Abstract
The TP53 tumor suppressor gene is frequently inactivated in human tumors by missense mutations in the DNA binding domain. TP53 mutations lead to protein unfolding, decreased thermostability and loss of DNA binding and transcription factor function. Pharmacological targeting of mutant p53 to restore its tumor suppressor function is a promising strategy for cancer therapy. The mutant p53 reactivating compound APR-246 (PRIMA-1Met) has been successfully tested in a phase I/IIa clinical trial. APR-246 is converted to the reactive electrophile methylene quinuclidinone (MQ), which binds covalently to p53 core domain. We identified cysteine 277 as a prime binding target for MQ in p53. Cys277 is also essential for MQ-mediated thermostabilization of wild-type, R175H and R273H mutant p53, while both Cys124 and Cys277 are required for APR-246-mediated functional restoration of R175H mutant p53 in living tumor cells. These findings may open opportunities for rational design of novel mutant p53-targeting compounds.
Similar content being viewed by others
Introduction
Tumor suppressor p53 is a transcription factor that responds to cellular stress, e.g., DNA damage and oncogene activation. p53 triggers cell cycle arrest, senescence and apoptosis1,2. More recent studies have shown that p53 also has roles in metabolism3, stem cell division4, fertility5 and cell death by ferroptosis6. The TP53 gene is inactivated in a large fraction of human tumors7,8. The majority of TP53 mutations are missense mutations resulting in substitution of amino acid residues that make direct contact with DNA, such as R248W and R273H, or residues that are important for the structural integrity of the core domain, e.g. R175H and R249S. This leads to loss of specific DNA binding9.
The high frequency of TP53 mutations in human tumors has stimulated efforts to develop therapeutic strategies for targeting mutant p53 in cancer. Several low molecular weight compounds have been reported to restore wild-type function to mutant p53, including PRIMA-1 and the PRIMA-1 analog APR-246 (PRIMA-1Met)10,11,12,13,14,15,16,17,18. In addition to targeting mutant p53, APR-246 inhibits the selenoprotein thioredoxin reductase 1 (TrxR1) and converts it to an active oxidase19, and depletes cellular gluthatione (GSH)20,21,22. This induces reactive oxygen species (ROS) and presumably contributes to the anticancer effect of APR-24623. Liu et al. showed that SLC7A11 expression is a new marker for sensitivity of mutant p53-carrying tumors to APR-246/MQ. Mutant p53 binds to the antioxidant transcription factor Nrf2, leading to decreased expression of SLC7A11, which sensitizes cells to TrxR1 inhibition and GSH depletion by APR-246/MQ22. APR-246 has been tested in a phase I/IIa clinical trial in patients with hematological malignancies or prostate cancer24. APR-246 is converted to methylene quinuclidinone (MQ), a Michael acceptor that reacts with cysteines in the p53 core domain25. However, the mechanism by which APR-246/MQ reactivates mutant p53 is not fully understood.
Many mutant p53 proteins are thermodynamically unstable at body temperature26. Studies of temperature-sensitive mutants suggest that stabilization of conformation is critical for regaining wild-type p53 activity27,28. Thus, pharmacological stabilization of mutant p53 should allow its functional rescue and efficient elimination of tumor cells29.
Here we examined the role of the Michael acceptor activity of MQ for thermostabilization of wild-type (wt) and mutant p53 core domains and refolding of R175H mutant p53 in living cells. We also show that Cys277 is essential for MQ-mediated thermostabilization of R175H and R273H mutant p53 core domains, and that both Cys124 and Cys277 are required for APR-246-mediated R175H mutant p53 reactivation in tumor cells.
Results
MQ binds to the p53 core domain via Michael addition
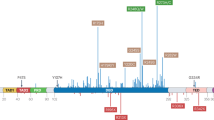
First, we studied MQ binding to p53 by Nanomate Linear Ion Trap Orbitrap hybrid mass spectrometry (LTQ-MS). Accurate masses of the p53 core domains are shown in Fig. 1a. We incubated wt and R273H mutant p53 core domains (20 µM) with increasing concentrations of MQ at 21 °C and assessed MQ adduct formation by LTQ-MS. The deconvoluted mass spectra showed that 32% of the wt and R273H p53 core domain proteins were modified by one MQ molecule when incubated with lower concentrations of MQ (Fig. 1b), and that all p53 protein molecules had two MQ adducts upon incubation with 200 µM MQ (Fig. 1d). We did not detect modification of all 10 cysteines in the p53 core at these concentrations of MQ.
Mass measurement of wild-type, R273H and R175H p53 core domains by LTQ-MS. a mass spectra of p53 core domains. b–d Reaction titration with MQ or MQ-H. p53 core domains were incubated with MQ at 50–200 µM (wt and R273H) or 10–50 µM (R175H) concentration ranges. One MQ adduct increased the molecular mass of p53 core domains by 137 Da. e Structure of MQ and MQ-H.
Due to low yield of the R175H p53 core domain, we analyzed it at lower protein concentration. LTQ-MS indicated that 20% of the R175H core was modified by one MQ molecule at 10 µM (Fig. 1c), and at 50 µM MQ, 35% of the R175H core domain protein had one MQ adduct and 14% had two MQ adducts (Fig. 1d). Thus, the number of MQ-modified cysteine residues increased in a concentration-dependent manner.
