Abstract
Galangin suppresses proliferation and induces apoptosis and autophagy in hepatocellular carcinoma (HCC) cells, but the precise mechanism is not clear. In this study, we demonstrated that galangin induced autophagy, enhanced the binding of SIRT1-LC3 and reduced the acetylation of endogenous LC3 in HepG2 cells. But this autophagy was inhibited by inactivation of SIRT1 meanwhile, galangin failed to reduce the acetylation of endogenous LC3 after SIRT1 was knocked-down. Collectively, these findings demonstrate a new mechanism by which galangin induces autophagy via the deacetylation of endogenous LC3 by SIRT1.
Similar content being viewed by others
Introduction
Galangin (3,5,7-trihydroxyflavone) is a flavonoid derived primarily from the rhizome Alpinia officinarum (Hance). It is used in traditional Chinese medicine and is contained in some foods1. Galangin likely undergoes oxidative metabolism mainly in the liver2. Galangin inhibits proliferation, induces apoptosis and promotes autophagy in hepatocellular carcinoma cells3,4,5,6,7,8, but the precise mechanism by which galangin induces autophagy remains unclear.
Autophagy is a self-digestion process, during which cytoplasmic components, such as protein aggregates, damaged organelles and lipid vesicles, are sequestered into double-membraned structures called autophagosomes. Autophagosomes then fuse with lysosomes for subsequent degradation of the content9,10,11,12.
The microtubule-associated protein 1 light chain 3 (LC3) is a key regulator of autophagy that controls major steps in the autophagic pathway, such as the growth of autophagic membranes, the recognition of autophagic cargoes and the fusion of autophagosomes with lysosomes13,14,15,16. The conversion of LC3 I to LC3 II via proteolytic cleavage and lipidation is a mark of mammalian autophagy12,17. Sequestosome 1 (p62/SQSTM1), which is degraded by autophagy, is another indicator of autophagy18,19. Other products of autophagy-related genes (ATGs), such as ATG3 and Beclin1 (also called ATG6/Vps30), also perform essential roles in autophagy20,21. Recent research has shown that the deacetylation of LC3, ATG5 and ATG7 by mammalian silent information regulator 2 homolog 1 (SIRT1) is essential for autophagosome formation under starvation conditions22,23.
SIRT1 is an NAD-dependent class III histone deacetylase that plays major roles in regulating gene expression, DNA damage repair, metabolism, tumor development, aging and autophagy22,23,24,25,26. SIRT1 promotes autophagy through the adenosine monophosphate-activated protein kinase (AMPK)/SIRT1 signaling pathway27 or via deacetylation of ATG5, ATG7, ATG8 (LC3)22,23 or forkhead box O1 (FOXO1)28.
In this study, we investigated the autophagy-inducing effect of galangin in HepG2 cells and explored the mechanism by which galangin induces autophagy via the deacetylation of endogenous LC3 by SIRT1.
Methods
Cell culture
The human liver cancer cell line HepG2 was obtained from the American Type Culture Collection (Rockville, Md., USA) and maintained at the Institute of Biochemistry and Molecular Biology at Guangdong Medical College. This cell line was cultured in Dulbecco’s modified Eagle’s medium (DMEM, Gibco BRL) supplemented with 10% fetal bovine serum (Sijiqing Laboratories, Hangzhou, China), 100 mg/mL penicillin and 100 mg/mL streptomycin. The cells were incubated at 37 °C in a humidified atmosphere containing 5% CO2.
Agents and chemicals
Galangin (PubChem CID: 5281616) was purchased from Sigma–Aldrich and dissolved in dimethyl sulfoxide (DMSO) before addition to the cell cultures. The final concentration of DMSO in the culture medium was kept below 0.1% (v/v) after the addition of galangin. TSA was purchased from Tokyo Chemical Industry. Rabbit or mouse polyclonal antibodies against Beclin1, LC3, acetylated lysine and GAPDH were purchased from Cell Signaling Technology. Rabbit or mouse polyclonal antibodies against SIRT1 and p62 were purchased from Abcam. A rabbit polyclonal antibody against β-actin was purchased from Beijing Biosynthesis Biotechnology.
Autophagy induction
To induce autophagy, HepG2 cells were treated with 130 μM galangin for 24 hours. For the co-immunoprecipitation experiments, HepG2 cells were treated with 130 μM galangin for 18 hours to induce autophagy. We repeated the experiments three times.
