Abstract
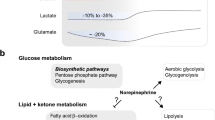
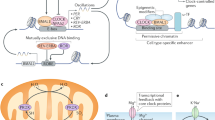
This review proposes a mechanistic link between cellular metabolic status, transcriptional regulatory changes and sleep. Sleep loss is associated with changes in cellular metabolic status in the brain. Metabolic sensors responsive to cellular metabolic status regulate the circadian clock transcriptional network. Modifications of the transcriptional activity of circadian clock genes affect sleep/wake state changes. Changes in sleep state reverse sleep loss-induced changes in cellular metabolic status. It is thus proposed that the regulation of circadian clock genes by cellular metabolic sensors is a critical intermediate step in the link between cellular metabolic status and sleep. Studies of this regulatory relationship may offer insights into the function of sleep at the cellular level.

Similar content being viewed by others
Abbreviations
- NREMS:
-
Non-rapid eye movement sleep
- SWA:
-
Slow wave activity
- EEG:
-
Electroencephalographic/electroencephalogram
- ATP:
-
Adenosine triphosphate
- ADP:
-
Adenosine diphosphate
- AMP:
-
Adenosine monophosphate
- AMPK:
-
Adenosine monophosphate-activated protein kinase
- cry:
-
Cryptochrome
- per:
-
Period
- NAD:
-
Nicotine adenine dinucleotide
- HDAC:
-
Histone deacetylase
- PARP:
-
Poly ADP-ribose polymerase
- GSK3b:
-
Glycogen synthase kinase 3b
- PPARs:
-
Peroxisome proliferator-activated receptors
References
Achermann P, Borbely AA (1997) Low-frequency (<1 Hz) oscillations in the human sleep electroencephalogram. Neuroscience 81:213–222
Ahnaou A, Drinkenburg WH (2010) Disruption of glycogen synthase kinase-3-beta activity leads to abnormalities in physiological measures in mice. Behav Brain Res 221:246–252
Alcain FJ, Villalba JM (2009) Sirtuin inhibitors. Expert Opin Ther Pat 19:283–294
Alle H, Roth A, Geiger JR (2009) Energy-efficient action potentials in hippocampal mossy fibers. Science 325:1405–1408
Asher G, Schibler U (2011) Crosstalk between components of circadian and metabolic cycles in mammals. Cell Metab 13:125–137
Asher G, Gatfield D, Stratmann M, Reinke H, Dibner C, Kreppel F, Mostoslavsky R, Alt FW, Schibler U (2008) SIRT1 regulates circadian clock gene expression through PER2 deacetylation. Cell 134:317–328
Asher G, Reinke H, Altmeyer M, Gutierrez-Arcelus M, Hottiger MO, Schibler U (2010) Poly(ADP-ribose) polymerase 1 participates in the phase entrainment of circadian clocks to feeding. Cell 142:943–953
Attwell D, Laughlin SB (2001) An energy budget for signaling in the grey matter of the brain. J Cereb Blood Flow Metab 21:1133–1145
Bechtold DA (2008) Energy-responsive timekeeping. J Genet 87:447–458
Benington JH, Heller HC (1995) Restoration of brain energy metabolism as the function of sleep. Prog Neurobiol 45:347–360
Boger DL, Henriksen SJ, Cravatt BF (1998) Oleamide: an endogenous sleep-inducing lipid and prototypical member of a new class of biological signaling molecules. Curr Pharm Des 4:303–314
Borbely AA, Achermann P (2004) Sleep homeostasis and models of sleep regulation. In: Kryger MH, Roth T, Dement WC (eds) Principles and practice of sleep medicine. Saunders, Philadelphia, pp 377–390