Next, we assessed the binding of MQ-H, a hydrogenated analog of MQ that lacks reactive carbon-carbon double bond (Fig. 1e). We did not detect any modification at concentrations up to 200 µM, indicating that MQ-H does not modify cysteine residues in p53. Thus, the Michael acceptor activity of MQ is required for modification of cysteines in p53.
MQ enhances thermostability of the p53 core domain
To test if MQ modification of cysteines increases thermostability of the p53 core domain, we applied differential scanning fluorimetry (DSF). This allows analysis of the interactions between a protein and a ligand based on changes in the protein melting temperature (Tm). DSF (Fig. 2a) demonstrates that the wt p53 core domain was the most stable of the three proteins with a Tm of 40.38 ± 0.06 °C. Tm for the DNA contact mutant R273H was 39.16 ± 0.52 °C, whereas the structural mutant R175H was the least stable protein with a Tm of 30.64 ± 0.46 °C, consistent with previously published studies26. To further validate this result, we used circular dichroism spectroscopy (CD), which allows assessment of α-helix and β-sheet structure content. We performed CD analysis at 218 nm since both α-helix and β-sheet structures are detected at this wavelength. In agreement with our DSF data, CD (Fig. 2b) measurements demonstrated that the wt p53 core domain is the most stable protein followed by the R273H core domain whereas the R175H core is considerably less stable. The Tm values are 44.74 ± 0.08 °C, 43.96 ± 0.37 °C and 36.24 ± 0.33 °C, respectively.
Melting curves of wt and R273H and R175H mutant p53 core domains as assessed by DSF (a) and CD at 218 nm (b). Changes in Tm after MQ or MQ-H modification as assessed by DSF (c) and CD at 218 nm (d) (mean ± SD, n = 3). All proteins were thermostabilized by MQ modification (red bars), but not by MQ-H (brown bars). * p < 0.05 (student t-test).
We then incubated wt, R273H and R175H p53 core domains with 2 mM MQ and performed DSF and CD analyses. This concentration of MQ was chosen to override the reducing agents DTT or TCEP included in the reaction buffer to preserve the structure of the p53 core domains. According to DSF, MQ increased the Tm value of wt, R273H and R175H p53 core domains by 3.44, 3.54 and 2.31 °C, respectively (Fig. 2c). This degree of thermal stabilization by small molecules is in agreement with previously published results16. CD analysis confirmed thermostabilization of all three cores by MQ as shown by the increase in Tm values by 1.20, 1.70, and 2.06 °C for wt, R273H and R175H p53, respectively (Fig. 2d). The inactive MQ analog MQ-H did not significantly change the thermal stability of the p53 core domains (Fig. 2c, d).
Thus, MQ but not MQ-H increases the thermostability of p53 core domains, confirming that cysteine binding by Michael addition is critical for p53 thermostabilization by MQ. The results from DSF and CD were fully consistent; the protein melting temperatures determined by the two methods correlated with each other (r = 0.999, p = 0.007). DSF was chosen for further studies.
Identification of MQ binding sites in the p53 core domain
The p53 core domain has 10 cysteine residues with varying solvent accessibility. Previous studies have indicated that in the absence of DNA, Cys277 has the highest solvent accessibility, followed by Cys182 and Cys229, whereas Cys135, Cys141 and Cys275 have poor solvent accessibility30. Cys124 is located at the center of the flexible L1/S3 pocket, which can be stabilized by second-site mutations to rescue mutant p53 folding31,32,33. Interestingly, Cys124 shows a nuclear magnetic resonance (NMR) chemical shift upon binding of the CDB3 peptide that stabilizes mutant p5334. In addition, substitution of Cys124 abrogated reactivation of R175H mutant p53 by PRIMA-131.
To investigate if the most solvent-exposed cysteine residues and Cys124 are critical for MQ binding to p53, we introduced single Cys to Ala substitutions at position 124, 182, 229 or 277 in the wt, R175H and R273H p53 core domains, and double substitutions at Cys124 and Cys277 in the same three core domains. The R175H p53 core domain has low intrinsic thermostability and additional amino acid substitutions might destabilize it further. This probably explains why we obtained negligible protein yield for the R175H–C124A and R175H–C124A–C277A core domains. LTQ-MS analysis of the R273H mutant p53 core domains identified one MQ adduct in the R273H, R273H–C124A, R273H–C182A, and R273H–C229A p53 cores after incubation with 50 µM MQ, whereas no MQ modification of the R273H–C277A and R273H–C124A–C277A core domains was detected (Fig. 3a). However, incubation with 100–200 µM MQ resulted in 2–5 MQ adducts in all p53 mutant core domains tested except R273H–C277A (Fig. 3b, c). This implies that MQ modifies several cysteines in p53, and suggests that Cys277 is the most reactive.
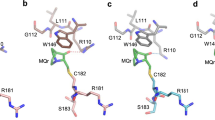
Mass measurement of p53 core domains by LTQ-MS. a–c Indicated p53 core domain proteins were incubated with MQ at increasing concentrations and mass shift was assessed. Proportion of MQ-modified proteins was calculated based on relative intensity of each detected mass. d Modification of Cys124 and Cys275/277 in the R273H mutant p53 core domain after treatment with 50 µM MQ as assessed by LC-MS after tryptic digestion of MQ-treated protein (mean ± SE, n = 3).