Transmission electron microscope
HepG2 cells were harvested by trypsinization, then washed twice with PBS, fixed with the buffer containing 3% glutaraldehyde and 0.1 M cacodylate and then re-fixed in osmium tetroxide. HepG2 cells were embedded in Epong and cut into sections with a thickness of 1.0 μm. Sections were stained with methylene buffer ArumeII and then viewed with a Philips electron microscope CM-120.
Inactivating SIRT1 and regulating its expression
EX-527 is a specific SIRT1 inhibitor that inhibits SIRT1 activity at a concentration of 50 μM22. Therefore, we treated HepG2 cells with 50 μM EX-527 for 2 hours to inhibit SIRT1 activity, induced autophagy by adding 130 μM galangin into the medium containing EX-527 and continued to cultivate the cells for 24 hours.
To regulate the expression of SIRT1, HepG2 cells were infected with relevant recombinant adenoviruses (Shanghai GeneChem Co., Ltd.) for 2 hours and then cultured in fresh complete medium for 24 hours. Next, the cells were treated with 130 μM galangin to induce autophagy. We repeated these experiments three times.
Detection of autophagy using GFP-LC3
The green fluorescent protein and LC3 (GFP-LC3) fusion protein provides a useful indicator of autophagy initiation through the evaluation of LC3 dots or punctae. When autophagy occurs, GFP-LC3 foci redistribute from a diffuse pattern to a punctate cytoplasmic pattern (GFP-LC3 punctae) and the percentage of cells with GFP-LC3 punctae increases29.
HepG2 cells were transfected with ptfLC3 (Addgene, Plasmid #21074), a highly specific fluorescent marker of autophagy, to measure autophagy levels. Lipofectamine® 3000 Reagent (Life Technologies) was used to transfect HepG2 cells. After the induction of autophagy with 130 μM galangin, the cellular localization of GFP-LC3 was visualized using a Nikon fluorescence microscope. We repeated these experiments three times.
Western blot and co-immunoprecipitation analysis
Western blot analysis was performed using whole cell extracts prepared by lysing the cells in lysis buffer (pH 8.0) containing 50 mM Tris–HCl, 150 mM NaCl, 5 mM EDTA, 1% NP40, 0.05% PMSF, 2 mg/mL aprotinin and 2 mg/mL leupeptin. Antibodies against GAPDH and β-actin were used to assess the purity of the respective fractions.
For the co-immunoprecipitation analyses, the cells were resuspended in Nonidet P40 lysis buffer containing 10 mM nicotinamide and 10 μM TSA. Cell lysates were mixed with antibodies at 4 °C overnight and PureProteome™ Protein A/G Mix Magnetic Beads (Millipore Corporation) were then added. The detailed protocol for this procedure can be found at www.millipore.com. The proteins were resolved by SDS-polyacrylamide gel electrophoresis (PAGE) and then transferred onto polyvinylidene fluoride membranes, which were blocked in 5% (w/v) nonfat milk and hybridized with specific primary antibodies. The resulting protein bands were visualized using ECL (Thermo Scientific) after hybridization with a secondary antibody (Thermo Scientific). All of the western blot results were quantified using Image J. We repeated these experiments three times.
Results
Galangin induced autophagy and upregulated SIRT1 expression in HepG2 cells
Our previous study demonstrated that 130 μM galangin induces autophagy in hepatocellular carcinoma cells5,7,8. In this study, we treated HepG2 cells with 130 μM galangin and then detected autophagy by transmission electron microscopy and western blot analysis. Results showed that galangin promoted the appearance of autophagic vacuoles, increased the expression of LC3 II, Beclin1 and the ratio of LC3 II to LC3 I and decreased the expression of p62 in a time-dependent manner (Fig. 1). The autophagy-inducing effects of galangin were consistent with those in our previous study5,7,8. During the induction of autophagy, galangin increased SIRT1 expression in a time-dependent manner (Fig. 1B).
Galangin induced autophagy and upregulated SIRT1 expression in HepG2 cells.
(A) HepG2 cells were treated with 130 μM galangin for 0, 6, 12 and 24 hours, then visualized by transmission electron microscopy (×5800). (B) The expression of SIRT1, Beclin1, p62 and LC3 were detected by western blot. (C) The relative (Rel.) levels of LC3 II/ LC3 I, **P < 0.01, n = 3.
