Abstract
Hutchinson-Gilford progeria syndrome (HGPS; MIM 176670) is a rare disease characterized by accelerated aging. In this study, light and immunofluorescence microscopy were used to assess morphological changes, measures of cell growth kinetics and gene expression profiles in HGPS cells and normal fibroblasts in culture. A filtering strategy was developed based on differentially expressed transcripts seen consistently across three culture stages based on cell passage number. This filtering strategy produced a list of 66 unique differentially expressed genes, of which ∼40% were upregulated in HGPS cells compared to normal fibroblasts. The increased mRNA expression in HGPS cells that was seen for one gene defined using this strategy—namely ANK3— was validated using quantitative reverse-transcriptase amplification, Western analysis and immunofluorescence microscopy, all of which showed significantly increased ankyrin G expression. These findings demonstrate differences in morphology, growth kinetics and mRNA expression profiles in HGPS cells compared to normal fibroblasts in culture, including increased expression of ANK3/ankyrin G. Furthermore, other genes that co-clustered with ANK3 might provide mechanistic clues regarding senescence in cultured HGPS cells.
Similar content being viewed by others
Introduction
Hutchinson-Gilford progeria syndrome (HGPS; MIM 176670) is an extremely rare disease that is characterized by accelerated aging and early death (Pollex and Hegele 2004). Organ systems degenerate to such an extent that the affected subject resembles an old person. Clinical features include short stature, micrognathia, alopecia, prominent scalp veins, prominent joints, hyperlipidemia and atherosclerosis, often with premature death from coronary artery disease. The genetic basis for >90% of cases of HGPS is a de novo recurrent c.2036C>T splicing mutation in LMNA, encoding nuclear lamin A/C, although there is some heterogeneity of mutations (Eriksson et al. 2003; De Sandre-Giovannoli et al. 2003; Cao and Hegele 2003; Csoka et al. 2004a).
In cultured HGPS fibroblasts, others have shown differential gene expression profiles, including changes in expression of genes within the ontologies of transcription factors and extracellular matrix proteins compared to normal fibroblasts (Csoka et al. 2004b). Since the LMNA mutation is the primary insult within the germline of HGPS fibroblasts, changes in gene expression profiles are secondary and may provide clues about dysregulated pathways that lead to abnormal phenotypes in cultured cells and perhaps within intact tissues and organs. Our current objectives were to: (1) develop a biological filtering approach to characterize cellular and molecular differences between cultured fibroblasts from a HGPS proband with the dominant LMNA c.2036C>T splicing mutation and fibroblasts from a normal subject; and (2) validate this approach at the protein level using Western analysis and immunocytochemistry to study the product of a candidate gene that showed marked differential mRNA expression. The filtering criteria included opposite patterns of expression between HGPS and normal fibroblasts at three different stages of cell culture maturity defined by passage number.
Materials and methods
Cell culture
Human normal (AG04456) and HGPS (AG03513, LMNA mutation c.2036C>T) fibroblast cell lines were obtained from the Coriell Cell Repository (Camden, NJ). Cells were grown in Dulbecco’s Modification of Eagle’s Medium (DMEM), supplemented with 15% non-inactivated fetal bovine serum (FBS), at 37°C in 5% CO2. Cells were passaged every 3 or 4 days and seeded at a density of 2×105 per 90 mm diameter dish. Cell growth curves were obtained by measuring accumulated population doublings (APD), using the formula (logH−logS)/log 2.0, where logH and logS are the logarithms of the numbers of cells harvested and seeded, respectively (Bridger and Kill 2004).
Genomic DNA isolation and direct sequencing
Genomic DNA was isolated from cultured cells using the Puregene kit (Gentra Systems, Minneapolis, MN). All coding exons, intron–exon boundaries and >300 bp of 5′ and 3′ untranslated regions of the LMNA gene were amplified as described (Cao and Hegele 2003) and were then sequenced on a ABI 3730 DNA Analyzer (Applied Biosystems, Mississauga, ON). Mutations were identified using Sequence Navigator software (Applied Biosystems).