Bouaboula M, Hilairet S, Marchand J, Fajas L, Le Fur G, Casellas P (2005) Anandamide induced PPARgamma transcriptional activation and 3T3-L1 preadipocyte differentiation. Eur J Pharmacol 517:174–181
Buchsbaum MS, Gillin JC, Wu J, Hazlett E, Sicotte N, Dupont RM, Bunney WE Jr (1989) Regional cerebral glucose metabolic rate in human sleep assessed by positron emission tomography. Life Sci 45:1349–1356
Buzsaki G, Kaila K, Raichle M (2007) Inhibition and brain work. Neuron 56:771–783
Canaple L, Rambaud J, Dkhissi-Benyahya O, Rayet B, Tan NS, Michalik L, Delaunay F, Wahli W, Laudet V (2006) Reciprocal regulation of brain and muscle Arnt-like protein 1 and peroxisome proliferator-activated receptor alpha defines a novel positive feedback loop in the rodent liver circadian clock. Mol Endocrinol 20:1715–1727
Chikahisa S, Tominaga K, Kawai T, Kitaoka K, Oishi K, Ishida N, Rokutan K, Sei H (2008) Bezafibrate, a PPARs agonist, decreases body temperature and enhances EEG delta oscillation during sleep in mice. Endocrinology 149:5262–5271
Chikahisa S, Fujiki N, Kitaoka K, Shimizu N, Sei H (2009) Central AMPK contributes to sleep homeostasis in mice. Neuropharmacology 57:369–374
Cirelli C (2006) Cellular consequences of sleep deprivation in the brain. Sleep Med Rev 10:307–321
Cirelli C, Tononi G (1998) Differences in gene expression between sleep and waking as revealed by mRNA differential display. Brain Res Mol Brain Res 56:293–305
Cravatt BF, Prospero-Garcia O, Siuzdak G, Gilula NB, Henriksen SJ, Boger DL, Lerner RA (1995) Chemical characterization of a family of brain lipids that induce sleep. Science 268:1506–1509
Dauvilliers Y, Tafti M (2008) The genetic basis of sleep disorders. Curr Pharm Des 14:3386–3395
DeBruyne JP, Noton E, Lambert CM, Maywood ES, Weaver DR, Reppert SM (2006) A clock shock: mouse CLOCK is not required for circadian oscillator function. Neuron 50:465
Destexhe A, Contreras D, Steriade M (1999) Spatiotemporal analysis of local field potentials and unit discharges in cat cerebral cortex during natural wake and sleep states. J Neurosci 19:4595–4608
Dworak M, McCarley RW, Kim T, Kalinchuk AV, Basheer R (2010) Sleep and brain energy levels: ATP changes during sleep. J Neurosci 30:9007–9016
Etchegaray JP, Machida KK, Noton E, Constance CM, Dallmann R, Di Napoli MN, DeBruyne JP, Lambert CM, Yu EA, Reppert SM, Weaver DR (2009) Casein kinase 1 delta regulates the pace of the mammalian circadian clock. Mol Cell Biol 29:3853–3866
Franken P, Dijk DJ (2009) Circadian clock genes and sleep homeostasis. Eur J Neurosci 29:1820–1829
Franken P, Thomason R, Heller HC, O'Hara BF (2007) A non-circadian role for clock-genes in sleep homeostasis: a strain comparison. BMC Neurosci 8:87
Fu J, Gaetani S, Oveisi F, Lo Verme J, Serrano A, Rodriguez De Fonseca F, Rosengarth A, Luecke H, Di Giacomo B, Tarzia G, Piomelli D (2003) Oleylethanolamide regulates feeding and body weight through activation of the nuclear receptor PPAR-alpha. Nature 425:90–93
Gamble KL, Allen GC, Zhou T, McMahon DG (2007) Gastrin-releasing peptide mediates light-like resetting of the suprachiasmatic nucleus circadian pacemaker through cAMP response element-binding protein and Per1 activation. J Neurosci 27:12078–12087
