Abstract
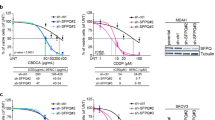
Pancreatic ductal adenocarcinoma (PDAC) is an aggressive and incurable disease. Poor prognosis is due to multiple reasons, including acquisition of resistance to gemcitabine, the first-line chemotherapeutic approach. Thus, there is a strong need for novel therapies, targeting more directly the molecular aberrations of this disease. We found that chronic exposure of PDAC cells to gemcitabine selected a subpopulation of cells that are drug-resistant (DR-PDAC cells). Importantly, alternative splicing (AS) of the pyruvate kinase gene (PKM) was differentially modulated in DR-PDAC cells, resulting in promotion of the cancer-related PKM2 isoform, whose high expression also correlated with shorter recurrence-free survival in PDAC patients. Switching PKM splicing by antisense oligonucleotides to favor the alternative PKM1 variant rescued sensitivity of DR-PDAC cells to gemcitabine and cisplatin, suggesting that PKM2 expression is required to withstand drug-induced genotoxic stress. Mechanistically, upregulation of the polypyrimidine-tract binding protein (PTBP1), a key modulator of PKM splicing, correlated with PKM2 expression in DR-PDAC cell lines. PTBP1 was recruited more efficiently to PKM pre-mRNA in DR- than in parental PDAC cells. Accordingly, knockdown of PTBP1 in DR-PDAC cells reduced its recruitment to the PKM pre-mRNA, promoted splicing of the PKM1 variant and abolished drug resistance. Thus, chronic exposure to gemcitabine leads to upregulation of PTBP1 and modulation of PKM AS in PDAC cells, conferring resistance to the drug. These findings point to PKM2 and PTBP1 as new potential therapeutic targets to improve response of PDAC to chemotherapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Stathis A, Moore MJ . Advanced pancreatic carcinoma: current treatment and future challenges. Nat Rev Clin Oncol 2010; 7: 163–1672.
Michl P, Gress TM . Current concepts and novel targets in advanced pancreatic cancer. Gut 2013; 62: 317–326.
Hidalgo M . Pancreatic cancer. N Engl J Med 2010; 362: 1605–1617.
Tang SC, Chen YC . Novel therapeutic targets for pancreatic cancer. World J Gastroenterol 2014; 20: 10825–10844.
David CJ, Manley JL . Alternative pre-mRNA splicing regulation in cancer: pathways and programs unhinged. Genes Dev 2010; 24: 2343–2364.
Zhang J, Manley JL . Misregulation of pre-mRNA alternative splicing in cancer. Cancer Discov 2013; 3: 1228–1237.
Bonomi S, Gallo S, Catillo M, Pignataro D, Biamonti G, Ghigna C . Oncogenic alternative splicing switches: role in cancer progression and prospects for therapy. Int J Cell Biol 2013; 2013: 962038.
Sette C, Ladomery M, Ghigna C . Alternative splicing: role in cancer development and progression. Int J Cell Biol 2013; 2013: 421606.
Cooper TA, Wan L, Dreyfuss G . RNA and disease. Cell 2009; 136: 777–793.
Ghigna C, Giordano S, Shen H, Benvenuto F, Castiglioni F, Comoglio PM et al. Cell motility is controlled by SF2/ASF through alternative splicing of the Ron protooncogene. Mol Cell. 2005; 20: 881–890.
Karni R, de Stanchina E, Lowe SW, Sinha R, Mu D, Krainer AR . The gene encoding the splicing factor SF2/ASF is a proto-oncogene. Nat Struct Mol Biol 2007; 14: 185–193.
Venables JP, Klinck R, Koh C, Gervais-Bird J, Bramard A, Inkel L et al. Cancer-associated regulation of alternative splicing. Nat Struct Mol Biol 2009; 16: 670–676.
Paronetto MP, Cappellari M, Busà R, Pedrotti S, Vitali R, Comstock C et al. Alternative splicing of the cyclin D1 proto-oncogene is regulated by the RNA-binding protein Sam68. Cancer Res 2010; 70: 229–239.
Zhou X, Li X, Cheng Y, Wu W, Xie Z, Xi Q et al. BCLAF1 and its splicing regulator SRSF10 regulate the tumorigenic potential of colon cancer cells. Nat Commun 2014; 5: 4581.
Dutertre M, Sanchez G, Barbier J, Corcos L, Auboeuf D . The emerging role of pre-messenger RNA splicing in stress responses: sending alternative messages and silent messengers. RNA Biol 2011; 8: 740–747.
Busà R, Geremia R, Sette C . Genotoxic stress causes the accumulation of the splicing regulator Sam68 in nuclear foci of transcriptionally active chromatin. Nucleic Acids Res 2010; 38: 3005–3018.