Total RNA isolation
RNA was isolated at the elapsed cell culture ages of APD 2.5 (young), APD 10 (middle-aged) for both the normal and HGPS fibroblast cell lines; and at APD 26 (old) for the normal cell line and APD 17 (old) for the HGPS fibroblasts. Cells were washed twice with PBS and total RNA was then isolated using an RNeasy kit (Qiagen, Mississauga, ON). Isolated total RNA was quantified by ultraviolet spectroscopy, and RNA integrity was evaluated using the BioAnalyzer 2100 Model G2938A (Agilent Technologies, Mississauga, ON).
Immunofluorescence studies
In order to determine proliferation differences between normal and HGPS fibroblasts, cells were grown on coverslips to 80% confluence. Cells on coverslips were washed twice with PBS, fixed with 4% paraformaldehyde on ice for 20 min, and incubated with 0.5% Triton X-100 at room temperature for 5 min. Fixed cells were next incubated with a 1:150 dilution of monoclonal mouse anti-human Ki-67 antibody (DakoCytomation, Mississauga, ON) at 37°C in a humidified chamber for 1 h, washed three times with PBS, then incubated with a 1:100 dilution of FITC goat anti-mouse IgG (Cedarlane Laboratories, Hornby, ON) at room temperature in a humidified chamber for 30 min. After washing three times with PBS, cells were stained with Hoechst dye (2.5 μg/ml in PBS) (Sigma-Aldrich, Oakville, ON) at room temperature for 5 min. Cells were then washed three times with PBS and mounted on glass slides using PermaFluor Aqueous Mounting Medium (Fisher, Markham, ON). Proliferating cells were identified by positive staining with the Ki-67 antibody.
In order to localize ankyrin G within fibroblasts, cultures grown on cover slips were fixed with 2% paraformaldehyde at room temperature for 10 min, followed by incubation with 0.5% Triton X-100 at room temperature for 2 min. Fixed cells were incubated with a 1:50 dilution of monoclonal mouse anti-human ankyrin G antibody (Santa Cruz Biotechnology, Santa Cruz, CA) at 37°C in a humidified chamber for 1 h, washed three times with PBS, then incubated with a 1:200 dilution of FITC goat anti-mouse IgG at room temperature in a humidified chamber for 30 min. After washing three times with PBS, cells were stained with Hoechst dye (2.5 μg/ml in PBS) at room temperature for 5 min. Cells were then washed three times with PBS and mounted on glass slides using PermaFluor Aqueous Mounting Medium (Fisher, Markham, ON).
Microarray studies
Standard procedures of the London Regional Genomics Centre were used (Carter et al. 2005). Total RNA isolates from three different cell ages—young, middle, and old—of both normal and HGPS cell lines were analyzed using the GeneChip Expression Array HU133A (Affymetrix, Santa Clara, CA). A 25 ng aliquot of total RNA from each sample was processed using the GeneChip Expression 3′-Amplification Reagents Two Cycle cDNA Synthesis Kit (Affymetrix, Santa Clara, CA). Sample quality was assessed using the BioAnalyzer 2100 Model G2938A (Agilent Technologies) and degradation software as described (Carter et al. 2005). Only samples that registered “no alert” were analyzed (Carter et al. 2005). A 600 ng sample of cRNA was used for the second cycle of cDNA and cRNA synthesis. BIOTINYLATED fragmented cRNA (15 μg) was then hybridized onto HU133A chips. GeneChips were washed and scanned on an Affymetrix Fluidics Module Model 400 and GeneChip Scanner Model 3000 controlled by GeneChip Operating System v1.3 (GCOS) (Affymetrix). The GCOS software acquired the image file and processed it to yield a .cel image file. This .cel file was then exported to third party software for expression profile analysis.