EX-527 inhibited SIRT1 and Galangin-induced autophagy
To confirm that SIRT1 participates in galangin-induced autophagy, we pretreated HepG2 cells with the specific SIRT1 inhibitor EX-527 (50 μM) for 2 hours, then, cells were treated with 130 μM galangin for 24 hours. Results showed that EX-527 inhibited galangin-induced autophagic vacuolization (Fig. 2A) and GFP-LC3 punctae formation (Fig. 2B). EX-527 also blocked the conversion of endogenous LC3 I to LC3 II and restored the galangin-induced downregulation of p62 (Fig. 2C). These data suggest that galangin-induced autophagy is inhibited by EX-527 in HepG2 cells.
EX-527 inhibited SIRT1 and Galangin-induced autophagy.
(A) Autophagic vacuolization (indicated by red arrows) in HepG2 cells treated with EX-527 and galangin. Cells were visualized using a phase contrast microscope (×200). (B) Distribution of GFP-LC3 in HepG2 cells treated with EX-527 and galangin. Cells were transfected with GFP-LC3 exposed to different treatment conditions and visualized by fluorescent microscopy (×200). (C) LC3 and p62 expression in HepG2 cells exposed to 50 μM EX-527 and 130 μM galangin.
SIRT1 is essential to Galangin-induced autophagy
We infected HepG2 cells with relevant recombinant adenoviruses to knockdown or upregulate SIRT1. Then, cells were treated with 130 μM galangin for 24 hours to induce autophagy. Compared to vector-infected cells (Vector) and uninfected cells (Blank), the conversion of LC3 I to LC3 II and downregulation of p62 was blocked in SIRT1-knockdown cells treated with galangin (Fig. 3A), indicating that galangin-induced autophagy was inhibited by SIRT1 downregulation. In addition, overexpressing SIRT1 increased the conversion of LC3 I to LC3 II and reduced the expression of p62, indicating that overexpressing SIRT1 stimulated basal autophagy of HepG2 cells under normal fed condition (Fig. 3B).
SIRT1 is essential to Galangin-induced autophagy.
(A) LC3 and p62 expression in DMSO treated or 130 μM galangin-treated HepG2 cells under following conditions: uninfected control (Blank), SIRT1-knockdown (SIRT1-KD), vector-infected control for SIRT1-KD (Vector). (B) LC3 and p62 expression in DMSO treated or 130 μM galangin-treated HepG2 cells under following conditions: uninfected control (Blank), SIRT1 over-expression (SIRT1-AD) and vector-infected control for SIRT1-AD (Vector).
Deacetylation of endogenous LC3 by SIRT1 was essential for galangin-induced autophagy
To determine the mechanism by which SIRT1 promotes galangin-induced autophagy, we used co-immunoprecipitation assays to detect the binding of endogenous SIRT1 and LC3. Results showed that the binding of SIRT1 and LC3 was enhanced and the acetylation of endogenous LC3 was reduced after HepG2 cells were treated with 130 μM galangin for 18 hours (Fig. 4A).
Deacetylation of endogenous LC3 by SIRT1 was essential for galangin-induced autophagy.
(A) The binding of endogenous SIRT1 to endogenous LC3 and endogenous LC3 acetylation were detected using co-immunoprecipitation assays and western blot analysis in HepG2 cells treated with DMSO or 130 μM galangin for 18 hours. (B) Endogenous LC3 acetylation was detected by co-immunoprecipitation assays and western blot analysis after HepG2 cells were treated with DMSO or 130 μM galangin in SIRT1-KD, Vector and uninfected cells (Blank).
To further explore whether endogenous LC3 was deacetylated by SIRT1 in galangin-induced autophagy, we knocked down SIRT1 expression in HepG2 cells and then treated cells with galangin for 18 hours. In vector-infected cells and uninfected cells, galangin promoted the conversion of LC3 I to LC3 II and decreased the acetylation of LC3. But in SIRT1-knockdown cells, galangin failed to reduce the acetylation of LC3 (Fig. 4B). These results suggest that deacetylation of endogenous LC3 by SIRT1 is essential for galangin-induced autophagy in HepG2 cells.
In conclusion, this study demonstrates a new mechanism by which galangin induces autophagy via the deacetylation of endogenous LC3 by SIRT1 in HepG2 cells (Fig. 4).
