Abstract
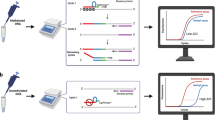
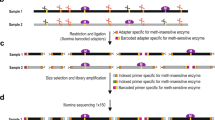
Methylation-specific polymerase chain reaction (PCR) (MSP) is frequently used to study gene silencing by promoter hypermethylation. However, non-specific primer design can lead to false-positive detection of methylation. We present a novel, web-based algorithm for the design of primers for bisulfite-PCRs (MSP, sequencing, COBRA and multiplex-MSP), allowing the determination of a specificity score, which is based on the thermodynamic characteristics of the primer 3′-end. PCR amplification with primers not reaching a high specificity score can result in false-positive findings. We used MSPprimer to design MSP primers for analysis of the ATM promoter. In 37 non-small cell lung cancer (NSCLC) samples and 43 breast cancer samples no promoter methylation was detected. Conversely, published MSP primers not reaching the required specificity score led to non-specific amplification of DNA not converted by bisulfite. The result was a false-positive incidence of ATM promoter methylation of 24% in NSCLC and 48% in breast cancers, similar to published studies. This highlights the critical need for specific primer design for MSP. MSPprimer is a convenient tool to achieve this goal, which is available free of charge to the scientific community.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ai L, Vo QN, Zuo C, Li L, Ling W, Suen JY et al. (2004). Ataxia-telangiectasia-mutated (ATM) gene in head and neck squamous cell carcinoma: promoter hypermethylation with clinical correlation in 100 cases. Cancer Epidemiol Biomarkers Prev 13: 150–156.
Allawi HT, SantaLucia Jr J . (1997). Thermodynamics and NMR of internal G. T mismatches in DNA. Biochemistry 36: 10581–10594.
Allawi HT, SantaLucia Jr J . (1998a). Nearest-neighbor thermodynamics of internal A.C mismatches in DNA: sequence dependence and pH effects. Biochemistry 37: 9435–9444.
Allawi HT, SantaLucia Jr J . (1998b). Nearest neighbor thermodynamic parameters for internal G. A mismatches in DNA. Biochemistry 37: 2170–2179.
Allawi HT, SantaLucia Jr J . (1998c). Thermodynamics of internal C.T mismatches in DNA. Nucleic Acids Res 26: 2694–2701.
Allinen M, Peri L, Kujala S, Lahti-Domenici J, Outila K, Karppinen SM et al. (2002). Analysis of 11q21–24 loss of heterozygosity candidate target genes in breast cancer: indications of TSLC1 promoter hypermethylation. Genes Chromosomes Cancer 34: 384–389.
Austen B, Powell JE, Alvi A, Edwards I, Hooper L, Starczynski J et al. (2005). Mutations in the ATM gene lead to impaired overall and treatment-free survival that is independent of IGVH mutation status in patients with B-CLL. Blood 106: 3175–3182.
Belinsky SA, Klinge DM, Dekker JD, Smith MW, Bocklage TJ, Gilliland FD et al. (2005). Gene promoter methylation in plasma and sputum increases with lung cancer risk. Clin Cancer Res 11: 6505–6511.
Bolt J, Vo QN, Kim WJ, McWhorter AJ, Thomson J, Hagensee ME et al. (2005). The ATM/p53 pathway is commonly targeted for inactivation in squamous cell carcinoma of the head and neck (SCCHN) by multiple molecular mechanisms. Oral Oncol 41: 1013–1020.
Brandes JC, Herman JG . (2006). p53 expression after treatment with zebularine is not due to demethylation. Cancer Res 66: 6892; author reply 6893.
Brandes JC, van Engeland M, Wouters KA, Weijenberg MP, Herman JG . (2005). CHFR promoter hypermethylation in colon cancer correlates with the microsatellite instability phenotype. Carcinogenesis 26: 1152–1156.
Clark SJ, Harrison J, Paul CL, Frommer M . (1994). High sensitivity mapping of methylated cytosines. Nucleic Acids Res 22: 2990–2997.
Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, Hidalgo OF, Vanaclocha V et al. (2000). Inactivation of the DNA-Repair Gene MGMT and the Clinical Response of Gliomas to Alkylating Agents. N Engl J Med 343: 1350–1354.
Esteller M, Sanchez-Cespedes M, Rosell R, Sidransky D, Baylin SB, Herman JG . (1999). Detection of aberrant promoter hypermethylation of tumor suppressor genes in serum DNA from non-small cell lung cancer patients. Cancer Res 59: 67–70.
Frommer M, McDonald LE, Millar DS, Collis CM, Watt F, Grigg GW et al. (1992). A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc Natl Acad Sci USA 89: 1827–1831.
Gronbaek K, Worm J, Ralfkiaer E, Ahrenkiel V, Hokland P, Guldberg P . (2002). ATM mutations are associated with inactivation of the ARF-TP53 tumor suppressor pathway in diffuse large B-cell lymphoma. Blood 100: 1430–1437.