Haque R, Ali FG, Biscoglia R, Abey J, Weller J, Klein D, Iuvone PM (2011) CLOCK and NPAS2 have overlapping roles in the circadian oscillation of arylalkylamine N-acetyltransferase mRNA in chicken cone photoreceptors. J Neurochem 113:1296–1306
Hardie DG (2007) AMP-activated/SNF1 protein kinases: conserved guardians of cellular energy. Nat Rev Mol Cell Biol 8:774–785
Hassa PO, Haenni SS, Elser M, Hottiger MO (2006) Nuclear ADP-ribosylation reactions in mammalian cells: where are we today and where are we going? Microbiol Mol Biol Rev 70:789–829
He Y, Jones CR, Fujiki N, Xu Y, Guo B, Holder JL Jr, Rossner MJ, Nishino S, Fu YH (2009) The transcriptional repressor DEC2 regulates sleep length in mammals. Science 325:866–870
Heiss WD, Pawlik G, Herholz K, Wagner R, Wienhard K (1985) Regional cerebral glucose metabolism in man during wakefulness, sleep, and dreaming. Brain Res 327:362–366
Hirayama J, Sahar S, Grimaldi B, Tamaru T, Takamatsu K, Nakahata Y, Sassone-Corsi P (2007) CLOCK-mediated acetylation of BMAL1 controls circadian function. Nature 450:1086–1090
Hong HK, Chong JL, Song W, Song EJ, Jyawook AA, Schook AC, Ko CH, Takahashi JS (2007) Inducible and reversible Clock gene expression in brain using the tTA system for the study of circadian behavior. PLoS Genet 3:e33
Hottiger MO, Boothby M, Koch-Nolte F, Luscher B, Martin NM, Plummer R, Wang ZQ, Ziegler M (2011) Progress in the function and regulation of ADP-ribosylation. Sci Signal 4:mr5
Huang W, Ramsey KM, Marcheva B, Bass J (2011) Circadian rhythms, sleep, and metabolism. J Clin Invest 121:2133–2141
Iitaka C, Miyazaki K, Akaike T, Ishida N (2005) A role for glycogen synthase kinase-3beta in the mammalian circadian clock. J Biol Chem 280:29397–29402
Johnson LA, Euston DR, Tatsuno M, McNaughton BL (2008) Stored-trace reactivation in rat prefrontal cortex is correlated with down-to-up state fluctuation density. J Neurosci 30:2650–2661
Jones BE (2004) Activity, modulation and role of basal forebrain cholinergic neurons innervating the cerebral cortex. Prog Brain Res 145:157–169
Karnovsky ML, Reich P, Anchors JM, Burrows BL (1983) Changes in brain glycogen during slow-wave sleep in the rat. J Neurochem 41:1498–1501
Kennedy C, Gillin JC, Mendelson W, Suda S, Miyaoka M, Ito M, Nakamura RK, Storch FI, Pettigrew K, Mishkin M, Sokoloff L (1982) Local cerebral glucose utilization in non-rapid eye movement sleep. Nature 297:325–327
Koethe D, Schreiber D, Giuffrida A, Mauss C, Faulhaber J, Heydenreich B, Hellmich M, Graf R, Klosterkotter J, Piomelli D, Leweke FM (2009) Sleep deprivation increases oleoylethanolamide in human cerebrospinal fluid. J Neural Transm 116:301–305
Krishnakumar R, Kraus WL The PARP side of the nucleus: molecular actions, physiological outcomes, and clinical targets. Mol Cell 39:8–24
Krueger JM, Rector DM, Roy S, Van Dongen HP, Belenky G, Panksepp J (2008) Sleep as a fundamental property of neuronal assemblies. Nat Rev Neurosci 9:910–919
Lamia KA, Sachdeva UM, DiTacchio L, Williams EC, Alvarez JG, Egan DF, Vasquez DS, Juguilon H, Panda S, Shaw RJ, Thompson CB, Evans RM (2009) AMPK regulates the circadian clock by cryptochrome phosphorylation and degradation. Science 326:437–440
Landgraf D, Shostak A, Oster H (2011) Clock genes and sleep. Pfluger's Archives Euro J Physiol, in press.
