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Näär, A.M., Lemon, B.D. & Tjian, R. Transcriptional coactivator complexes. Annu. Rev. Biochem. 70, 475–501 (2001).
Poss, Z.C., Ebmeier, C.C. & Taatjes, D.J. The Mediator complex and transcription regulation. Crit. Rev. Biochem. Mol. Biol. 48, 575–608 (2013).
Ansari, S.A. & Morse, R.H. Mechanisms of Mediator complex action in transcriptional activation. Cell. Mol. Life Sci. 70, 2743–2756 (2013).
Carlsten, J.O., Zhu, X. & Gustafsson, C.M. The multitalented Mediator complex. Trends Biochem. Sci. 38, 531–537 (2013).
Myers, L.C. et al. The Med proteins of yeast and their function through the RNA polymerase II carboxy-terminal domain. Genes Dev. 12, 45–54 (1998).
Spahr, H. et al. Mediator influences Schizosaccharomyces pombe RNA polymerase II–dependent transcription in vitro. J. Biol. Chem. 278, 51301–51306 (2003).
Knuesel, M.T., Meyer, K.D., Bernecky, C. & Taatjes, D.J. The human CDK8 subcomplex is a molecular switch that controls Mediator coactivator function. Genes Dev. 23, 439–451 (2009).
Näär, A.M., Taatjes, D.J., Zhai, W., Nogales, E. & Tjian, R. Human CRSP interacts with RNA polymerase II CTD and adopts a specific CTD-bound conformation. Genes Dev. 16, 1339–1344 (2002).
Andrau, J.-C. et al. Genome-wide location of the coactivator mediator: Binding without activation and transient Cdk8 interaction on DNA. Mol. Cell 22, 179–192 (2006).
Zhu, X. et al. Genome-wide occupancy profile of mediator and the Srb8–11 module reveals interactions with coding regions. Mol. Cell 22, 169–178 (2006).
Conaway, R.C. & Conaway, J.W. The Mediator complex and transcription elongation. Biochim. Biophys. Acta 1829, 69–75 (2013).
Kuras, L. & Struhl, K. Binding of TBP to promoters in vivo is stimulated by activators and requires Pol II holoenzyme. Nature 399, 609–613 (1999).
Cantin, G.T., Stevens, J.L. & Berk, A.J. Activation domain–mediator interactions promote transcription preinitiation complex assembly on promoter DNA. Proc. Natl. Acad. Sci. USA 100, 12003–12008 (2003).
Bhaumik, S.R., Raha, T., Aiello, D.P. & Green, M.R. In vivo target of a transcriptional activator revealed by fluorescence resonance energy transfer. Genes Dev. 18, 333–343 (2004).
Wang, G. et al. Mediator requirement for both recruitment and postrecruitment steps in transcription initiation. Mol. Cell 17, 683–694 (2005).
Baek, H.J., Kang, Y.K. & Roeder, R.G. Human Mediator enhances basal transcription by facilitating recruitment of transcription factor IIB during preinitiation complex assembly. J. Biol. Chem. 281, 15172–15181 (2006).
He, Q., Battistella, L. & Morse, R.H. Mediator requirement downstream of chromatin remodeling during transcriptional activation of CHA1 in yeast. J. Biol. Chem. 283, 5276–5286 (2008).
Rana, R., Surapureddi, S., Kam, W., Ferguson, S. & Goldstein, J.A. Med25 is required for RNA polymerase II recruitment to specific promoters, thus regulating xenobiotic and lipid metabolism in human liver. Mol. Cell. Biol. 31, 466–481 (2011).
Lin, J.J. et al. Mediator coordinates PIC assembly with recruitment of CHD1. Genes Dev. 25, 2198–2209 (2011).
Chen, X.F. et al. Mediator and SAGA have distinct roles in Pol II preinitiation complex assembly and function. Cell Reports 2, 1061–1067 (2012).
Takahashi, H. et al. Human mediator subunit MED26 functions as a docking site for transcription elongation factors. Cell 146, 92–104 (2011).
Galbraith, M.D. et al. HIF1A employs CDK8-mediator to stimulate RNAPII elongation in response to hypoxia. Cell 153, 1327–1339 (2013).