To confirm modification of Cys124 and Cys277 by MQ, we incubated recombinant R273H mutant p53 core domain with 50 µM MQ, digested with trypsin, and analyzed peptides by LC-MS (liquid chromatography-mass spectrometry). We detected modification of p53 peptides SVTCTYSPALNK and NSFEVHVCACPGR that contain Cys124 and Cys275/277, respectively. The degree of modification of Cys124 and Cys275/277 could be estimated to 0.14 ± 0.03% and 0.04 ± 0.02 (mean ± SE, n = 3), respectively. Amount of each peptide is expressed as a percentage of the amount of the corresponding unmodified peptide detected in the same sample according to LC-MS (Fig. 3d).
Cys277 is essential for MQ-mediated thermostabilization of p53 core domains
To assess the role of selected Cys residues in MQ-mediated thermostabilization, we incubated wt, R175H and R273H p53 core domains carrying indicated Cys to Ala substitutions with MQ and performed DSF. The Tm values of the wt and p53 core domains carrying C124A, C277A, C182A, C229A, and C124A–C277A substitutions were 42.36, 36.67, 43.07, 42.43, 40.59 and 40.91 °C, respectively (Table 1). MQ modification increased the Tm values of the C124A, C182A, and C229A p53 core domains by 5.82, 1.35 and 1.00 °C (Fig. 4a), respectively. In contrast, MQ caused a slight destabilization of the C277A core domain by −0.06 °C, and only stabilized the C124A–C277A core domain by 0.16 °C, indicating that Cys277 is critical for MQ-mediated thermostabilization of p53.
C277A substitution completely abolished MQ-mediated thermostabilization whereas other substitutions had little or no effect on wt a, R273H b and R175H c core domains at 2 mM concentration. Higher concentrations of MQ (1, 2 or 4 mM) did not thermostabilize the indicated p53 core domain proteins with C277A substitution d.
We then analyzed R273H p53 core domains with the same C124A, C277A, C182A and C229A substitutions. The melting temperatures of the R273H, R273H–C124A, R273H–C227A, R273H–C182A, and R273H–C229A p53 core domains were 40.49, 39.17, 41.48, 40.52, and 38.60 °C, respectively (Table 1). MQ shifted the Tm values by 0.93, 0.64, −0.19, 1.03 and 0.76 °C (Fig. 4b), respectively. The Tm of the R273H core domain with C124A–C277A double substitution was 39.48 °C and only changed by −0.07 °C upon MQ modification.
As indicated above, we were not able to obtain sufficient amounts of the R175H–C124A and R175H–C124A–C277A p53 core domains. Thus, only the R175H, R175H–C277A, R175H–C182A and R175H–C229A core domains were further analyzed. Their respective melting temperatures were 31.09, 29.05, 29.49, and 28.16 °C (Table 1). MQ modification shifted the melting temperatures by 0.71, −0.12, 0.73, and 0.45 °C (Fig. 4c), respectively. Thus, C227A substitution abrogated MQ-induced thermostabilization in wt and R273H and R175H mutant p53 core domain backgrounds.
Our MS data demonstrate that MQ binds to the p53 core domain in a concentration-dependent manner and that high concentrations of MQ lead to modification of more than one cysteine residue. This raises the question whether high concentrations of MQ might induce p53 thermostabilization even in the absence of Cys277. To address this, we incubated wt, C277A, C124A–C277A, R175H–C277A and R273H–C277A p53 core domains with 1, 2 or 4 mM MQ and performed DSF (Fig. 4d). The highest concentration of MQ (4 mM) destabilized all core domains (Fig. 4d). The wt p53 core was stabilized by MQ at 1 and 2 mM, as observed previously. However, p53 core domains with the C277A substitution were not stabilized at the same concentrations. This further supports our conclusion that Cys277 is indispensable for MQ-mediated p53 core domain stabilization.
APR-246 and MQ induce R175H mutant p53 refolding to wild-type conformation
Proper folding of p53 is crucial for sequence-specific DNA binding and transactivation of p53 target genes. Most TP53 mutations lead to unfolding and loss of sequence-specific DNA binding. PRIMA-1 was shown to promote R175H mutant p53 refolding to wild-type conformation in SKOV-His175 cells11. However, this has yet to be demonstrated for APR-246 and MQ.
The monoclonal antibody PAb1620 detects properly folded p53 but not unfolded p53. We verified the specificity of this antibody in immunostaining by treating HCT116 human colon carcinoma cells with doxorubicin to induce the levels of wild-type p53. HCT116 cells expressing wild-type p53 or Saos-2 cells expressing R273H mutant p53, which retains wild-type-like conformation, showed positive PAb1620 staining. In contrast, H1299 cells expressing R175H mutant p53 were PAb1620 negative. Treatment with APR-246 induced positive PAb1620 staining in H1299–R175H cells, suggesting that APR-246 via MQ restores wild-type conformation of this mutant (Supplementary Figure 1).
