Abstract
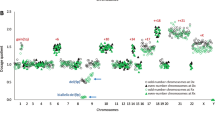
We have carried out a high-resolution whole genome DNA profiling analysis on 100 bone marrow samples from a consecutive series of de novo acute myeloid leukemia (AML) cases. After discarding copy number changes that are known to be genetic polymorphisms, we found that genomic aberrations (GA) in the form of gains or losses of genetic material were present in 74% of the samples, with a median of 2 GA per case (range 0–35). In addition to the cytogenetically detected aberration, GA were present in cases from all cytogenetic prognostic groups: 79% in the favorable group, 60% in the intermediate group (including 59% of cases with normal karyotype) and 83% in the adverse group. Five aberrant deleted regions were recurrently associated with cases with a highly aberrant genome (e.g., a 1.5 Mb deletion at 17q11.2 and a 750 kb deletion at 5q31.1). Different degrees of genomic instability showed a statistically significant impact on survival curves, even within the normal karyotype cases. This association was independent of other clinical and genetic parameters. Our study provides, for the first time, a detailed picture of the nature and frequency of DNA copy number aberrations in de novo AML.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Lowenberg B, Downing JR, Burnett A . Acute myeloid leukemia. N Engl J Med 1999; 341: 1051–1062.
Bacher U, Kern W, Schnittger S, Hiddemann W, Schoch C, Haferlach T . Further correlations of morphology according to FAB and WHO classification to cytogenetics in de novo acute myeloid leukemia: a study on 2,235 patients. Ann Hematol 2005; 84: 785–791.
Grimwade D, Walker H, Oliver F, Wheatley K, Harrison C, Harrison G et al. The importance of diagnostic cytogenetics on outcome in AML: analysis of 1,612 patients entered into the MRC AML 10 trial. The Medical Research Council Adult and Children's Leukaemia Working Parties. Blood 1998; 92: 2322–2333.
Byrd JC, Mrozek K, Dodge RK, Carroll AJ, Edwards CG, Arthur DC et al. Pretreatment cytogenetic abnormalities are predictive of induction success, cumulative incidence of relapse, and overall survival in adult patients with de novo acute myeloid leukemia: results from Cancer and Leukemia Group B (CALGB 8461). Blood 2002; 100: 4325–4336.
Mrozek K, Heerema NA, Bloomfield CD . Cytogenetics in acute leukemia. Blood Rev 2004; 18: 115–136.
Smith M, Barnett M, Bassan R, Gatta G, Tondini C, Kern W . Adult acute myeloid leukaemia. Crit Rev Oncol Hematol 2004; 50: 197–222.
Wolman SR, Gundacker H, Appelbaum FR, Slovak ML . Impact of trisomy 8 (+8) on clinical presentation, treatment response, and survival in acute myeloid leukemia: a Southwest Oncology Group study. Blood 2002; 100: 29–35.
Kelly L . Genetics of Myeloid Leukemias. Annu Rev Genomics Hum Genet 2002; 3: 179–198.
Stirewalt DL, Kopecky KJ, Meshinchi S, Engel JH, Pogosova-Agadjanyan EL, Linsley J et al. Size of FLT3 internal tandem duplication has prognostic significance in patients with acute myeloid leukemia. Blood 2006; 107: 3724–3726.
Frohling S, Schlenk RF, Breitruck J, Benner A, Kreitmeier S, Tobis K et al. Prognostic significance of activating FLT3 mutations in younger adults (16 to 60 years) with acute myeloid leukemia and normal cytogenetics: a study of the AML Study Group Ulm. Blood 2002; 100: 4372–4380.
Schnittger S, Schoch C, Kern W, Mecucci C, Tschulik C, Martelli MF et al. Nucleophosmin gene mutations are predictors of favorable prognosis in acute myelogenous leukemia with a normal karyotype. Blood 2005; 106: 3733–3739.
Suzuki T, Kiyoi H, Ozeki K, Tomita A, Yamaji S, Suzuki R et al. Clinical characteristics and prognostic implications of NPM1 mutations in acute myeloid leukemia. Blood 2005; 106: 2854–2861.
Haferlach T, Kohlmann A, Schnittger S, Dugas M, Hiddemann W, Kern W et al. Global approach to the diagnosis of leukemia using gene expression profiling. Blood 2005; 106: 1189–1198.