Discussion
Many flavonoids, such as quercetin, sudachitin, kaempferol and luteolin, have been reported to upregulate SIRT1 expression or activate SIRT130,31,32,33. Furthermore, certain flavonoids, such as catechin, epicatechin, quercetin and myricetin, can induce autophagy34. However, the role of SIRT1 in flavonoid-induced autophagy had not been reported. In this study, we showed that SIRT1 expression was upregulated in a time-dependent manner in HepG2 cells during galangin-induced autophagy, which was inhibited when SIRT1 was inactivated or knocked down. In addition, overexpressing SIRT1 stimulated basal autophagy of HepG2 cells. Our data suggest that galangin, which has a simple flavonoid structure, can induce autophagy and upregulate SIRT1 in HepG2 cells and that SIRT1 is essential for galangin-induced autophagy. This study provides a connection between flavonoids, autophagy and SIRT1.
Then, we explored the mechanism by which SIRT1 regulates galangin-induced autophagy. Researches have demonstrated the relationship between SIRT1 and autophagy; for example, SIRT1 regulates autophagy via the deacetylation of several ATGs22,23 and FoxO128 and via the AMPK/SIRT1 signaling pathway27. However, the mechanism by which SIRT1 influences flavonoid-induced autophagy had not been reported. ATGs perform important roles in autophagy and the deacetylation of ATG5, ATG7 and ATG8 (LC3) by SIRT1 is necessary for the induction of starvation-induced autophagy22,23. Thus, we speculated that SIRT1 may participate in galangin-induced autophagy through deacetylation of ATG8 (LC3). In this study, we demonstrated that LC3-SIRT1 binding and endogenous LC3 deacetylation by SIRT1 play important roles in galangin-induced autophagy. These results provide a new mechanism by which galangin induces autophagy via the deacetylation of endogenous LC3 by SIRT1 in HepG2 cells.
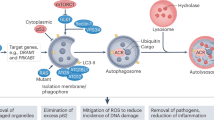
Based on previous experimental results, we demonstrate the mechanisms by which galangin induces autophagy as shown in Fig. 5. Galangin induces autophagy in hepatocellular carcinoma cells by activating the TGF-β receptor/Smad pathway8, by activating AMPK via increasing the AMP/TAN ratio7, by upregulating p535 and by deacetylating LC3. All these pathways have been reported to share a close relationship with SIRT1. AMPK is a key energy sensors of the cell. Simultaneously, SIRT1/AMPK signaling pathway has been reported to induce autophagy in oxidative stress35, glucose starvation36, chronic colitis37 and resveratrol treatment38,39,40. So galangin may induce glucose starvation and then activiates SIRT1/AMPK pathway, leads to autophagy finally. Further more, the deacetylation of p53 by SIRT1 is essential for capsaicin induced autophagy41. Zerr et al. have reported that SIRT1 can activate TGF-β/Smad pathway42. And then Smad3 also in turn activates SIRT1 transcription43. Basing on our results that galangin-induced autophagy was block by inactiviating or down-regulating SIRT1, we think that the autophagy-inducing effect of galangin may depend on SIRT1. As many flavonoids can activiate SIRT1 and induce aotuphagy, we believe that some kind of flavonoids may induce autophagy mainly through SIRT1.
Autophagy is a highly conserved process that involves degradation and recycling of proteins and organelles to generate nucleotides, amino acids, fatty acids, sugars and ATP to support metabolism and survival under adverse microenvironmental conditions11,44. Autophagy is thought to play dual roles in cancer because it enables the survival of tumor cells by promoting metabolite turnover and absorption, inhibiting apoptosis and reactive oxygen species production and increasing drug resistance45; in contrast, it can prevent tumor initiation by suppressing chronic tissue damage, inflammation and genome instability46 and many studies have shown that autophagy induction can contribute to apoptosis47,48,49. Similarly, SIRT1 has opposing effects in cancer. On the one hand, SIRT1 inactivates tumor suppressors, activates proto-oncogenes and promotes cancer cell proliferation, invasion, migration and chemoresistance, which confer a survival advantage to cancer cells50,51,52,53,54,55,56. On the other hand, SIRT1 suppresses tumors by inhibiting inflammation and the activity of transcription factors that exacerbate carcinogenesis and by preserving genomic stability57,58,59,60,61,62. In our study, galangin inhibited the growth of HepG2 cells mainly by potently inducing continuous autophagy, thereby inhibiting cell proliferation and inducing apoptosis7,8. Here, we demonstrated that SIRT1 is essential for galangin-induced autophagy, suggesting that SIRT1 and autophagy play an anti-cancer role under the treatment of galangin.