To determine if APR-246 and MQ can refold endogenous R175H mutant p53 in TOV-112D ovarian carcinoma cells, we treated the cells with APR-246 or MQ and performed co-immunostaining with PAb1620 and anti-p53 polyclonal antibody FL-393. In parallel, the cells were stained with the monoclonal HO3.5 antibody that specifically detects unfolded p53 similarly to the PAb240 antibody35. APR-246 treatment increased PAb1620 staining (Fig. 5a, c) and decreased HO3.5 staining (Fig. 5b, d), indicating refolding of R175H mutant p53 to wild-type-like conformation.
a Immunofluorescence staining of TOV-112D cells treated with APR-246, MQ or MQ-H using wild-type p53 conformation-specific antibody PAb1620 and co-immunostaining with general p53 antibody FL-393. b Immunofluorescence staining of TOV-112D cells treated with APR-246, MQ or MQ-H using the mutant p53 conformation-specific antibody HO3.5 and co-immunostaining with general p53 antibody FL-393. c Quantification of PAb1620 immunostaining. d Quantification of HO3.5 immunostaining. PAb1620 or HO3.5-positive cells were counted in 15 fields from 3 independent experiments. The number of PAb1620 or HO3.5-positive cells was divided by the total cell number to obtain percentage of positive cells.
Next, we treated TOV-112D cells with MQ and stained with PAb1620 and HO3.5 antibodies. Like APR-246, MQ-induced PAb1620 staining (Fig. 5a), which coincided with decreased HO3.5 staining (Fig. 5b). We did not detect any changes in PAb1620 and HO3.5 staining after treatment with MQ-H, confirming that the Michael acceptor activity of MQ is crucial for mutant p53 refolding in living tumor cells.
Cysteines 124 and 277 are important for APR-246/MQ-mediated R175H mutant p53 reactivation
To investigate the role of cysteine residues in mutant p53 reactivation by APR-246/MQ in tumor cells, we transiently transfected p53 null H1299 cells with vectors encoding R175H, R175H–C124A, R175H–C277A or R175H–C124A–C277A p53 mutant proteins, or with control vector (pCMV). Western blotting confirmed similar levels of p53 expression in all transfectants (Supplementary Figure 2). Next, the cells were treated with 45 µM APR-246. We chose this concentration since it induced cell death in R175H mutant p53-transfected cells but only marginally affected empty vector-transfected cells. We then assessed apoptosis and expression of p53 targets p21, Fas and Bax by flow cytometry. Signals obtained in mock-treated control cells were subtracted from the signals obtained in APR-246-treated cells, and the values are presented as relative increase (ΔAnnexin V, Δp21, ΔFas, and ΔBax). Cells transfected with R175H showed substantial induction (61.06%) of Annexin V staining after APR-246 treatment when compared to cells transfected with empty vector (3.91%). We observed a relatively small induction of Annexin V staining by APR-246 in cells transfected with R175H–C124A p53 (13.96%), and cells expressing R175H–C277A or R175H–C124A–C277A p53 showed no induction of Annexin V staining as compared to empty vector-transfected cells (Fig. 6a). The p53 targets p21 (Fig. 6b) and Bax (Fig. 6c) were highly induced by APR-246 in cells expressing R175H p53 but only slightly or moderately increased in cells expressing R175H–C124A and R175H–C124A–C277A mutants. Cells expressing R175H–C277A mutant p53 showed no induction of p21 or Bax. The p53 target Fas (Fig. 6d) was strongly induced by APR-246 in R175H-transfected cells but no induction compared to control-transfected cells was detected in cells expressing R175H–C124A, R175H–C277A or R175H–C124A–C277A p53 proteins, consistent with the observed absence of apoptosis induction.
To determine whether the C124A and C277A substitutions themselves affect wild-type p53 function, H1299 cells transfected with wt, C124A, C277A, R175H and R273H p53 constructs were assessed for apoptosis and expression of p53 target p21 by flow cytometry. pCMV vector was used as a control. Induction of Annexin V (Supplementary Figure 3a) was detected in cells expressing wt, C124A or C277A p53 proteins, which coincided with slight induction of p21 (Supplementary Figure 3b), but not in cells transfected with R175H or R273H p53 constructs. Thus, C124A or C277A substitution per se does not inactivate p53 in this experimental system, supporting our conclusion that these cysteines play a key role in APR-246/MQ-mediated mutant p53 reactivation.
Discussion
Small molecules that reactivate mutant p53 by restoring wild-type conformation have been identified by various approaches10,23,36. APR-246 has been tested in a phase I/IIa clinical trial in patients with hematological malignancies or prostate cancer24,37, and is currently tested in a phase II trial in patients with high-grade serous (HGS) ovarian cancer (http://www.clinicaltrials.gov/ct2/show/NCT02098343). We previously demonstrated thiol modifications in the p53 core domain by PRIMA-1 conversion products25. This led us to conclude that APR-246-mediated mutant p53 reactivation involves covalent binding of MQ. Other mutant p53-reactivating compounds, such as MIRA-138, CP-31398 and STIMA-139, 3-benzoylacrylic acid14, 2-sulfonylpyrimidines16, and the curcumin analog HO-386718 possess similar thiol reactivity, indicating that the observed association between thiol reactivity and mutant p53 reactivation is not coincidental.
