Abstract
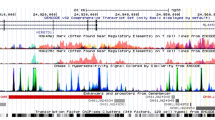
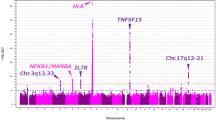
In the United States, biliary atresia (BA) is the most frequent indication for liver transplantation in pediatric patients. BA is a complex disease, with suspected environmental and genetic risk factors. A genome-wide association study in Chinese patients identified association to the 10q24.2 (hg18) genomic region. This signal was upstream of two genes, XPNPEP1 and ADD3, both expressed in intrahepatic bile ducts. We tested association to this region in 171 BA patients and 1,630 controls of European descent and found the strongest signal to be at rs7099604 (p = 2.5 × 10−3) in intron 1 of the ADD3 gene. Moreover, expression data suggest that ADD3, but not XPNPEP1, is differentially expressed in BA patients. The role of ADD3 in biliary development is unclear, but our findings suggest that this gene may be functionally relevant for the development of BA.



Similar content being viewed by others
References
1000 Genomes Project Consortium (2012) An integrated map of genetic variation from 1,092 human genomes. Nature 491:56–65
Barrett JC, Fry B, Maller J, Daly MJ (2005) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Cheng G, Tang CS, Wong EH, Cheng WW, So MT, Miao X, Zhang R, Cui L, Liu X, Ngan ES, Lui VC, Chung PH, Chan IH, Liu J, Zhong W, Xia H, Yu J, Qiu X, Wu XZ, Wang B, Dong X, Tou J, Huang L, Yi B, Ren H, Chan EK, Ye K, O’Reilly PF, Wong KK, Sham PC, Cherny SS, Tam PK, Garcia-Barcelo MM (2013) Common genetic variants regulating ADD3 gene expression alter biliary atresia risk. J Hepatol. doi:10.1016/j.jhep.2013.07.021
Citterio L, Tizzoni L, Catalano M, Zerbini G, Bianchi G, Barlassina C (2003) Expression analysis of the human adducin gene family and evidence of ADD2 beta 4 multiple splicing variants. Biochem Biophys Res Commun 309:359–367
Cui S, Leyva-Vega M, Tsai EA, Eauclaire SF, Glessner JT, Hakonarson H, Devoto M, Haber BA, Spinner NB, Matthews RP (2013) Evidence from human and zebrafish that GPC1 is a biliary atresia susceptibility Gene. Gastroenterology 144(5):1107–1115
Cusi D, Barlassina C, Azzani T, Casari G, Citterio L, Devoto M, Glorioso N, Lanzani C, Manunta P, Righetti M, Rivera R, Stella P, Troffa C, Zagato L, Bianchi G (1997) Polymorphisms of alpha-adducin and salt sensitivity in patients with essential hypertension. Lancet 349:1353–1357
Erlichman J, Hohlweg K, Haber BA (2009) Biliary atresia: how medical complications and therapies impact outcome. Expert Rev Gastroenterol Hepatol 3:425–434
Ersahin C, Szpaderska AM, Orawski AT, Simmons WH (2005) Aminopeptidase P isozyme expression in human tissues and peripheral blood mononuclear cell fractions. Arch Biochem Biophys 435:303–310
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B, Higgins J, DeFelice M, Lochner A, Faggart M, Liu-Cordero SN, Rotimi C, Adeyemo A, Cooper R, Ward R, Lander ES, Daly MJ, Altshuler D (2002) The structure of haplotype blocks in the human genome. Science 296:2225–2229
Garcia-Barcelo MM, Yeung MY, Miao XP, Tang CS, Chen G, So MT, Ngan ES, Lui VC, Chen Y, Liu XL, Hui KJ, Li L, Guo WH, Sun XB, Tou JF, Chan KW, Wu XZ, Song YQ, Chan D, Cheung K, Chung PH, Wong KK, Sham PC, Cherny SS, Tam PK (2010) Genome-wide association study identifies a susceptibility locus for biliary atresia on 10q24.2. Hum Mol Genet 19:2917–2925
Howie BN, Donnelly P, Marchini J (2009) A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet 5:e1000529
International HapMap 3 Consortium (2010) Integrating common and rare genetic variation in diverse human populations. Nature 467:52–58
Kaewkiattiyot S, Honsawek S, Vejchapipat P, Chongsrisawat V, Poovorawan Y (2011) Association of X-prolyl aminopeptidase 1 rs17095355 polymorphism with biliary atresia in Thai children. Hepatol Res 41:1249–1252
Kowlessar OD, Haeffner LJ, Riley EM, Sleisenger MH (1961) Comparative study of serum leucine aminopeptidase, 5-nucleotidase and non-specific alkaline phosphatase in diseases affecting the pancreas, hepatobiliary tree and bone. Am J Med 31:231–237