Virtaneva K, Wright FA, Tanner SM, Yuan B, Lemon WJ, Caligiuri MA et al. Expression profiling reveals fundamental biological differences in acute myeloid leukemia with isolated trisomy 8 and normal cytogenetics. Proc Natl Acad Sci USA 2001; 98: 1124–1129.
Raghavan M, Lillington DM, Skoulakis S, Debernardi S, Chaplin T, Foot NJ et al. Genome-wide single nucleotide polymorphism analysis reveals frequent partial uniparental disomy due to somatic recombination in acute myeloid leukemias. Cancer Res 2005; 65: 375–378.
Martinez-Ramirez A, Urioste M, Calasanz MJ, Cigudosa JC, Benitez J . Array comparative genomic hybridization analysis of myelodysplastic syndromes with complex karyotypes. A technical evaluation. Cancer Genet Cytogenet 2003; 144: 87–89.
Martinez-Ramirez A, Urioste M, Melchor L, Blesa D, Valle L, de Andres SA et al. Analysis of myelodysplastic syndromes with complex karyotypes by high-resolution comparative genomic hybridization and subtelomeric CGH array. Genes Chromosomes Cancer 2005; 42: 287–298.
Paulsson K, Heidenblad M, Strombeck B, Staaf J, Jonsson G, Borg A et al. High-resolution genome-wide array-based comparative genome hybridization reveals cryptic chromosome changes in AML and MDS cases with trisomy 8 as the sole cytogenetic aberration. Leukemia 2006; 20: 840–846.
Rucker FG, Bullinger L, Schwaenen C, Lipka DB, Wessendorf S, Frohling S et al. Disclosure of Candidate Genes in Acute Myeloid Leukemia With Complex Karyotypes Using Microarray-Based Molecular Characterization. J Clin Oncol 2006; 24: 3887–3894.
Barrett MT, Scheffer A, Ben-Dor A, Sampas N, Lipson D, Kincaid R et al. Comparative genomic hybridization using oligonucleotide microarrays and total genomic DNA. Proc Natl Acad Sci USA 2004; 101: 17765–17770.
Kiyoi H, Naoe T, Nakano Y, Yokota S, Minami S, Miyawaki S et al. Prognostic implication of FLT3 and N-RAS gene mutations in acute myeloid leukemia. Blood 1999; 93: 3074–3080.
Suarez L, Vidriales MB, Moreno MJ, Lopez A, Garcia-Larana J, Perez-Lopez C et al. Differences in anti-apoptotic and multidrug resistance phenotypes in elderly and young acute myeloid leukemia patients are related to the maturation of blast cells. Haematologica 2005; 90: 54–59.
Vaquerizas JM, Conde L, Yankilevich P, Cabezon A, Minguez P, Diaz-Uriarte R et al. GEPAS, an experiment-oriented pipeline for the analysis of microarray gene expression data. Nucleic Acids Res 2005; 33 (Web Server issue): W616–W620.
The Database of Genomic Variants 2005, Accession Date: 20th December 2006. http://projects.tcag.ca/variation.
Carrasco DR, Tonon G, Huang Y, Zhang Y, Sinha R, Feng B et al. High-resolution genomic profiles define distinct clinico-pathogenetic subgroups of multiple myeloma patients. Cancer Cell 2006; 9: 313–325.
Cigudosa JC, Odero MD, Calasanz MJ, Sole F, Salido M, Arranz E et al. De novo erythroleukemia chromosome features include multiple rearrangements, with special involvement of chromosomes 11 and 19. Genes Chromosomes Cancer 2003; 36: 406–412.
Wimmer K, Yao S, Claes K, Kehrer-Sawatzki H, Tinschert S, De Raedt T et al. Spectrum of single- and multiexon NF1 copy number changes in a cohort of 1,100 unselected NF1 patients. Genes Chromosomes Cancer 2006; 45: 265–276.
Farag SS, Archer KJ, Mrozek K, Ruppert AS, Carroll AJ, Vardiman JW et al. Pretreatment cytogenetics add to other prognostic factors predicting complete remission and long-term outcome in patients 60 years of age or older with acute myeloid leukemia: results from cancer and leukemia group B 8461. Blood 2006; 108: 63–73.