Microarray data analysis
The GeneChip experiment was comprised of six microarrays in total, namely preparations from young, middle-aged and old time points from normal and HGPS fibroblast cell lines. In our laboratory the Pearson correlation coefficient was 0.9917 for technical replicates. The .cel files were imported into DNA Chip Analyzer (dCHIP) (Li and Wong 2001) for outlier detection. The array intensities were normalized to the median intensity array and percent present calls, percent array outliers and percent single probe outliers were computed by the model-based analysis algorithm using perfect match only and perfect match mismatch difference probe pairs. There were no arrays flagged as having outlier values >5% from the median intensity, providing a high degree of confidence that all six arrays had no technical or biological artifacts.
Filtering strategy
Agilent GX v7.2 (formerly GeneSping; Agilent Technologies, Redwood City, CA) was used for analysis of differential gene expression. For these analyses, the three microarrays at different ages from each of normal and HGPS RNA were treated as groups to produce lists of genes that were differentially expressed between HGPS and normal fibroblast lines. The .cel processed raw image files produced by GCOS software were imported into GX with robust multi-array average (RMA) pre-processing (Irizarry et al. 2003), a summary measure to normalize arrays that simultaneously adjusted background, normalized and log-transformed perfect match intensity values from the .cel files. A gene list was generated with all Affymetrix technical quality assurance probes, marked using the “Affx” prefix.
A compound filter was used to create the final gene list: (1) remove all “Affx” prefix genes; (2) remove all intensity values ≤150 units in at least three out of six conditions; (3) remove all values ≤ 2-fold difference between normal–young and HGPS–young expression values; and (4) filter on confidence of association with nominal P≤0.1. This filter yielded a set of 66 unique genes that were up-regulated in normal and down-regulated in the HGPS cell line and vice versa (Table 1). A similar compound filter, with the exception of not filtering on confidence, yielded a list of 462 genes that satisfied the filter criteria. This list was the used for quality threshold (QT) clustering (Heyer et al. 1999). ANK3 (ankyrin G; Affymetrix probe set identifier 206385_a_t; NCBI identifier NM_020987) had the greatest magnitude of differential gene expression between normal and HGPS cell lines using both filtering strategies. Using a minimum cluster size of ten and a minimum similarity (Pearson correlation coefficient) >0.99, we identified ten genes that co-clustered with ANK3 based on the expression profile (Table 2).
Quantitative real time polymerase chain reaction analysis
Total RNA isolated from HGPS and normal fibroblasts was reverse transcribed for 2 h at 37°C to generate cDNA using the High Capacity cDNA Archive Kit (Applied Biosystems). Quantitative real time PCR (Q-RT-PCR) was performed using TaqMan Gene Expression Assays (Applied Biosystems) with gene specific primer and probe mix (Hs01088367_m1, Hs00193657_m1, Hs99999905_m1) and TaqMan Universal PCR Master Mix (Applied Biosystems) according to the manufacturer’s instructions for the standard curve method. cDNA (100 ng) was used for each target PCR reaction. To confirm microarray findings, ANK3 and SEPP1 were chosen for verification studies because of their extreme positions on the differentially regulated gene list for HGPS cells and normal fibroblasts (Table 1). Expression of these two genes was determined using the manufacturer’s protocol for Q-RT-PCR and analyzed using ABI 7900HT Fast Real Time PCR System with SDS version 2.2.1 software (Applied Biosystems). GAPDH expression was used as the endogenous control for target normalization.
