Abstract
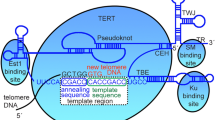
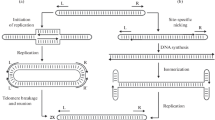
The terminal structure of the linear mitochondrial DNA (mtDNA) from the yeast Candida parapsilosis was investigated. This mtDNA, 30 kb long, has symmetrical ends forming inverted terminal repeats. These repeats are made up of a variable number of tandemly repeating units of 738 by each; the terminal nucleotide corresponds to a precise position within the last repeat unit sequence. The ends had an open structure accessible to enzymes, with a 5′ single-stranded extension of about 110 nucleotides. No circular forms were detected in the DNA preparations. Two other unrelated species, Pichia philodendra and Candida salmanticensis also appear to have a linear mtDNA of similar organization. These linear DNAs (which we name Type 2 linear mtDNAs) are distinct from the previously described linear mtDNAs of yeasts whose termini are formed by a closed hairpin loop (Type 1 linear mtDNA). The terminal structure of C. parapsilosis mtDNA is reminiscent of the linear mitochondrial genomes of the ciliate Tetrahymena although, in the latter, the telomeric tandem repeat unit is considerably shorter.
Similar content being viewed by others
References
Allen NE, MacQuillan AM (1969) Target analysis of mitochondrial genetic units in yeast. J Bacteriol 97:1142–1148
Baroudy BM, Venkatesan S, Moss B (1982) Incompletely basepaired flip-flop terminal loops link the two DNA strands of the vaccinia virus genome into one uninterrupted polynucleotide chain. Cell 28:315–324
Batesman AJ (1975) Simplification of palindromic telomere theory. Nature 253:379–380
Blackburn EH (1984) Telomeres: do the ends justify the means? Cell 37:7–8
Camougrand N, Mila B, Velours G, Lazowska J, Guerin B (1988) Discrimination between different groups of Candida parapsilosis by mitochondrial DNA restriction analysis. Curr Genet 13:445–449
Casey J, Hsu H, Rabinowitz M, Getz GS, Fukuhara H (1974) Transfer RNA genes in the mitochondrial DNA of cytoplasmic petite mutants of Saccharomyces cerevisiae. J Mol Biol 88:717–733
Cavalier-Smith T (1974) Palindromic sequences and replication of eukaryotic chromosomes. Nature 250:467–470
Cummings DJ, Pritchard AE (1982) Replication mechanism of mitochondrial DNA from Paramecium aurelia: Sequence of the cross-linked origin. In: Slonimski PP, Attardi G, Borst P (eds) Mitochondrial genes. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, pp 441–447
Dinouël N, Drissi R, Miyakawa I, Sor F, Rousset S, Fukuhara H (1993) Linear mitochondrial DNAs of yeasts. Closed hairpin structure of the termini and possible linear-circular conversion mechanisms. Mol Cell Biol 13:2315–1323
Dunn B, Szauter P, Purdue ML, Szostak JW (1984) Transfer of yeast telomeres to linear plasmids by recombination. Cell 39:191–201
Fukuhara H, Sor F, Drissi R, Dinouël N, Miyakawa I, Rousset S, Viola AM (1993) Linear mitochondrial DNAs of yeasts. Frequency of occurrence and general features. Mol Cell Biol 13:2309–2314
Gargouri A (1989) A rapid and simple method for extracting yeast mitochondrial DNA. Curr Genet 15:235–237
Goddard JM, Cummings DJ (1975) Structure and replication of mitochondrial DNA from Paramecium aurelia. J Mol Biol 97:593–609
Guélin E, Velours J, Guerin M (1990) Cloning and sequencing of a fragment of the linear mitochondrial DNA of the yeast Candida parapsilosis. Nucleic Acids Res 18:4267
Guélin E, Guérin M, Velours J (1991) Isolation of the ATP synthase subunit 6 and sequence of the mitochondrial ATP6 gene of the yeast Candida parapsilosis. Eur J Biochem 197:105–111
Guilham N (1978) Organelle heredity. Raven Press, New York
Gunge N, Tamaru A, Ozawa F, Sakaguchi K (1981) Isolation and characterization of linear deoxyribonucleic acid plasmids from Kluyveromyces lactis and the plasmid-associated killer character. J Bacteriol 145:382–390
Horowitz H, Haber JE (1985) Identification of autonomously replicating circular subtelomeric Y' elements in Saccharomyces cerevisiae. Mol Cell Biol 5:2369–2380
Kovac L, Lazowska J, Slonimski PP (1984) A yeast with linear molecules of mitochondrial DNA. Mol Gen Genet 197:420–424
Maleszka R, Skelly PJ, Clark-Walker GD (1991) Rolling circle replication of DNA in yeast mitochondria. EMBO J 10:3923–3929
Morin GB, Cech TR (1986) The telomeres of the linear mitochondrial DNA of Tetrahymena thermophila consist of 53 by tandem repeats. Cell 46:873–883
Morin GB, Cech TR (1988) Mitochondrial telomeres: surprising diversity of repeated telomeric sequences among six species of Tetrahymena. Cell 52:367–374
Nosek J, Fukuhara H (1994) Mitochondrial transfer RNA genes of the yeast Candida parapsilosis. Gene 142:307–308
Nosek J, Fukuhara H (1994) The NADH dehydrogenase subunit genes in the mitochondrial DNA of yeasts. J Bacteriol 176:5622–5634
Ragnini A, Fukuhara H (1989) Genetic instability of an oligomycin resistance mutation in yeast is associated with an amplification of a mitochondrial DNA segment. Nucleic Acids Res 17:6927–6937
Ryan R, Grant D, Chiang KS, Swift H (1978) Isolation and characterization of mitochondrial DNA from Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 75:3268–3272
Takano H, Kawano S, Kuroiwa T (1991) Telomeric structures in a linear mitochondrial plasmid from Physarum polycephalum. Curr Genet 20:315–317
Vahrenholz C, Riemen G, Pratje E, Dujon B, Michaelis G (1993) Mitochondrial DNA of Chlamydomonas reinhardtii: The structure of replication. Curr Genet 24:241–247
Volkert F, Wilson C, Broach JR (1989) Deoxyribonucleic acid plasmids in yeasts. Microbiol Rev 53:299–317
Williamson DH, Johnston LH, Richmond KMV, Game JC (1977) Mitochondrial DNA and the heritable unit of the yeast mitochondrial genome. A review. In: Bandlow W, Schweyen RJ, Wolf K, Kaudewitz F (eds) Mitochondria 1977. Genetics and biogenesis of mitochondria. Walter de Gruyer, Berlin, pp 1–24
Author information
Authors and Affiliations
Additional information
Communicated by C. Hollenberg
Rights and permissions
About this article
Cite this article
Nosek, J., Dinouël, N., Kovac, L. et al. Linear mitochondrial DNAs from yeasts: telomeres with large tandem repetitions. Molec. Gen. Genet. 247, 61–72 (1995). https://doi.org/10.1007/BF00425822
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/BF00425822