Here we show that the MQ analog MQ-H that lacks a reactive carbon-carbon double bond and therefore lacks Michael acceptor activity, does not modify cysteine residues in the p53 core domain, does not enhance p53 thermostability and does not induce R175H mutant p53 refolding according to PAb1620 staining. Thus, by using several approaches, we demonstrate that the electrophilic properties of MQ are essential for cysteine modification, thermostabilization and refolding of mutant p53.
Although our previous studies indicated that PRIMA-1 conversion products bind covalently to the p53 core domain25, the exact cysteine targets for MQ have remained unknown. We applied LTQ-MS analysis to a set of Cys to Ala mutants to identify cysteine residues that are critical for MQ binding and MQ-mediated stabilization of mutant p53. The reactivity of cysteine residues in a protein is largely affected by their solvent accessibility. Among 10 cysteines in p53 core domain, Cys176, Cys238, and Cys242 coordinate a zinc ion which is responsible for holding p53 loops together9, making them less likely targets for modification. Cys135, Cys141, and Cys275 are poorly accessible to solvent based on the X-ray crystal structure of the p53 core domain. Cys277 and Cys182 have the highest solvent accessibility, followed by Cys22930. Interestingly, Cys277 has the lowest pKa of all p53 cysteines, making it the strongest nucleophile in the protein. Thus, Cys277 combines the greatest solvent accessibility with the highest nucleophilicity, suggesting that it might be a prime target for MQ.
Indeed, we found that Cys277 to Ala substitution abolishes MQ binding to p53 core domain, at lower concentrations. Moreover, the ability of MQ to thermostabilize p53 core domains is impaired in Cys277 to Ala mutants. Thus, a good correlation exists between the extent of MQ adduct formation and MQ-mediated thermostabilization of the p53 core domain.
Cys182 and Cys277 have recently been identified as prime binding sites for PK11000, a 2-sulfonylpyrimidine compound that reacts with cysteines and thermostabilizes p5316. It is noteworthy that although Cys277 interacts directly with DNA, modification of this residue by PK11000 did not change p53 DNA binding and transactivation of target genes. The mutant p53-targeting curcumin analog HO-3867 also binds covalently to Cys182 and Cys27718. However, our results demonstrate that substitution of Cys182 to Ala does not affect p53 modification and thermostabilization by MQ in any significant way.
Kaar and colleagues14 identified 3-benzoylacrylic acid as a thiol-binding compound that reacts first with Cys124 and Cys141 and to a lesser extent with Cys135, Cys182 and Cys277 in p53. Cys124 was also identified as a target for PRIMA-1 by molecular modeling, and Cys124 to Ala substitution abolished PRIMA-1-induced reactivation of mutant p53 in human tumor cells31. Here we found that substitution of this cysteine did not impair MQ binding and p53 thermostabilization. However, Cys124 to Ala substitution abrogated R175H reactivation by APR-246/MQ in tumor cells, in agreement with the results of Wassmann et al.31.
It is unclear why the C124A mutant core domain was more efficiently stabilized by MQ than wt p53 or R273H–C124A p53 (Fig. 4). Notably, Cys124 to Ala substitution decreases p53 core domain stability more extensively in a wt background. Although the structures of wt and R273H p53 core domains are presumably similar, it is difficult to predict local structural changes after introduction of C124A. Thus, this question can probably only be resolved by NMR and/or X-ray crystallography analysis of MQ-modified p53.
To exclude the possibility that the Cys to Ala substitutions themselves impair wild-type p53 function in our experimental setting, we examined whether the Cys to Ala substitutions affect the ability of p53 to transactivate p21 and induce apoptosis. We confirmed that these substitutions do not affect normal p53 function to any major extent, supporting the notion that the observed effects are indeed due to an important role of Cys124 and Cys277 for APR-246/MQ-mediated mutant p53 reactivation.
In conclusion, our data demonstrate that specific cysteines are critical targets for mutant p53 reactivation by APR-246/MQ. Our findings may open opportunities for designing novel compounds targeting mutant p53 based on a similar mechanism of nucleophilic addition at the identified binding sites.
Materials and Methods
Cell lines and reagents
Human lung adenocarcinoma cells H1299 and osteosarcoma cells Saos-2 are p53 null. The sub-lines H1299–R175H and Saos-2-R273H stably express the indicated mutants11,38. Human HCT116 colon carcinoma cells express wild-type p53. Human epithelial ovarian cancer cells TOV-112D express R175H mutant p53. All cells were cultured at 37 °C, 5% CO2 in IMDM medium (Hyclone, Logan, Utah) supplemented with 10% FBS (Thermo Fisher Scientific, Waltham, MA).
APR-246, MQ, and MQ-H were obtained from Aprea Therapeutics AB, Stockholm, Sweden. Methanol, formaldehyde, and acetonitrile were purchased from Thermo Fisher Scientific (Waltham, MA). Formic acid was purchased from Sigma-Aldrich (St. Louis, MO). Lipofectamine 2000 was from Thermo Fisher Scientific (Waltham, MA). All solvents were of analytical grade and are commercially available.