Western blot analysis
HGPS and control cells taken at the three APD time points were washed twice with PBS, lysed for 5 min in SDS buffer [50 mM Tris-Cl (pH 6.8), 2% SDS, 10% glycerol] at 95°C. Lysates were then heated to 95°C for 5 min and stored at −20°C. Protein concentrations were determined using the DC Protein Assay Kit (Bio-Rad Laboratories, Mississauga, ON). Lysates were supplemented with 50 mM Tris-Cl (pH 6.8), 100 mM DTT, 2% SDS, 0.1% bromophenol blue and 10% glycerol, heated to 95°C for 5 min, loaded on 4–12% gradient SDS-PAGE gel (Invitrogen, Burlington, ON) and transferred to nitrocellulose membranes in a transfer module (Bio-Rad Laboratories). Hybridization to a 1:100 dilution of ankyrin G antibody (EMD Biosciences, Pasadena, CA) was performed and signals were visualized using the SuperSignal West Pico Chemiluminescent Substrate kit (Pierce Biotechnology, Rockford, IL). Densitometer analysis was performed on a GS-700 Imaging Densitometer (Bio-Rad Laboratories). Membranes were washed with TBS and stripped with stripping buffer (25 mM glycine, 1% SDS, pH 2.0) for 30 min, followed by a TBS wash and hybridization to a 1:25,000 dilution of anti-tubulin antibody (Sigma, Mississauga, ON). The level of tubulin protein expression was used as the internal standard for normalizing ankyrin protein expression.
Results
DNA sequence analysis
Genomic DNA sequencing for the LMNA coding region and intron-exon boundaries confirmed heterozygosity for the LMNA c.2036C>T splicing mutation only in the HGPS cell line, with no other LMNA sequence abnormality found in either cell line.
Cell and nuclear morphology
When compared to normal fibroblasts, HGPS fibroblasts had shrunken cell bodies with coarse cell membranes beginning in early passages (Fig. 1). HGPS fibroblasts lost inhibition of growth from cell-cell contact and grew in clustered cell stacks. HGPS fibroblasts had numerous abnormalities of nuclear size and shape compared to normal fibroblasts (Fig. 2), including larger nuclear size with herniations into cytoplasm, nuclear membrane folding, crevices and fragmentation.
a, b Morphology of Hutchinson-Gilford progeria syndrome (HGPS) and normal fibroblasts. Hoffman-contrast photomicrographs showing primary culture of normal (a) and HGPS fibroblasts (b). Fibroblasts were cultured as described, seeded at 2×105 per 100 mm dish and were passaged twice weekly. Compared with normal fibroblasts, HGPS fibroblasts showed shrunken cell bodies, coarse cell membranes, and grew in clustered cell stacks. c, d Ki-67 antigen detection in proliferating cells for normal and HGPS fibroblasts. Fluorescent photomicrographs of immunostaining using anti-Ki-67 antibody in normal (c) and HGPS (d) fibroblasts (APD13), visualized with FITC-conjugated secondary antibody (green). Nuclei are stained with Hoechst dye (blue). Ki-67 antigen is detected only in proliferating cells and is associated with reforming nuclei
Nuclear morphology in normal and HGPS cultured fibroblasts. Images from phase-contrast photomicrographs show the nuclear morphology from normal fibroblasts (a) and HGPS fibroblasts (b–e). Cells were grown on glass coverslips, fixed and stained with Hoechst dye, which binds to nuclear DNA. Abnormalities such as herniation (b), crevices (c), fragmentation (d), and severe fragmentation (e) were observed in HGPS fibroblasts
Cell growth
At passages 1 to 5, HGPS fibroblasts showed an accelerated growth rate, as assessed by APD, compared to normal fibroblasts (Fig. 3). Thereafter, HGPS cells failed to thrive at later passages, with a rapid decrease in APD compared to normal fibroblasts (Fig. 3). This was independently confirmed by an observed decrease in the fraction of proliferating cells as detected by indirect immunofluorescence staining with anti-Ki67 antibody (Kill 1996; Bridger and Kill 2004).