Wilson CS, Davidson GS, Martin SB, Andries E, Potter J, Harvey R et al. Gene expression profiling of adult acute myeloid leukemia identifies novel biologic clusters for risk classification and outcome prediction. Blood 2006; 108: 685–696.
Bullinger L, Dohner K, Bair E, Frohling S, Schlenk RF, Tibshirani R et al. Use of gene-expression profiling to identify prognostic subclasses in adult acute myeloid leukemia. N Engl J Med 2004; 350: 1605–1616.
Avivi I, Rowe JM . Prognostic factors in acute myeloid leukemia. Curr Opin Hematol 2005; 12: 62–67.
Galm O, Wilop S, Luders C, Jost E, Gehbauer G, Herman JG et al. Clinical implications of aberrant DNA methylation patterns in acute myelogenous leukemia. Ann Hematol 2005; 84 (Suppl 13): 39–49.
Alvarez S, Cigudosa JC . Gains, losses and complex karyotypes in myeloid disorders: a light at the end of the tunnel. Hematol Oncol 2005; 23: 18–25.
Trost D, Hildebrandt B, Beier M, Muller N, Germing U, Royer-Pokora B . Molecular cytogenetic profiling of complex karyotypes in primary myelodysplastic syndromes and acute myeloid leukemia. Cancer Genet Cytogenet 2006; 165: 51–63.
Tyybakinoja A, Elonen E, Piippo K, Porkka K, Knuutila S . Oligonucleotide array-CGH reveals cryptic gene copy number alterations in karyotypically normal acute myeloid leukemia. Leukemia 2007; 21: 571–574.
Gorletta TA, Gasparini P, D'Elios MM, Trubia M, Pelicci PG, Di Fiore PP . Frequent loss of heterozygosity without loss of genetic material in acute myeloid leukemia with a normal karyotype. Genes Chromosomes Cancer 2005; 44: 334–337.
Mitelman F JBaMFE . Mitelman Database of Chromosome Aberrations in Cancer. (2005), Accession Date: 2006. http://cgap.nci.nih.gov/Chromosomes/Mitelman.
Sanderson RN, Johnson PR, Moorman AV, Roman E, Willett E, Taylor PR et al. Population-based demographic study of karyotypes in 1709 patients with adult acute myeloid leukemia. Leukemia 2006; 20: 444–450.
Blaveri E, Brewer JL, Roydasgupta R, Fridlyand J, DeVries S, Koppie T et al. Bladder cancer stage and outcome by array-based comparative genomic hybridization. Clin Cancer Res 2005; 11: 7012–7022.
Fridlyand J, Snijders AM, Ylstra B, Li H, Olshen A, Segraves R et al. Breast tumor copy number aberration phenotypes and genomic instability. BMC Cancer 2006; 6: 96.
Monzo M, Brunet S, Urbano-Ispizua A, Navarro A, Perea G, Esteve J et al. Genomic polymorphisms provide prognostic information in intermediate-risk acute myeloblastic leukemia. Blood 2006; 107: 4871–4879.
Beyer V, Castagne C, Muhlematter D, Parlier V, Gmur J, Hess U et al. Systematic screening at diagnosis of -5/del(5)(q31), -7, or chromosome 8 aneuploidy by interphase fluorescence in situ hybridization in 110 acute myelocytic leukemia and high-risk myelodysplastic syndrome patients: concordances and discrepancies with conventional cytogenetics. Cancer Genet Cytogenet 2004; 152: 29–41.
Chowdhury D, Keogh MC, Ishii H, Peterson CL, Buratowski S, Lieberman J . gamma-H2AX dephosphorylation by protein phosphatase 2A facilitates DNA double-strand break repair. Mol Cell 2005; 20: 801–809.
Reynolds RM, Browning GG, Nawroz I, Campbell IW . Von Recklinghausen's neurofibromatosis: neurofibromatosis type 1. Lancet 2003; 361: 1552–1554.
Stephens K, Weaver M, Leppig KA, Maruyama K, Emanuel PD, Le Beau MM et al. Interstitial uniparental isodisomy at clustered breakpoint intervals is a frequent mechanism of NF1 inactivation in myeloid malignancies. Blood 2006; 108: 1684–1689.