Rabbit polyclonal anti-p53 FL-393, rabbit polyclonal anti-GAPDH, mouse monoclonal anti-p53 DO-1 and mouse monoclonal PAb1620, Alexa Fluor 647 conjugated FL-393, Alexa Fluor 488 conjugated anti-p21 antibodies were from Santa Cruz Biotechnology (Heidelberg, Germany). Mouse monoclonal antibody HO3.5 was a gift from Professor Thierry Soussi, Karolinska Institutet. Polyclonal rabbit anti-Bax Biotin OAAF02999 and QdotTM 605 streptavidin were from Nordic Biosite (Stockholm, Sweden). BV510 mouse anti-human CD95 (Fas) and BD Horizon V450 Annexin V were from BD Biosciences (Stockhlom, Sweden).
Site-directed mutagenesis
Prokaryotic and eukaryotic plasmid constructs were produced by Genscript, Piscataway, NJ.
Expression and purification of proteins
p53 cores (94–292) were cloned into pNIC28-Bsa4 that adds an N-terminal hexahistidine tag and transformed into E. coli strain Rosetta2 (DE3). Bacteria were grown in TB medium supplemented with 8 g/l glycerol at 37 °C with shaking. Protein expression was induced with 0.5 mM IPTG at 18 °C overnight. Afterwards bacteria were pelleted by centrifugation and lyzed in cold IMAC lysis buffer (50 mM TRIS, 300 mM NaCl, 10% glycerol, 0.05 mM ZnCl, 0.5 mM TCEP, pH 8.0) supplemented with complete protease mix (complete EDTA-free (protease inhibitor) and 5 μl benzonase nuclease (250 U) and stored at −80 °C. After thawing, the cells were lyzed by pulsed sonication (4 s/4 s 3 min, 80% amplitude), centrifuged (20 min at 49,000×g) and the soluble fractions were decanted and filtered through 0.45μm filters. The samples were loaded onto the ÄKTA Xpress LC and purified overnight. His-tag was cleaved with Thrombin. Sample homogeneity was confirmed by mass spectrometry and the concentration was measured by nanodrop. The proteins were aliquoted and stored at −80 °C in storage buffer (50 mM TRIS, 800 mM NaCl, 10% glycerol, 2.0 mM TCEP, pH 8.0).
Mass spectrometry
Wild-type and R273H p53 core domains were de-salted against 20 mM ammonium acetate buffer by using 10 K concentration columns (Vivaspin, GE Healthacare, Chicago, IL). Twenty µM of the purified protein were incubated with 0 µM (control), 50, 100 or 200 µM MQ for 15 min at 21 °C. R175H core domains were de-salted by ZipTip C4 resin tips for MALDI-ToF MS (Merck Millipore, Billerica, MA) following the manufacturer’s protocol. 3.2 µM of R175H protein were treated with 0 µM (control), 10, 25 or 50 µM of MQ for 15 min at 21 °C. 5% formic acid (1:1 volume ratio) was added to the samples to increase the ionization sensitivity. Samples were analyzed by LTQ XL mass spectrometry (Thermo Fisher Scientific, Waltham, MA) fitted with an automated nanospray source (TriVersa Nanomate, Advion Biosciences, Ithaca, NY) using nanoelectrospray chips with spraying nozzels. The ion source was controlled using the Chipsoft 8.3.1 software (Advion Biosciences, Ithaca NY). Three microliters of each sample were loaded into a 96-well plate and injection volume was one and a half microliters. Full scan spectra were collected at the m/z 500–2000 in positive ion mode. The mass spectra of each sample were acquired in profile mode over 4 min. The spectra were analyzed using XCaliburTM Software (Thermo Fisher Scientific, Waltham, MA). Deconvoluted ESI spectra are presented.
LC-MS
30 μg of R273H p53 core domain protein was treated with 50 μM MQ in 20 mM ammonium bicarbonate pH 8.0 for 1 h at room temperature. Samples were then precipitated with acetone and pellets were digested with trypsin at 37 °C for 3 h. 10 µl of each sample was injected onto Waters Alliance HPLC system (Waters, Sollentuna, Sweden) and resolved on XSelect® Peptide CHSTM C18, XP column, 130 Å, 2.5 μm, 2.1 × 150 mm (Waters, Sollentuna, Sweden). The peptide separation was performed by water-acetonitrile gradient elution supplemented with 0.1% formic acid. Peptides were detected with Acquity QDa MS detector in ESI + mode. Data were analyzed by MassLynx software (Waters, Sollentuna, Sweden).
Circular dichroism
75 μg p53 core domain proteins were incubated with or without 2 mM MQ in 250 µl 40 mM potassium phosphate buffer (pH 7.5) and 1 mM DTT for 1 h at 21 °C. The concentration of p53 was 13.3 μM. CD measurements were performed on Jasco-810 (Jasco Inc., Tokyo, Japan) with 1.00 mm pathlength. Denaturation curves were obtained by measuring the CD spectra at 218 nm. Melting temperatures were analyzed by GraphPad Prism 6 (Graphpad Software Inc, La Jolla, CA) according to the Boltzmann equation \(y = \frac{{A_1 - A_2}}{{1 + e^{\left( {x - x_0} \right)/dx}}} + A_2\), xo - inflection point.
Differential scanning fluorimetry
5 μg p53 core domain proteins were incubated with 1–4 mM MQ in 25 μl 40 mM potassium phosphate buffer (pH 7.5) with 1 mM DTT for 1 h at 21 °C under controlled conditions. The concentration of p53 was 8.8 μM. 1 μl of 25× Sypro orange were added to each well. The fluorescence was assessed by Bio-Rad iCycler (Bio-Rad Laboratories, CA) at increasing temperature from 10 to 75 °C with a rate of 1 °C per min. Tm values were calculated by GraphPad Prism 6 (Graphpad Software Inc, La Jolla, CA) in the same way as for CD.