Paronetto MP, Miñana B, Valcárcel J . The Ewing sarcoma protein regulates DNA damage-induced alternative splicing. Mol Cell 2011; 43: 353–368.
Hayes GM, Carrigan PE, Miller LJ . Serine-arginine protein kinase 1 overexpression is associated with tumorigenic imbalance in mitogen-activated protein kinase pathways in breast, colonic, and pancreatic carcinomas. Cancer Res 2007; 67: 2072–2080.
Naro C, Sette C . Phosphorylation-mediated regulation of alternative splicing in cancer. Int J Cell Biol 2013; 2013: 151839.
Amin EM, Oltean S, Hua J, Gammons MV, Hamdollah-Zadeh M, Welsh GI et al. WT1 mutants reveal SRPK1 to be a downstream angiogenesis target by altering VEGF splicing. Cancer Cell. 2011; 20: 768–780.
Omenn GS, Yocum AK, Menon R . Alternative splice variants, a new class of protein cancer biomarker candidates: findings in pancreatic cancer and breast cancer with systems biology implications. Dis Markers 2010; 28: 241–251.
Christofk HR, Vander Heiden MG, Harris MH, Ramanathan A, Gerszten RE, Wei R et al. The M2 splice isoform of pyruvate kinase is important for cancer metabolism and tumour growth. Nature 2008; 452: 230–233.
Yang W, Xia Y, Hawke D, Li X, Liang J, Xing D et al. PKM2 phosphorylates histone H3 and promotes gene transcription and tumorigenesis. Cell 2012; 150: 685–696.
Yang W, Xia Y, Cao Y, Zheng Y, Bu W, Zhang L et al. EGFR-induced and PKCɛ monoubiquitylation-dependent NF-κB activation upregulates PKM2 expression and promotes tumorigenesis. Mol Cell 2012; 48: 771–784.
Shultz JC, Goehe RW, Murudkar CS, Wijesinghe DS, Mayton EK, Massiello A et al. SRSF1 regulates the alternative splicing of caspase 9 via a novel intronic splicing enhancer affecting the chemotherapeutic sensitivity of non-small cell lung cancer cells. Mol Cancer Res 2011; 9: 889–900.
Droin N, Rébé C, Bichat F, Hammann A, Bertrand R, Solary E . Modulation of apoptosis by procaspase-2 short isoform: selective inhibition of chromatin condensation, apoptotic body formation and phosphatidylserine externalization. Oncogene 2001; 20: 260–269.
Mercatante DR, Mohler JL, Kole R . Cellular response to an antisense-mediated shift of Bcl-x pre-mRNA splicing and antineoplastic agents. J Biol Chem 2002; 277: 49374–49382.
Anczuków O, Rosenberg AZ, Akerman M, Das S, Zhan L, Karni R et al. The splicing factor SRSF1 regulates apoptosis and proliferation to promote mammary epithelial cell transformation. Nat Struct Mol Biol 2012; 19: 220–228.
Proussakova OV, Rabaya NA, Moshnikova AB, Telegina ES, Turanov A, Nanazashvili MG et al. Oligomerization of soluble Fas antigen induces its cytotoxicity. J Biol Chem 2003 Sep; 278: 36236–36241.
Nakajima S, Lan L, Wei L, Hsieh CL, Rapić-Otrin V, Yasui A et al. Ubiquitin-specific protease 5 is required for the efficient repair of DNA double-strand breaks. PLoS One 2014; 9: e84899.
Adesso L, Calabretta S, Barbagallo F, Capurso G, Pilozzi E, Geremia R et al. Gemcitabine triggers a pro-survival response in pancreatic cancer cells through activation of the MNK2/eIF4E pathway. Oncogene 2013; 32: 2848–2857.
Maimon A, Mogilevsky M, Shilo A, Golan-Gerstl R, Obiedat A, Ben-Hur V et al. Mnk2 alternative splicing modulates the p38-MAPK pathway and impacts Ras-induced transformation. Cell Rep 2014; 7: 501–513.
Boccaccio C, Comoglio PM . Invasive growth: a MET-driven genetic programme for cancer and stem cells. Nat Rev Cancer 2006; 6: 637–645.
Sawa H, Ohshima TA, Ukita H, Murakami H, Chiba Y, Kamada H et al. Alternatively spliced forms of cyclin D1 modulate entry into the cell cycle in an inverse manner. Oncogene 1998; 16: 1701–1712.
David CJ, Chen M, Assanah M, Canoll P, Manley JL . HnRNP proteins controlled by c-Myc deregulate pyruvate kinase mRNA splicing in cancer. Nature 2010; 463: 364–368.