Mukundan, B. & Ansari, A. Novel role for mediator complex subunit Srb5/Med18 in termination of transcription. J. Biol. Chem. 286, 37053–37057 (2011).
Mukundan, B. & Ansari, A. Srb5/Med18-mediated termination of transcription is dependent on gene looping. J. Biol. Chem. 288, 11384–11394 (2013).
Huang, Y. et al. Mediator complex regulates alternative mRNA processing via the MED23 subunit. Mol. Cell 45, 459–469 (2012).
Fan, X., Chou, D.M. & Struhl, K. Activator-specific recruitment of Mediator in vivo. Nat. Struct. Mol. Biol. 13, 117–120 (2006).
Fan, X. & Struhl, K. Where does mediator bind in vivo? PLoS ONE 4, e5029 (2009).
Ansari, S.A., He, Q. & Morse, R.H. Mediator complex association with constitutively transcribed genes in yeast. Proc. Natl. Acad. Sci. USA 106, 16734–16739 (2009).
Strässer, K. et al. TREX is a conserved complex coupling transcription with messenger RNA export. Nature 417, 304–308 (2002).
Teytelman, L., Thurtle, D.M., Rine, J. & van Oudenaarden, A. Highly expressed loci are vulnerable to misleading ChIP localization of multiple unrelated proteins. Proc. Natl. Acad. Sci. USA 110, 18602–18607 (2013).
Park, D., Lee, Y., Bhupindersingh, G. & Iyer, V.R. Widespread misinterpretable ChIP-seq bias in yeast. PLoS ONE 8, e83506 (2013).
Drouin, S. et al. DSIF and RNA polymerase II CTD phosphorylation coordinate the recruitment of Rpd3S to actively transcribed genes. PLoS Genet. 6, e1001173 (2010).
Soontorngun, N. et al. Genome-wide location analysis reveals an important overlap between the targets of the yeast transcriptional regulators Rds2 and Adr1. Biochem. Biophys. Res. Commun. 423, 632–637 (2012).
Liesen, T., Hollenberg, C.P. & Heinisch, J.J. ERA, a novel cis-acting element required for autoregulation and ethanol repression of PDC1 transcription in Saccharomyces cerevisiae. Mol. Microbiol. 21, 621–632 (1996).
Donner, A.J., Ebmeier, C.C., Taatjes, D.J. & Espinosa, J.M. CDK8 is a positive regulator of transcriptional elongation within the serum response network. Nat. Struct. Mol. Biol. 17, 194–201 (2010).
Loewith, R. & Hall, M.N. Target of rapamycin (TOR) in nutrient signaling and growth control. Genetics 189, 1177–1201 (2011).
Haruki, H., Nishikawa, J. & Laemmli, U.K. The anchor-away technique: rapid, conditional establishment of yeast mutant phenotypes. Mol. Cell 31, 925–932 (2008).
Nonet, M.L. & Young, R.A. Intragenic and extragenic suppressors of mutations in the heptapeptide repeat domain of Saccharomyces cerevisiae RNA Polymerase II. Genetics 123, 715–724 (1989).
Thompson, C.M., Koleske, A.J., Chao, D.M. & Young, R.A. A multisubunit complex associated with the RNA polymerase II CTD and TATA-binding protein in yeast. Cell 73, 1361–1375 (1993).
Søgaard, T.M. & Svejstrup, J.Q. Hyperphosphorylation of the C-terminal repeat domain of RNA polymerase II facilitates dissociation of its complex with mediator. J. Biol. Chem. 282, 14113–14120 (2007).
Liu, Y. et al. Two cyclin-dependent kinases promote RNA polymerase II transcription and formation of the scaffold complex. Mol. Cell. Biol. 24, 1721–1735 (2004).
Kim, Y.J., Bjorklund, S., Li, Y., Sayre, M.H. & Kornberg, R.D. A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell 77, 599–608 (1994).
Rhee, H.S. & Pugh, B.F. Genome-wide structure and organization of eukaryotic pre-initiation complexes. Nature 483, 295–301 (2012).
Bataille, A.R. et al. A universal RNA polymerase II CTD cycle is orchestrated by complex interplays between kinase, phosphatase, and isomerase enzymes along genes. Mol. Cell 45, 158–170 (2012).