Immunofluorescence staining
Cells were plated into a 16-well chamber slide at a density of 3000 cells per well, allowed to attach overnight, and treated with 25 µM APR-246 or 5 µM MQ/MQ-H for 16 h. Cells were washed, fixed with 4% formaldehyde and permealized with 0.2% Triton X. Mouse PAb1620 or HO3.5 antibody were co-incubated with rabbit FL-393 antibody, all were diluted 1:200 in 2% BSA for 1 h at 4 °C. Anti-rabbit Alexa 488 and anti-mouse Alexa 594 conjugates were used as secondary antibody with 1:200 dilution in 2% BSA.
Flow cytometry
Cells were grown on 6-well plates at an initial density of 500,000 cells/well. Sixteen hours later cells were transfected for 24 h with p53 expression vectors or empty vector using Lipofectamine 2000 according to the manufacturer’s protocol (Life Technology, Waltham, MA). The medium was then replaced with fresh medium, the cells were reseeded at a density of 20,000 cells/well after 6 h culture, and treated with APR-246 on the following day. Cells were collected 24 h post-treatment, stained with Annexin V, fixed with 4% formaldehyde, permealized with 90% methanol and stained with Fas, Bax, and p21 antibodies. Cells were analyzed on a NovoCyte Flow Cytometer (ACEA Biosciences, Solna, Sweden).
Change history
10 October 2019
Since publication of this article, the authors have noticed that there was an error in Fig. 1d, third panel from left, “R273H + 200 μM MQ-H” should be “R273H + 200 μM MQ”. Our corrections do not affect the original conclusions of this paper.
References
Vousden, K. H. & Prives, C. Blinded by the light: the growing complexity of p53. Cell 137, 413–431 (2009).
Kastenhuber, E. R. & Lowe, S. W. Putting p53 in context. Cell 170, 1062–1078 (2017).
Puzio-Kuter, A. M. The role of p53 in metabolic regulation. Genes Cancer 2, 385–391 (2011).
Bonizzi, G., Cicalese, A., Insinga, A. & Pelicci, P. G. The emerging role of p53 in stem cells. Trends Mol. Med 18, 6–12 (2012).
Hu, W., Zheng, T. & Wang, J. Regulation of fertility by the p53 family members. Genes Cancer 2, 420–430 (2011).
Jiang, L. et al. Ferroptosis as a p53-mediated activity during tumour suppression. Nature 520, 57–62 (2015).
Soussi, T. & Wiman, K. G. TP53: an oncogene in disguise. Cell Death Differ. 22, 1239–1249 (2015).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Cho, Y., Gorina, S., Jeffrey, P. D. & Pavletich, N. P. Crystal structure of a p53 tumor suppressor-DNA complex: understanding tumorigenic mutations. Science 265, 346–355 (1994).
Bykov, V. J. & Wiman, K. G. Mutant p53 reactivation by small molecules makes its way to the clinic. FEBS Lett. 588, 2622–2627 (2014).
Bykov, V. J. et al. Restoration of the tumor suppressor function to mutant p53 by a low-molecular-weight compound. Nat. Med. 8, 282–288 (2002).
Bykov, V. J., Issaeva, N., Selivanova, G. & Wiman, K. G. Mutant p53-dependent growth suppression distinguishes PRIMA-1 from known anticancer drugs: a statistical analysis of information in the National Cancer Institute database. Carcinogenesis 23, 2011–2018 (2002).
Bykov, V. J. et al. PRIMA-1(MET) synergizes with cisplatin to induce tumor cell apoptosis. Oncogene 24, 3484–3491 (2005).
Kaar, J. L. et al. Stabilization of mutant p53 via alkylation of cysteines and effects on DNA binding. Protein Sci. 19, 2267–2278 (2010).
Liu, X. et al. Small molecule induced reactivation of mutant p53 in cancer cells. Nucleic Acids Res. 41, 6034–6044 (2013).
Bauer, M. R., Joerger, A. C. & Fersht, A. R. 2-Sulfonylpyrimidines: mild alkylating agents with anticancer activity toward p53-compromised cells. Proc. Natl. Acad. Sci. USA 113, E5271–E5280 (2016).
Yu, X., Vazquez, A., Levine, A. J. & Carpizo, D. R. Allele-specific p53 mutant reactivation. Cancer Cell 21, 614–625 (2012).
Madan E., et al. The curcumin analog HO-3867 selectively kills cancer cells by converting mutant p53 protein to transcriptionally active wildtype p53. J. Biol. Chem. https://doi.org/10.1074/jbc.RA117.000950. 2018.
Peng, X. et al. APR-246/PRIMA-1MET inhibits thioredoxin reductase 1 and converts the enzyme to a dedicated NADPH oxidase. Cell Death Dis. 4, e881 (2013).
Mohell, N. et al. APR-246 overcomes resistance to cisplatin and doxorubicin in ovarian cancer cells. Cell Death Dis. 6, e1794 (2015).