These findings suggest that the SIRT1-activating and autophagy-inducing functions of flavonoids are worth further exploration for cancer therapy, as are the autophagy-regulating functions of SIRT1. In conclusion, galangin induces autophagy via the deacetylation of LC3 by SIRT1 in HepG2 cells. Combined with the results of our previous reports, the current data suggest that galangin is a potential anti-cancer drug.
Additional Information
How to cite this article: Li, X. et al. Galangin Induces Autophagy via Deacetylation of LC3 by SIRT1 in HepG2 Cells. Sci. Rep. 6, 30496; doi: 10.1038/srep30496 (2016).
References
Devadoss, D., Ramar, M. & Chinnasamy, A. Galangin, a dietary flavonol inhibits tumor initiation during experimental pulmonary tumorigenesis by modulating xenobiotic enzymes and antioxidant status. Archives of pharmacal research, doi: 10.1007/s12272-014-0330-8 (2014).
Otake, Y. & Walle, T. Oxidation of the flavonoids galangin and kaempferide by human liver microsomes and CYP1A1, CYP1A2 and CYP2C9. Drug Metab Dispos 30, 103–105 (2002).
Zhang, H. T., Wu, J., Wen, M., Su, L. J. & Luo, H. Galangin induces apoptosis in hepatocellular carcinoma cells through the caspase 8/t-Bid mitochondrial pathway. J Asian Nat Prod Res 14, 626–633, doi: 10.1080/10286020.2012.682152 (2012).
Zhang, H.-T. Galangin induces apoptosis of hepatocellular carcinoma cells via the mitochondrial pathway. World Journal of Gastroenterology 16, 3377, doi: 10.3748/wjg.v16.i27.3377 (2010).
Wen, M., Wu, J., Luo, H. & Zhang, H. Galangin induces autophagy through upregulation of p53 in HepG2 cells. Pharmacology 89, 247–255, doi: 10.1159/000337041 (2012).
Su, L. et al. Galangin inhibits proliferation of hepatocellular carcinoma cells by inducing endoplasmic reticulum stress. Food and chemical toxicology : an international journal published for the British Industrial Biological Research Association 62, 810–816, doi: 10.1016/j.fct.2013.10.019 (2013).
Zhang, H. et al. Galangin inhibits proliferation of HepG2 cells by activating AMPK via increasing the AMP/TAN ratio in a LKB1-independent manner. European journal of pharmacology 718, 235–244, doi: 10.1016/j.ejphar.2013.08.026 (2013).
Wang, Y. et al. Galangin suppresses HepG2 cell proliferation by activating the TGF-beta receptor/Smad pathway. Toxicology 326, 9–17, doi: 10.1016/j.tox.2014.09.010 (2014).
Klionsky, D. J. Autophagy: from phenomenology to molecular understanding in less than a decade. Nat Rev Mol Cell Biol 8, 931–937, doi: 10.1038/nrm2245 (2007).
Mizushima, N. Autophagy: process and function. Genes Dev 21, 2861–2873, doi: 10.1101/gad.1599207 (2007).
Galluzzi, L., Pietrocola, F., Levine, B. & Kroemer, G. Metabolic control of autophagy. Cell 159, 1263–1276, doi: 10.1016/j.cell.2014.11.006 (2014).
Mizushima, N., Yoshimori, T. & Levine, B. Methods in mammalian autophagy research. Cell 140, 313–326, doi: 10.1016/j.cell.2010.01.028 (2010).
Pankiv, S. et al. FYCO1 is a Rab7 effector that binds to LC3 and PI3P to mediate microtubule plus end-directed vesicle transport. J Cell Biol 188, 253–269, doi: 10.1083/jcb.200907015 (2010).
Ichimura, Y. et al. A ubiquitin-like system mediates protein lipidation. Nature 408, 488–492, doi: 10.1038/35044114 (2000).
Nakatogawa, H., Ichimura, Y. & Ohsumi, Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell 130, 165–178, doi: 10.1016/j.cell.2007.05.021 (2007).
Noda, N. N., Ohsumi, Y. & Inagaki, F. Atg8-family interacting motif crucial for selective autophagy. FEBS Lett 584, 1379–1385, doi: 10.1016/j.febslet.2010.01.018 (2010).