Gumy-Pause F, Wacker P, Maillet P, Betts DR, Sappino AP . (2006). ATM promoter analysis in childhood lymphoid malignancies: a brief communication. Leuk Res 30: 335–337.
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M et al. (2005). MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med 352: 997–1003.
Herman JG, Baylin SB . (2003). Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 349: 2042–2054.
Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB . (1996). Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 93: 9821–9826.
Karpf AR, Moore BC, Ririe TO, Jones DA . (2001). Activation of the p53 DNA damage response pathway after inhibition of DNA methyltransferase by 5-aza-2′-deoxycytidine. Mol Pharmacol 59: 751–757.
Kim WJ, Vo QN, Shrivastav M, Lataxes TA, Brown KD . (2002). Aberrant methylation of the ATM promoter correlates with increased radiosensitivity in a human colorectal tumor cell line. Oncogene 21: 3864–3871.
Li LC, Dahiya R . (2002). MethPrimer: designing primers for methylation PCRs. Bioinformatics 18: 1427–1431.
Luo L, Lu FM, Hart S, Foroni L, Rabbani H, Hammarstrom L et al. (1998). Ataxia-telangiectasia and T-cell leukemias: no evidence for somatic ATM mutation in sporadic T-ALL or for hypermethylation of the ATM-NPAT/E14 bidirectional promoter in T-PLL. Cancer Res 58: 2293–2297.
Miura F, Uematsu C, Sakaki Y, Ito T . (2005). A novel strategy to design highly specific PCR primers based on the stability and uniqueness of 3′-end subsequences. Bioinformatics 21: 4363–4370.
Owczarzy R, You Y, Moreira BG, Manthey JA, Huang L, Behlke MA et al. (2004). Effects of sodium ions on DNA duplex oligomers: improved predictions of melting temperatures. Biochemistry 43: 3537–3554.
Pellise M, Castells A, Gines A, Agrelo R, Sole M, Castellvi-Bel S et al. (2004). Detection of lymph node micrometastases by gene promoter hypermethylation in samples obtained by endosonography-guided fine-needle aspiration biopsy. Clin Cancer Res 10: 4444–4449.
Peyret N, Seneviratne PA, Allawi HT, SantaLucia Jr J . (1999). Nearest-neighbor thermodynamics and NMR of DNA sequences with internal A.A, C.C, G.G, and T.T mismatches. Biochemistry 38: 3468–3477.
Roy K, Wang L, Makrigiorgos GM, Price BD . (2006). Methylation of the ATM promoter in glioma cells alters ionizing radiation sensitivity. Biochem Biophys Res Commun 344: 821–826.
Safar AM, Spencer III H, Su X, Coffey M, Cooney CA, Ratnasinghe LD et al. (2005). Methylation profiling of archived non-small cell lung cancer: a promising prognostic system. Clin Cancer Res 11: 4400–4405.
SantaLucia Jr J, Allawi HT, Seneviratne PA . (1996). Improved nearest-neighbor parameters for predicting DNA duplex stability. Biochemistry 35: 3555–3562.
Stresemann C, Brueckner B, Musch T, Stopper H, Lyko F . (2006). Functional diversity of DNA methyltransferase inhibitors in human cancer cell lines. Cancer Res 66: 2794–2800.
Tusnady GE, Simon I, Varadi A, Aranyi T . (2005). BiSearch: primer-design and search tool for PCR on bisulfite-treated genomes. Nucleic Acids Res 33: e9.
Vallone PM, Butler JM . (2004). AutoDimer: a screening tool for primer-dimer and hairpin structures. Biotechniques 37: 226–231.
Vo QN, Kim WJ, Cvitanovic L, Boudreau DA, Ginzinger DG, Brown KD . (2004). The ATM gene is a target for epigenetic silencing in locally advanced breast cancer. Oncogene 23: 9432–9437.
Xiong Z, Laird PW . (1997). COBRA: a sensitive and quantitative DNA methylation assay. Nucleic Acids Res 25: 2532–2534.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brandes, J., Carraway, H. & Herman, J. Optimal primer design using the novel primer design program: MSPprimer provides accurate methylation analysis of the ATM promoter. Oncogene 26, 6229–6237 (2007). https://doi.org/10.1038/sj.onc.1210433
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1210433
Keywords
This article is cited by
-
Technical considerations in PCR-based assay design for diagnostic DNA methylation cancer biomarkers
Clinical Epigenetics (2022)
-
Bisulfite profiling of the MGMT promoter and comparison with routine testing in glioblastoma diagnostics
Clinical Epigenetics (2022)
-
MSP-HTPrimer: a high-throughput primer design tool to improve assay design for DNA methylation analysis in epigenetics
Clinical Epigenetics (2016)
-
Quantitative methodology is critical for assessing DNA methylation and impacts on correlation with patient outcome
Clinical Epigenetics (2014)
-
A critical re-assessment of DNA repair gene promoter methylation in non-small cell lung carcinoma
Scientific Reports (2014)