Tamada M, Suematsu M, Saya H . Pyruvate kinase M2: multiple faces for conferring benefits on cancer cells. Clin Cancer Res 2012; 18: 5554–5561.
Jiang Y, Li X, Yang W, Hawke DH, Zheng Y, Xia Y et al. PKM2 regulates chromosome segregation and mitosis progression of tumor cells. Mol Cell. 2014; 53: 75–87.
Wang Z, Jeon HY, Rigo F, Bennett CF, Krainer AR . Manipulation of PK-M mutually exclusive alternative splicing by antisense oligonucleotides. Open Biol 2012; 2: 120133.
Goldberg MS, Sharp PA . Pyruvate kinase M2-specific siRNA induces apoptosis and tumor regression. J Exp Med 2012; 209: 217–224.
Kole R, Krainer AR, Altman S . RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nat Rev Drug Discov 2012; 11: 125–140.
Xue Y, Zhou Y, Wu T, Zhu T, Ji X, Kwon YS et al. Genome-wide analysis of PTB-RNA interactions reveals a strategy used by the general splicing repressor to modulate exon inclusion or skipping. Mol Cell 2009; 36: 996–1006.
Carstens RP, Wagner EJ, Garcia-Blanco MA . An intronic splicing silencer causes skipping of the IIIb exon of fibroblast growth factor receptor 2 through involvement of polypyrimidine tract binding protein. Mol Cell Biol 2000; 20: 7388–7400.
Llorian M, Schwartz S, Clark TA, Hollander D, Tan LY, Spellman R et al. Position-dependent alternative splicing activity revealed by global profiling of alternative splicing events regulated by PTB. Nat Struct Mol Biol 2010; 17: 1114–1123.
Costello BA, Borad MJ, Qi Y, Kim GP, Northfelt DW, Erlichman C et al. Phase I trial of everolimus, gemcitabine and cisplatin in patients with solid tumors. Invest New Drugs 2014; 32: 710–716.
Braunschweig U, Gueroussov S, Plocik AM, Graveley BR, Blencowe BJ . Dynamic integration of splicing within gene regulatory pathways. Cell 2013; 152: 1252–1269.
Cerwenka H, Aigner R, Bacher H, Werkgartner G, el-Shabrawi A, Quehenberger F et al. TUM2-PK (pyruvate kinase type tumor M2), CA19-9 and CEA in patients with benign, malignant and metastasizing pancreatic lesions. Anticancer Res 1999; 19: 849–851.
Shi HS, Li D, Zhang J, Wang YS, Yang L, Zhang HL et al. Silencing of pkm2 increases the efficacy of docetaxel in human lung cancer xenografts in mice. Cancer Sci 2010; 101: 1447–1453.
Grosso AR, Martins S, Carmo-Fonseca M . The emerging role of splicing factors in cancer. EMBO Rep 2008; 9: 1087–1093.
Paronetto MP, Achsel T, Massiello A, Chalfant CE, Sette C . The RNA-binding protein Sam68 modulates the alternative splicing of Bcl-x. J Cell Biol 2007; 176: 929–939.
Bielli P, Busà R, Di Stasi SM, Munoz MJ, Botti F, Kornblihtt AR et al. The transcription factor FBI-1 inhibits SAM68-mediated BCL-X alternative splicing and apoptosis. EMBO Rep 2014; 15: 419–427.
Bielli P, Bordi M, Di Biasio V, Sette C . Regulation of BCL-X splicing reveals a role for the Polypyrimidine-tract binding protein (PTBP1/hnRNP I) in alternative 5 ‘splice site selection. Nucleic Acids Res 2014; 42: 12070–12081.
Acknowledgements
We thank Andrea D’Ascenzo and Enrico Duranti for help with immunohistochemistry; Dr Chiara Naro for helpful suggestions throughout the study; and Dr Vittoria Pagliarini for critical reading of the manuscript. This work was supported by Association for International Cancer Research (AICR) (grant 12-0150), Associazione Italiana Ricerca sul Cancro (AIRC) (grant 14581) and Fondazione Santa Lucia Ricerca Corrente.
Author contributions
SC, PB and IP performed the experiments and analyzed the data. EP and VF performed IHC experiments and score analysis. GC performed statistical analysis of patients; SC, GDF, GC and CS wrote the manuscript. SC and CS designed the study.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
About this article
Cite this article
Calabretta, S., Bielli, P., Passacantilli, I. et al. Modulation of PKM alternative splicing by PTBP1 promotes gemcitabine resistance in pancreatic cancer cells. Oncogene 35, 2031–2039 (2016). https://doi.org/10.1038/onc.2015.270
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2015.270