Kuma, A. et al. The role of autophagy during the early neonatal starvation period. Nature 432, 1032–1036, doi: 10.1038/nature03029 (2004).
Komatsu, M. et al. Homeostatic levels of p62 control cytoplasmic inclusion body formation in autophagy-deficient mice. Cell 131, 1149–1163, doi: 10.1016/j.cell.2007.10.035 (2007).
Komatsu, M., Kageyama, S. & Ichimura, Y. p62/SQSTM1/A170: physiology and pathology. Pharmacol Res 66, 457–462, doi: 10.1016/j.phrs.2012.07.004 (2012).
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell 19, 2092–2100, doi: 10.1091/mbc.E07-12-1257 (2008).
Tanida, I., Tanida-Miyake, E., Ueno, T. & Kominami, E. The human homolog of Saccharomyces cerevisiae Apg7p is a Protein-activating enzyme for multiple substrates including human Apg12p, GATE-16, GABARAP and MAP-LC3. J Biol Chem 276, 1701–1706, doi: 10.1074/jbc.C000752200 (2001).
Huang, R. et al. Deacetylation of nuclear LC3 drives autophagy initiation under starvation. Molecular cell 57, 456–466, doi: 10.1016/j.molcel.2014.12.013 (2015).
Lee, I. H. et al. A role for the NAD-dependent deacetylase Sirt1 in the regulation of autophagy. Proceedings of the National Academy of Sciences 105, 3374–3379, doi: 10.1073/pnas.0712145105 (2008).
Haigis, M. C. & Sinclair, D. A. Mammalian sirtuins: biological insights and disease relevance. Annu Rev Pathol 5, 253–295, doi: 10.1146/annurev.pathol.4.110807.092250 (2010).
Ou, X. et al. SIRT1 deficiency compromises mouse embryonic stem cell hematopoietic differentiation and embryonic and adult hematopoiesis in the mouse. Blood 117, 440–450, doi: 10.1182/blood-2010-03-273011 (2011).
Chae, H. D. & Broxmeyer, H. E. SIRT1 deficiency downregulates PTEN/JNK/FOXO1 pathway to block reactive oxygen species-induced apoptosis in mouse embryonic stem cells. Stem Cells Dev 20, 1277–1285, doi: 10.1089/scd.2010.0465 (2011).
Jin, X. et al. Delphinidin-3-glucoside protects human umbilical vein endothelial cells against oxidized low-density lipoprotein-induced injury by autophagy upregulation via the AMPK/SIRT1 signaling pathway. Mol Nutr Food Res 58, 1941–1951, doi: 10.1002/mnfr.201400161 (2014).
Hariharan, N. et al. Deacetylation of FoxO by Sirt1 Plays an Essential Role in Mediating Starvation-Induced Autophagy in Cardiac Myocytes. Circulation Research 107, 1470–1482, doi: 10.1161/circresaha.110.227371 (2010).
Mizushima, N., Yamamoto, A., Matsui, M., Yoshimori, T. & Ohsumi, Y. In vivo analysis of autophagy in response to nutrient starvation using transgenic mice expressing a fluorescent autophagosome marker. Mol Biol Cell 15, 1101–1111, doi: 10.1091/mbc.E03-09-0704 (2004).
Rafacho, B. P. et al. Inhibition of diethylnitrosamine-initiated alcohol-promoted hepatic inflammation and precancerous lesions by flavonoid luteolin is associated with increased sirtuin 1 activity in mice. Hepatobiliary Surg Nutr 4, 124–134, doi: 10.3978/j.issn.2304-3881.2014.08.06 (2015).
Tsutsumi, R. et al. Sudachitin, a polymethoxylated flavone, improves glucose and lipid metabolism by increasing mitochondrial biogenesis in skeletal muscle. Nutr Metab (Lond) 11, 32, doi: 10.1186/1743-7075-11-32 (2014).
Casuso, R. A. et al. Quercetin supplementation does not enhance cerebellar mitochondrial biogenesis and oxidative status in exercised rats. Nutr Res 35, 585–591, doi: 10.1016/j.nutres.2015.05.007 (2015).
Guo, Z. et al. Kaempferol protects cardiomyocytes against anoxia/reoxygenation injury via mitochondrial pathway mediated by SIRT1. Eur J Pharmacol 761, 245–253, doi: 10.1016/j.ejphar.2015.05.056 (2015).