Mackiewicz M, Shockley KR, Romer MA, Galante RJ, Zimmerman JE, Naidoo N, Baldwin DA, Jensen ST, Churchill GA, Pack AI (2007) Macromolecule biosynthesis: a key function of sleep. Physiol Genomics 31:441–457
Maquet P (1995) Sleep function(s) and cerebral metabolism. Behav Brain Res 69:75–83
Maquet P, Dive D, Salmon E, Sadzot B, Franco G, Poirrier R, von Frenckell R, Franck G (1990) Cerebral glucose utilization during sleep–wake cycle in man determined by positron emission tomography and [18F]2-fluoro-2-deoxy-d-glucose method. Brain Res 513:136–143
Martinek S, Inonog S, Manoukian AS, Young MW (2001) A role for the segment polarity gene shaggy/GSK-3 in the Drosophila circadian clock. Cell 105:769–779
McMurry J, Castellion ME (1976) The generation of biochemical energy, organic and biological chemistry. Prentice-Hall, Upper Saddle River, NJ, pp 590–619
Morgenthaler FD, Lanz BR, Petit JM, Frenkel H, Magistretti PJ, Gruetter R (2009) Alteration of brain glycogen turnover in the conscious rat after 5 h of prolonged wakefulness. Neurochem Int 55:45–51
Mukovski M, Chauvette S, Timofeev I, Volgushev M (2007) Detection of active and silent states in neocortical neurons from the field potential signal during slow-wave sleep. Cereb Cortex 17:400–414
Naylor E, Aillon DV, Gabbert S, Harmon H, Johnson DA, Wilson GS, Petillo PA (2011) Real-time measurement of EEG/EMG and l-glutamate in mice: a biosensor study of neuronal activity during sleep. J Electroanal Chem 656:106–113
Nelson DL, Cox MM (2008) Bioenergetics and biochemical reaction types. Lehninger principles of biochemistry. Freeman, New York, pp 485–519
Nelson DL, Cox MM (2008) The citric acid cycle. Lehninger principles of biochemistry. Freeman, New York, pp 615–646
Nelson DL, Cox MM (2008) Principles of metabolic regulation. Lehninger principles of biochemistry. Freeman, New York, pp 569–614
Nir Y, Staba RJ, Andrillon T, Vyazovskiy VV, Cirelli C, Fried I, Tononi G (2011) Regional slow waves and spindles in human sleep. Neuron 70:153–169
Nonaka K, Nakazawa Y, Kotorii T (1983) Effects of antibiotics, minocycline and ampicillin, on human sleep. Brain Res 288:253–259
Oishi K, Miyazaki K, Kadota K, Kikuno R, Nagase T, Atsumi G, Ohkura N, Azama T, Mesaki M, Yukimasa S, Kobayashi H, Iitaka C, Umehara T, Horikoshi M, Kudo T, Shimizu Y, Yano M, Monden M, Machida K, Matsuda J, Horie S, Todo T, Ishida N (2003) Genome-wide expression analysis of mouse liver reveals CLOCK-regulated circadian output genes. J Biol Chem 278:41519–41527
Panda S, Antoch MP, Miller BH, Su AI, Schook AB, Straume M, Schultz PG, Kay SA, Takahashi JS, Hogenesch JB (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109:307–320
Panossian L, Fenik P, Zhu Y, Zhan G, McBurney MW, Veasey S (2010) SIRT1 regulation of wakefulness and senescence-like phenotype in wake neurons. J Neurosci 31:4025–4036
Petit JM, Tobler I, Kopp C, Morgenthaler F, Borbely AA, Magistretti PJ (2010) Metabolic response of the cerebral cortex following gentle sleep deprivation and modafinil administration. Sleep 33:901–908
Raizen DM, Wu MN (2010) Genome-wide association studies of sleep disorders. Chest 139:446–452
Ramanathan L, Hu S, Frautschy SA, Siegel JM (2010) Short-term total sleep deprivation in the rat increases antioxidant responses in multiple brain regions without impairing spontaneous alternation behavior. Behav Brain Res 207:305–309
Rector DM (2010) Local functional state differences between rat cortical columns. Curr Topics Med Chem, in press
Rutter J, Reick M, Wu LC, McKnight SL (2001) Regulation of clock and NPAS2 DNA binding by the redox state of NAD cofactors. Science 293:510–514