Leyva-Vega M, Gerfen J, Thiel BD, Jurkiewicz D, Rand EB, Pawlowska J, Kaminska D, Russo P, Gai X, Krantz ID, Kamath BM, Hakonarson H, Haber BA, Spinner NB (2010) Genomic alterations in biliary atresia suggest region of potential disease susceptibility in 2q37.3. Am J Med Genet A 152A:886–895
Li J, Bessho K, Shivakumar P, Mourya R, Mohanty SK, Dos Santos JL, Miura IK, Porta G, Bezerra JA (2011a) Th2 signals induce epithelial injury in mice and are compatible with the biliary atresia phenotype. J Clin Invest 121:4244–4256
Li X, Bartlam M, Rao Z (2011b) Human cytosolic X-Prolyl aminopeptidase. Encyclopedia of inorganic and bioinorganic chemistry. Wiley, New York
Mack CL, Sokol RJ (2005) Unraveling the pathogenesis and etiology of biliary atresia. Pediatr Res 57:87R–94R
Marchini J, Howie B, Myers S, McVean G, Donnelly P (2007) A new multipoint method for genome-wide association studies by imputation of genotypes. Nat Genet 39:906–913
Muraji T, Suskind DL, Irie N (2009) Biliary atresia: a new immunological insight into etiopathogenesis. Expert Rev Gastroenterol Hepatol 3:599–606
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
R Core Team (2012) R: a language and environment for statistical computing. http://www.R-project.org
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, Wong MC, Maddren M, Fang R, Heitner SG, Lee BT, Barber GP, Harte RA, Diekhans M, Long JC, Wilder SP, Zweig AS, Karolchik D, Kuhn RM, Haussler D, Kent WJ (2013) ENCODE data in the UCSC genome browser: year 5 update. Nucleic Acids Res 41:D56–D63
Shaikh TH, Gai X, Perin JC, Glessner JT, Xie H, Murphy K, O’Hara R, Casalunovo T, Conlin LK, D’Arcy M, Frackelton EC, Geiger EA, Haldeman-Englert C, Imielinski M, Kim CE, Medne L, Annaiah K, Bradfield JP, Dabaghyan E, Eckert A, Onyiah CC, Ostapenko S, Otieno FG, Santa E, Shaner JL, Skraban R, Smith RM, Elia J, Goldmuntz E, Spinner NB, Zackai EH, Chiavacci RM, Grundmeier R, Rappaport EF, Grant SF, White PS, Hakonarson H (2009) High-resolution mapping and analysis of copy number variations in the human genome: a data resource for clinical and research applications. Genome Res 19:1682–1690
Shivakumar P, Campbell KM, Sabla GE, Miethke A, Tiao G, McNeal MM, Ward RL, Bezerra JA (2004) Obstruction of extrahepatic bile ducts by lymphocytes is regulated by IFN-gamma in experimental biliary atresia. J Clin Invest 114:322–329
Shneider BL, Brown MB, Haber B, Whitington PF, Schwarz K, Squires R, Bezerra J, Shepherd R, Rosenthal P, Hoofnagle JH, Sokol RJ (2006) A multicenter study of the outcome of biliary atresia in the United States, 1997 to 2000. J Pediatr 148:467–474
Shneider BL, Abel B, Haber B, Karpen SJ, Magee JC, Romero R, Schwarz K, Bass LM, Kerkar N, Miethke AG, Rosenthal P, Turmelle Y, Robuck PR, Sokol RJ (2012) Portal hypertension in children and young adults with biliary atresia. J Pediatr Gastroenterol Nutr 55:567–573
Acknowledgments
We would like to thank the patients and their families who donated their samples for this study. We also would like to acknowledge R.P. Matthews for his preliminary data on the biliary phenotype of xpnpep1 and add3a knockdown constructs in zebrafish, and M. Leyva-Vega, A. Hutchinson, and L.D. Leonard for their help in obtaining the BA samples from ChiLDREN. This work was supported by The Fred and Suzanne Biesecker Liver Center at CHOP as well as R01-DK090045 to N.B.S, U01-DK062481 to K.M.L., and R01-DK083781 to J.A.B. E.A.T. was also supported by training grant T32-HG000046.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Tsai, E.A., Grochowski, C.M., Loomes, K.M. et al. Replication of a GWAS signal in a Caucasian population implicates ADD3 in susceptibility to biliary atresia. Hum Genet 133, 235–243 (2014). https://doi.org/10.1007/s00439-013-1368-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-013-1368-2