Pietrocola, F. et al. Pro-autophagic polyphenols reduce the acetylation of cytoplasmic proteins. Cell Cycle 11, 3851–3860, doi: 10.4161/cc.22027 (2012).
Han, X. et al. AMPK activation protects cells from oxidative stress-induced senescence via autophagic flux restoration and intracellular NAD(+) elevation. Aging cell 15, 416–427, doi: 10.1111/acel.12446 (2016).
Chang, C. et al. AMPK-Dependent Phosphorylation of GAPDH Triggers Sirt1 Activation and Is Necessary for Autophagy upon Glucose Starvation. Molecular cell 60, 930–940, doi: 10.1016/j.molcel.2015.10.037 (2015).
Talero, E. et al. Expression patterns of sirtuin 1-AMPK-autophagy pathway in chronic colitis and inflammation-associated colon neoplasia in IL-10-deficient mice. International immunopharmacology 35, 248–256, doi: 10.1016/j.intimp.2016.03.046 (2016).
Guo, H., Chen, Y., Liao, L. & Wu, W. Resveratrol protects HUVECs from oxidized-LDL induced oxidative damage by autophagy upregulation via the AMPK/SIRT1 pathway. Cardiovascular drugs and therapy / sponsored by the International Society of Cardiovascular Pharmacotherapy 27, 189–198, doi: 10.1007/s10557-013-6442-4 (2013).
Wang, X. H. et al. Resveratrol attenuated TNF-alpha-induced MMP-3 expression in human nucleus pulposus cells by activating autophagy via AMPK/SIRT1 signaling pathway. Experimental biology and medicine, doi: 10.1177/1535370216637940 (2016).
Wu, Y. et al. Resveratrol-activated AMPK/SIRT1/autophagy in cellular models of Parkinson’s disease. Neuro-Signals 19, 163–174, doi: 10.1159/000328516 (2011).
Lee, Y. H., Chen, H. Y., Su, L. J. & Chueh, P. J. Sirtuin 1 (SIRT1) Deacetylase Activity and NAD(+)/NADH Ratio Are Imperative for Capsaicin-Mediated Programmed Cell Death. Journal of agricultural and food chemistry 63, 7361–7370, doi: 10.1021/acs.jafc.5b02876 (2015).
Zerr, P. et al. Sirt1 regulates canonical TGF-beta signalling to control fibroblast activation and tissue fibrosis. Annals of the rheumatic diseases 75, 226–233, doi: 10.1136/annrheumdis-2014-205740 (2016).
Warburton, D., Shi, W. & Xu, B. TGF-beta-Smad3 signaling in emphysema and pulmonary fibrosis: an epigenetic aberration of normal development? American journal of physiology. Lung cellular and molecular physiology 304, L83–85, doi: 10.1152/ajplung.00258.2012 (2013).
Rabinowitz, J. D. & White, E. Autophagy and metabolism. Science 330, 1344–1348, doi: 10.1126/science.1193497 (2010).
Yang, S. et al. Pancreatic cancers require autophagy for tumor growth. Genes Dev 25, 717–729, doi: 10.1101/gad.2016111 (2011).
White, E. Deconvoluting the context-dependent role for autophagy in cancer. Nat Rev Cancer 12, 401–410, doi: 10.1038/nrc3262 (2012).
Law, B. Y. et al. Natural small-molecule enhancers of autophagy induce autophagic cell death in apoptosis-defective cells. Scientific reports 4, 5510, doi: 10.1038/srep05510 (2014).
Kumar, D., Shankar, S. & Srivastava, R. K. Rottlerin-induced autophagy leads to the apoptosis in breast cancer stem cells: molecular mechanisms. Molecular cancer 12, 171, doi: 10.1186/1476-4598-12-171 (2013).
He, P. X., Che, Y. S., He, Q. J., Chen, Y. & Ding, J. G226, a novel epipolythiodioxopiperazine derivative, induces autophagy and caspase-dependent apoptosis in human breast cancer cells in vitro. Acta pharmacologica Sinica 35, 1055–1064, doi: 10.1038/aps.2014.47 (2014).
Chen, W. Y. et al. Tumor suppressor HIC1 directly regulates SIRT1 to modulate p53-dependent DNA-damage responses. Cell 123, 437–448, doi: 10.1016/j.cell.2005.08.011 (2005).