Scharf MT, Naidoo N, Zimmerman JE, Pack AI (2008) The energy hypothesis of sleep revisited. Prog Neurobiol 86:264–280
Siegel JM (2011) REM sleep: a biological and psychological paradox. Sleep Med 15:139–142
Silva RH, Abilio VC, Takatsu AL, Kameda SR, Grassl C, Chehin AB, Medrano WA, Calzavara MB, Registro S, Andersen ML, Machado RB, Carvalho RC, Ribeiro Rde A, Tufik S, Frussa-Filho R (2004) Role of hippocampal oxidative stress in memory deficits induced by sleep deprivation in mice. Neuropharmacology 46:895–903
Smith SA (2002) Peroxisome proliferator-activated receptors and the regulation of mammalian lipid metabolism. Biochem Soc Trans 30:1086–1090
Szymusiak R (2010) Hypothalamic versus neocortical control of sleep. Curr Opin Pulm Med 16:530–535
Thakkar M, Mallick BN (1993) Rapid eye movement sleep-deprivation-induced changes in glucose metabolic enzymes in rat brain. Sleep 16:691–694
Tu BP, McKnight SL (2006) Metabolic cycles as an underlying basis of biological oscillations. Nat Rev Mol Cell Biol 7:696–701
Van den Noort S, Brine K (1970) Effect of sleep on brain labile phosphates and metabolic rate. Am J Physiol 218:1434–1439
Vaziri H, Dessain SK, Ng Eaton E, Imai SI, Frye RA, Pandita TK, Guarente L, Weinberg RA (2001) hSIR2(SIRT1) functions as an NAD-dependent p53 deacetylase. Cell 107:149–159
Vyazovskiy VV, Cirelli C, Pfister-Genskow M, Faraguna U, Tononi G (2008) Molecular and electrophysiological evidence for net synaptic potentiation in wake and depression in sleep. Nat Neurosci 11:200–208
Vyazovskiy VV, Olcese U, Lazimy YM, Faraguna U, Esser SK, Williams JC, Cirelli C, Tononi G (2009) Cortical firing and sleep homeostasis. Neuron 63:865–878
Walker JW, Jijon HB, Madsen KL (2006) AMP-activated protein kinase is a positive regulator of poly(ADP-ribose) polymerase. Biochem Biophys Res Commun 342:336–341
Wang N, Yang G, Jia Z, Zhang H, Aoyagi T, Soodvilai S, Symons JD, Schnermann JB, Gonzalez FJ, Litwin SE, Yang T (2008) Vascular PPARgamma controls circadian variation in blood pressure and heart rate through Bmal1. Cell Metab 8:482–491
Wigren HK, Rytkonen KM, Porkka-Heiskanen T (2009) Basal forebrain lactate release and promotion of cortical arousal during prolonged waking is attenuated in aging. J Neurosci 29:11698–11707
Wisor JP, Clegern WC (2011) Quantification of short-term slow wave sleep homeostasis and its disruption by minocycline in the laboratory mouse. Neurosci Lett 490:165–169
Wisor JP, Kilduff TS (2005) Molecular genetic advances in sleep research and their relevance to sleep medicine. Sleep 28:357–367
Wisor JP, O'Hara BF, Terao A, Selby CP, Kilduff TS, Sancar A, Edgar DM, Franken P (2002) A role for cryptochromes in sleep regulation. BMC Neurosci 3:20
Wisor JP, Pasumarthi RK, Gerashchenko D, Thompson CL, Pathak S, Sancar A, Franken P, Lein ES, Kilduff TS (2008) Sleep deprivation effects on circadian clock gene expression in the cerebral cortex parallel electroencephalographic differences among mouse strains. J Neurosci 28:7193–7201
Wisor JP, Schmidt MA, Clegern WC (2011) Evidence for neuroinflammatory and microglial changes in the cerebral response to sleep loss. Sleep 34:261–272
Acknowledgements
The author is supported by DARPA Young Faculty Award N66001-09-1-2117 and NINDS 1R15NS070734.
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published as part of the Special Issue on Sleep.
Rights and permissions
About this article
Cite this article
Wisor, J.P. A metabolic–transcriptional network links sleep and cellular energetics in the brain. Pflugers Arch - Eur J Physiol 463, 15–22 (2012). https://doi.org/10.1007/s00424-011-1030-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00424-011-1030-6