Tessoulin, B. et al. PRIMA-1Met induces myeloma cell death independent of p53 by impairing the GSH/ROS balance. Blood 124, 1626–1636 (2014).
Liu, D. S. et al. Inhibiting the system xC-/glutathione axis selectively targets cancers with mutant-p53 accumulation. Nat. Commun. 8, 14844 (2017).
Bykov, V. J. N., Eriksson, S. E., Bianchi, J. & Wiman, K. G. Targeting mutant p53 for efficient cancer therapy. Nat. Rev. Cancer 18, 89–102 (2018).
Lehmann, S. et al. Targeting p53 in vivo: a first-in-human study with p53-targeting compound APR-246 in refractory hematologic malignancies and prostate cancer. J. Clin. Oncol. 30, 3633–3639 (2012).
Lambert, J. M. et al. PRIMA-1 reactivates mutant p53 by covalent binding to the core domain. Cancer Cell 15, 376–388 (2009).
Bullock, A. N. et al. Thermodynamic stability of wild-type and mutant p53 core domain. Proc. Natl Acad. Sci. USA 94, 14338–14342 (1997).
Friedlander, P., Legros, Y., Soussi, T. & Prives, C. Regulation of mutant p53 temperature-sensitive DNA binding. J. Biol. Chem. 271, 25468–25478 (1996).
Michalovitz, D., Halevy, O. & Oren, M. Conditional inhibition of transformation and of cell proliferation by a temperature-sensitive mutant of p53. Cell 62, 671–680 (1990).
Joerger, A. C. & Fersht, A. R. Structure-function-rescue: the diverse nature of common p53 cancer mutants. Oncogene 26, 2226–2242 (2007).
Scotcher, J. et al. Identification of Two Reactive Cysteine Residues in the Tumor Suppressor Protein p53 Using Top-Down FTICR Mass Spectrometry. J. Am. Soc. Mass Spectr. 22, 888–897 (2011).
Wassman, C. D. et al. Computational identification of a transiently open L1/S3 pocket for reactivation of mutant p53. Nat. Commun. 4, 1407 (2013).
Brachmann, R. K., Yu, K., Eby, Y., Pavletich, N. P. & Boeke, J. D. Genetic selection of intragenic suppressor mutations that reverse the effect of common p53 cancer mutations. EMBO J. 17, 1847–1859 (1998).
Baroni, T. E. et al. A global suppressor motif for p53 cancer mutants. Proc. Natl Acad. Sci. USA 101, 4930–4935 (2004).
Friedler, A. et al. A peptide that binds and stabilizes p53 core domain: chaperone strategy for rescue of oncogenic mutants. Proc. Natl Acad. Sci. USA 99, 937–942 (2002).
Legros, Y., Meyer, A., Ory, K. & Soussi, T. Mutations in p53 produce a common conformational effect that can be detected with a panel of monoclonal antibodies directed toward the central part of the p53 protein. Oncogene 9, 3689–3694 (1994).
Khoo, K. H., Verma, C. S. & Lane, D. P. Drugging the p53 pathway: understanding the route to clinical efficacy. Nat. Rev. Drug Discov. 13, 217–236 (2014).
Deneberg, S. et al. An open-label phase I dose-finding study of APR-246 in hematological malignancies. Blood Cancer J. 6, e447 (2016).
Bykov, V. J. et al. Reactivation of mutant p53 and induction of apoptosis in human tumor cells by maleimide analogs. J. Biol. Chem. 280, 30384–30391 (2005).
Zache, N. et al. Mutant p53 targeting by the low molecular weight compound STIMA-1. Mol. Oncol. 2, 70–80 (2008).
Acknowledgements
This work was supported by the Swedish Research Council (VR), Cancerfonden, Radiumhemmets Forskningsfonder, Åke Wiberg Stiftelse, The Strategic Research Program in Cancer, and Karolinska Institutet. We thank Prof. Thierry Soussi, Karolinska Institutet, for the HO3.5 antibody, and the Protein Science Facility (PSF) and Proteomics Karolinska, Karolinska Institutet, for valuable help with protein purification and mass spectrometry.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
K.G.W. and V.J.N.B. are co-founders and shareholders of Aprea Therapeutics AB, a company that develops p53-based cancer therapy including APR-246. K.G.W. is a member of its Clinical Advisory Board. Research in the K.G.W. lab has received financial support from Aprea Therapeutics AB. K.G.W. has received a salary from Aprea Therapeutics AB.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, Q., Bykov, V.J.N., Wiman, K.G. et al. APR-246 reactivates mutant p53 by targeting cysteines 124 and 277. Cell Death Dis 9, 439 (2018). https://doi.org/10.1038/s41419-018-0463-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0463-7
This article is cited by
-
p53 biology and reactivation for improved therapy in MDS and AML
Biomarker Research (2024)
-
Translating p53-based therapies for cancer into the clinic
Nature Reviews Cancer (2024)
-
Pharmacological reactivation of p53 in the era of precision anticancer medicine
Nature Reviews Clinical Oncology (2024)
-
50 years on and still very much alive: ‘Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics’
British Journal of Cancer (2023)
-
The effects of free Cys residues on the structure, activity, and tetrameric stability of mammalian uricase
Applied Microbiology and Biotechnology (2023)