Ikenoue, T., Inoki, K., Zhao, B. & Guan, K. L. PTEN acetylation modulates its interaction with PDZ domain. Cancer Res 68, 6908–6912, doi: 10.1158/0008-5472.CAN-08-1107 (2008).
Sundaresan, N. R. et al. The deacetylase SIRT1 promotes membrane localization and activation of Akt and PDK1 during tumorigenesis and cardiac hypertrophy. Sci Signal 4, ra46, doi: 10.1126/scisignal.2001465 (2011).
Orecchia, A. et al. Sirtinol treatment reduces inflammation in human dermal microvascular endothelial cells. PLoS One 6, e24307, doi: 10.1371/journal.pone.0024307 (2011).
Eades, G. et al. miR-200a regulates SIRT1 expression and epithelial to mesenchymal transition (EMT)-like transformation in mammary epithelial cells. J Biol Chem 286, 25992–26002, doi: 10.1074/jbc.M111.229401 (2011).
Zhang, Y. et al. Deacetylation of cortactin by SIRT1 promotes cell migration. Oncogene 28, 445–460, doi: 10.1038/onc.2008.388 (2009).
Chen, H. C., Jeng, Y. M., Yuan, R. H., Hsu, H. C. & Chen, Y. L. SIRT1 promotes tumorigenesis and resistance to chemotherapy in hepatocellular carcinoma and its expression predicts poor prognosis. Ann Surg Oncol 19, 2011–2019, doi: 10.1245/s10434-011-2159-4 (2012).
Yeung, F. et al. Modulation of NF-kappaB-dependent transcription and cell survival by the SIRT1 deacetylase. EMBO J 23, 2369–2380, doi: 10.1038/sj.emboj.7600244 (2004).
Nie, Y. et al. STAT3 inhibition of gluconeogenesis is downregulated by SirT1. Nat Cell Biol 11, 492–500, doi: 10.1038/ncb1857 (2009).
Gao, Z. & Ye, J. Inhibition of transcriptional activity of c-JUN by SIRT1. Biochem Biophys Res Commun 376, 793–796, doi: 10.1016/j.bbrc.2008.09.079 (2008).
Firestein, R. et al. The SIRT1 deacetylase suppresses intestinal tumorigenesis and colon cancer growth. PLoS One 3, e2020, doi: 10.1371/journal.pone.0002020 (2008).
Oberdoerffer, P. et al. SIRT1 redistribution on chromatin promotes genomic stability but alters gene expression during aging. Cell 135, 907–918, doi: 10.1016/j.cell.2008.10.025 (2008).
Palacios, J. A. et al. SIRT1 contributes to telomere maintenance and augments global homologous recombination. J Cell Biol 191, 1299–1313, doi: 10.1083/jcb.201005160 (2010).
Acknowledgements
This study was supported by grants from the National Natural Science Foundation of China (Grant Nos 81273549 and 81400023), Science and Technology Projects of Guangdong Province, China (No. 2014A020212292) and Medical Science and Technology Research Fund of Guangdong (No. A2015338).
Author information
Authors and Affiliations
Contributions
X.L. and Y.J.W. contributed to the study design, western blot assays, co-IP assays, data collection, data analysis and manuscript preparation. Y.Z.X. performed the plasmid transformations and the GFP-LC3 visualizations. J.W. contributed to the adenovirus infections. H.D., X.Y.C. and L.B.L. contributed to the discussion section and interpreted some of the results. H.Z. designed the study and revised the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Li, X., Wang, Y., Xiong, Y. et al. Galangin Induces Autophagy via Deacetylation of LC3 by SIRT1 in HepG2 Cells. Sci Rep 6, 30496 (2016). https://doi.org/10.1038/srep30496
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep30496
This article is cited by
-
Galangin, as a Potential Anticancer Agent
Revista Brasileira de Farmacognosia (2022)
-
Silibinin-induced autophagy mediated by PPARα-sirt1-AMPK pathway participated in the regulation of type I collagen-enhanced migration in murine 3T3-L1 preadipocytes
Molecular and Cellular Biochemistry (2019)
-
The photocytotoxicity effect of cationic sulfonated corrole towards lung cancer cells: in vitro and in vivo study
Lasers in Medical Science (2019)
-
Autophagic flux is essential for the downregulation of D-dopachrome tautomerase by atractylenolide I to ameliorate intestinal adenoma formation
Journal of Cell Communication and Signaling (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.