Kim, H. et al. Gene-specific RNA polymerase II phosphorylation and the CTD code. Nat. Struct. Mol. Biol. 17, 1279–1286 (2010).
Tietjen, J.R. et al. Chemical-genomic dissection of the CTD code. Nat. Struct. Mol. Biol. 17, 1154–1161 (2010).
Guidi, B.W. et al. Mutual targeting of mediator and the TFIIH kinase Kin28. J. Biol. Chem. 279, 29114–29120 (2004).
Jeronimo, C., Bataille, A.R. & Robert, F. The writers, readers, and functions of the RNA polymerase II C-terminal domain code. Chem. Rev. 113, 8491–8522 (2013).
McCracken, S. et al. 5′-capping enzymes are targeted to pre-mRNA by binding to the phosphorylated carboxy-terminal domain of RNA polymerase II. Genes Dev. 11, 3306–3318 (1997).
Fong, N., Bird, G., Vigneron, M. & Bentley, D.L. A 10 residue motif at the C-terminus of the RNA pol II CTD is required for transcription, splicing and 3′ end processing. EMBO J. 22, 4274–4282 (2003).
Hsin, J.P., Sheth, A. & Manley, J.L. RNAP II CTD phosphorylated on threonine-4 is required for histone mRNA 3′ end processing. Science 334, 683–686 (2011).
Thompson, C.M. & Young, R.A. General requirement for RNA polymerase II holoenzymes in vivo. Proc. Natl. Acad. Sci. USA 92, 4587–4590 (1995).
Holstege, F.C.P. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998).
Takagi, Y. & Kornberg, R.D. Mediator as a general transcription factor. J. Biol. Chem. 281, 80–89 (2006).
Kagey, M.H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Whyte, W.A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Svejstrup, J.Q. et al. Evidence for a mediator cycle at the initiation of transcription. Proc. Natl. Acad. Sci. USA 94, 6075–6078 (1997).
Robinson, P.J., Bushnell, D.A., Trnka, M.J., Burlingame, A.L. & Kornberg, R.D. Structure of the mediator head module bound to the carboxy-terminal domain of RNA polymerase II. Proc. Natl. Acad. Sci. USA 109, 17931–17935 (2012).
Koleske, A.J. & Young, R.A. An RNA polymerase II holoenzyme responsive to activators. Nature 368, 466–469 (1994).
Liao, S.M. et al. A kinase-cyclin pair in the RNA polymerase II holoenzyme. Nature 374, 193–196 (1995).
Ren, B. et al. Genome-wide location and function of DNA binding proteins. Science 290, 2306–2309 (2000).
Guillemette, B. et al. Variant histone H2A.Z is globally localized to the promoters of inactive yeast genes and regulates nucleosome positioning. PLoS Biol. 3, e384 (2005).
Szilard, R.K. et al. Systematic identification of fragile sites via genome-wide location analysis of γ-H2AX. Nat. Struct. Mol. Biol. 17, 299–305 (2010).
Rufiange, A., Jacques, P.-É., Bhat, W., Robert, F. & Nourani, A. Genome-wide replication-independent histone H3 exchange occurs predominantly at promoters and implicates H3 K56 acetylation and Asf1. Mol. Cell 27, 393–405 (2007).
Hardy, S. et al. The euchromatic and heterochromatic landscapes are shaped by antagonizing effects of transcription on H2A.Z deposition. PLoS Genet. 5, e1000687 (2009).
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009).
Harbison, C.T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Acknowledgements
We thank K. Struhl (Harvard Medical School) for discussing unpublished data and P. Collin (Robert laboratory) for assistance with the CTD mutant constructs, as well as R. Young (Whitehead Institute for Biomedical Research), S. Hahn (Fred Hutchinson Cancer Research Center) and L. Myers (Dartmouth-Hitchcock Medical Center) for sharing strains and antibodies. We are also grateful to C. Kaplan and N. Francis for their critical reading of the manuscript. This work was funded by a Canadian Institutes of Health Research grant (MOP-82891) to F.R.
Author information
Authors and Affiliations
Contributions
C.J. and F.R. designed the project and wrote the manuscript. C.J. performed most of the experiments. F.R. performed most of the analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Enrichment in highly transcribed coding regions is often observed in ChIP experiments and has to be controlled for.