Lu D, Nounou R, Beran M, Estey E, Manshouri T, Kantarjian H et al. The prognostic significance of bone marrow levels of neurofibromatosis-1 protein and ras oncogene mutations in patients with acute myeloid leukemia and myelodysplastic syndrome. Cancer 2003; 97: 441–449.
Chao RC, Pyzel U, Fridlyand J, Kuo YM, Teel L, Haaga J et al. Therapy-induced malignant neoplasms in Nf1 mutant mice. Cancer Cell 2005; 8: 337–348.
Yang G, Khalaf W, van de Locht L, Jansen JH, Gao M, Thompson MA et al. Transcriptional repression of the Neurofibromatosis-1 tumor suppressor by the t(8;21) fusion protein. Mol Cell Biol 2005; 25: 5869–5879.
Crescenzi B, La Starza R, Romoli S, Beacci D, Matteucci C, Barba G et al. Submicroscopic deletions in 5q- associated malignancies. Haematologica 2004; 89: 281–285.
Acknowledgements
We thank our colleagues Carmen Mateos (Hospital Virgen del Camino, Pamplona), José Rifon (Clinica Universitaria de Navarra, Pamplona), Maria Jose Najera (Hospital San Millan, Logroño), Filomena Floristan (Hospital Cruces, Bilbao), Maite Zudaire (Hospital de Navarra Pamplona), Gerardo Hermida (Hospital General Yague, Burgos), Angel Pereda (Hospital Santiago, Vitoria), Gemma Azaceta (Hospital Clinico Lozano Blesa, Zaragoza) and Alfonso G. Pineda (Hospital U. Gregorio Marañon, Madrid) for providing samples and clinical information. We also thank M. José Larrayoz and Iria Gonzalez for managing DNA samples; Joaquin Dopazo (Centro Investigacion Principe Felipe, Valencia) for bioinformatic advice; M Carmen Martín, Carmen Carralero and Gloria Soler for their technical support; and Roger Milne (CeGen, CNIO, Madrid) for his help with the multivariate statistical analysis.
Research grants and financial support: This work was partially financed by grant PI040555 from Fondo Investigaciones Sanitarias, Ministerio de Sanidad y Consumo to JCC and a grant from Fundacion Cientifica de la Asociación Española contra el Cáncer to MDO. JS has a PhD fellowship from Ministerio de Educacion y Ciencia. CL has a PhD fellowship from Gobierno de Navarra. BF has a Marie Curie PhD Early Stage Research Training Fellowship.
Contributions: JS, FC, CL and BF performed the laboratory work, JS, SA, DB, and JCC analyzed the data; MA, RG, JAM, MDO and MJC contributed vital new reagents, and collected clinical data; JCC, MJC and SA designed the study; JS and JCC wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Leukemia website (http://www.nature.com/leu)
Supplementary information
Rights and permissions
About this article
Cite this article
Suela, J., Álvarez, S., Cifuentes, F. et al. DNA profiling analysis of 100 consecutive de novo acute myeloid leukemia cases reveals patterns of genomic instability that affect all cytogenetic risk groups. Leukemia 21, 1224–1231 (2007). https://doi.org/10.1038/sj.leu.2404653
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.leu.2404653
Keywords
This article is cited by
-
Array-based comparative genomic hybridization detects copy number variations with prognostic relevance in 80% of ALL with normal karyotype or failed chromosome analysis
Leukemia (2016)
-
Submicroscopic deletion of 5q involving tumor suppressor genes (CTNNA1, HSPA9) and copy neutral loss of heterozygosity associated with TET2 and EZH2 mutations in a case of MDS with normal chromosome and FISH results
Molecular Cytogenetics (2014)
-
Evidence-based genomic diagnosis characterized chromosomal and cryptic imbalances in 30 elderly patients with myelodysplastic syndrome and acute myeloid leukemia
Molecular Cytogenetics (2011)
-
Identification of copy number alterations by array comparative genomic hybridization in patients with late chronic or accelerated phase chronic myeloid leukemia treated with imatinib mesylate
International Journal of Hematology (2011)
-
Identification of acquired copy number alterations and uniparental disomies in cytogenetically normal acute myeloid leukemia using high-resolution single-nucleotide polymorphism analysis
Leukemia (2010)