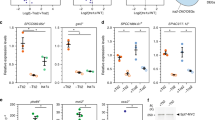
(a) Enrichment of various Mediator subunits at the highly transcribed PMA1 gene as determined by ChIP-chip using a panel of rabbit polyclonal antibodies (green), as well as using a control IgG antibody (black). All ChIPs were hybridized (and therefore normalized) against input DNA. (b) A scatter plot of the enrichment over coding regions in a control IgG ChIP and an Rpb3 ChIP. (c) Enrichment of various Mediator subunits as determined by ChIP-chip using epitope (Myc)-tagged Mediator subunits (green). Except for the last trace, all ChIPs were hybridized (and therefore normalized) against input DNA as in a. The last profile, however, shows an experiment were the MED15-Myc ChIP sample was hybridized (and therefore normalized) against a control ChIP sample performed using the same anti-Myc antibody but from an isogenic non-tagged strain. (d) TFIIB enrichment as determined by ChIP-chip using a rabbit polyclonal anti-TFIIB antibody (purple) as well as using a control IgG antibody (black) as in a. The middle panel shows TFIIB enrichment after subtracting the signal from the control (black) experiment. Overall, these experiments show that ChIP experiments tend to enrich highly transcribed coding regions regardless of the antibody used. It also shows that most of this systematic error can be removed either by normalizing against controls ChIPs performed in a non-tagged strain (in the case where a tagged protein is tested) or against a control ChIP performed using IgG (when a polyclonal antibody is used). Because each antibody varies with regards to the amount of noise it generates, we favor, when possible, using the "tagged versus non-tagged" strategy.
Supplementary Figure 2 A complement to Figure 1.
(a) A heat map representation of the data from Figure 1a. Genes were sorted by decreasing Rpb3 occupancy and the data were aligned on the TSS. (b) The average TFIIB (purple), RNAPII (Rpb3, red) and Mediator (Gal11, green) occupancies are shown around the TSS of RP genes. (c) Same as in b but for a group of control genes chosen based on the fact that they exhibit similar transcription rates as the RP genes from b. (d) A heat map representation of Mediator (Gal11) occupancy over the same genes as in b and c.
Supplementary Figure 3 A complement to Figure 3.
(a-c) Average Mediator (Gal11, green) occupancy in wild type (WT, solid traces) and the indicated CTD kinase mutants (dotted and dashed traces) around the TSS of all genes with an Rpb3 average ORF occupancy >1 in WT cells. TFIIB (purple) from WT cells is shown as a place-holder for core promoters. Data are shown for kin28-as mutant treated with NAPP1 (n=266) (a), ctk1Δ and bur2Δ mutants (n=299) (b) and srb10Δ mutant (n=295) (c). (d) Average Mediator (Gal11, green) and RNAPII (Rpb3, red) occupancy in wild type (WT, solid traces) and kin28-as mutant (kin28-as, dotted traces) cells, both treated with NAPP1, around the TSS for groups of genes with different RNAPII occupancies. The genes were grouped based on the average Rpb3 enrichment (log2 ratio) over their ORF (>1.5, Very high n=142; 1-1.5, High n=135; 0.5-1, Medium n=337; 0-0.5, Low n=827; <0, Very low n=1764).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Table 1 (PDF 998 kb)
Rights and permissions
About this article
Cite this article
Jeronimo, C., Robert, F. Kin28 regulates the transient association of Mediator with core promoters. Nat Struct Mol Biol 21, 449–455 (2014). https://doi.org/10.1038/nsmb.2810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2810
This article is cited by
-
PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
Nature Communications (2023)
-
A novel tRNA-derived fragment AS-tDR-007333 promotes the malignancy of NSCLC via the HSPB1/MED29 and ELK4/MED29 axes
Journal of Hematology & Oncology (2022)
-
The Mediator complex as a master regulator of transcription by RNA polymerase II
Nature Reviews Molecular Cell Biology (2022)
-
CDK7 inhibitors as anticancer drugs
Cancer and Metastasis Reviews (2020)
-
SAGA DUBm-mediated surveillance regulates prompt export of stress-inducible transcripts for proteostasis
Nature Communications (2019)