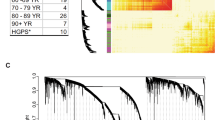
Growth kinetics of cultured fibroblasts. a Growth curves for normal and HGPS (progeria) fibroblasts shown in the colors indicated. Accumulated population doublings (APDs) were calculated using the formula (logH−logS)/log2.0, where logH is and logS are logarithms of the number of cells harvested and seeded, respectively. HGPS fibroblasts showed early accelerated growth followed by rapid decline in proliferation in later passages compared to normal fibroblasts. b Histogram of number of population doublings achieved at the indicated passage number for normal and HGPS fibroblasts. Cells were seeded at a density of 2×105 per dish and passaged twice weekly to maintain the cells in log-phase growth
DNA microarray
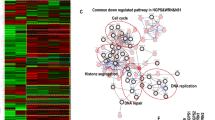
Total RNA isolated from three different ages—young, middle-aged and old—of normal and HGPS fibroblasts was studied using GeneChip Expression Arrays HU133A. A total of 66 unique genes (see Table 1) were differentially expressed in HGPS compared to normal fibroblasts based on greater than 2-fold differences in expression. Of these, 27 and 39 were up- and down-regulated, respectively. Gene ontology analysis found that genes with differential expression levels between HGPS and normal cells were associated with cell growth and cell morphology, such as cell development, proliferation, adhesion, cytoskeletal anchoring and regulation of cell cycle. Among these genes, ANK3, encoding ankyrin G, had markedly differential expression between HGPS and normal cells at all three ages: 112-, 101- and 72-fold up-regulated in HGPS young, middle and old age compared to normal cells (Table 1, Fig. 4).
Microarray data. Microarrays were normalized to the 50th percentile (per chip) and normalized to median intensity (per gene). Robust multi-array average (RMA) normalization was used for pre-processing. The experiment was set up to compare normal versus HGPS samples. The relative expression profile of ANK3 is bracketed by arrows
Quantitative reverse-transcriptase polymerase chain reaction
After target normalization, the lower expressed target was used as calibrator (baseline) to calculate the fold-difference between two targets in HGPS and normal fibroblasts. For ANK3, HGPS cells had 750-fold more target RNA than the normal fibroblasts. In contrast, for SEPP1, normal fibroblasts had 892-fold more target RNA of compared to HGPS fibroblasts.
Ankyrin G expression and localization
In order to confirm the microarray findings at the protein level, total protein extracts from cultured HGPS and normal fibroblasts were used for Western analysis with an anti-ankyrin G antibody. Compared to normal fibroblasts, ankyrin G concentration in HGPS cells was dramatically increased in repeat assays from samples taken at different passage numbers. Six groups of protein extracts from HGPS and normal fibroblasts were studied, with tubulin mass used as the internal control. The fold-increase of normalized ankyrin G mass in HGPS compared to normal cells was 9.9±3.9 (Fig. 5; P =0.008).
We next examined the cellular localization of ankyrin G by indirect immunostaining using the anti-ankyrin G antibody. We observed that in HGPS fibroblasts, ankyrin G was widely distributed throughout the cytoplasm, while in normal fibroblasts ankyrin G was not seen in any cellular compartment (Fig. 6).
Ankyrin G localization in cultured fibroblasts. Fluorescent photomicrographs showing ankyrin G immunostaining of HGPS fibroblasts visualized with FITC-conjugated secondary antibody (green) (b–e). Ankyrin G is distributed throughout the cytoplasm in HGPS fibroblasts. Nuclei are stained with Hoechst dye (blue). No ankyrin G staining at all was seen in triplicates from control fibroblasts (a)
Discussion
The principal novel findings of this study are: (1) documentation of widespread cellular and nuclear morphological abnormalities and abnormal growth kinetics of HGPS fibroblasts compared with normal fibroblasts in culture; (2) development of a filtering strategy based upon differential mRNA expression between HGPS and normal fibroblasts at three distinct ages in culture; (3) creating a list of genes with different expression profiles between HGPS and normal fibroblasts from this filtering strategy; (4) validation of the mRNA overexpression in HGPS cells of one gene so identified, namely ANK3, using Q-RT-PCR of total mRNA, Western analysis of cell lysates and immunostaining of intact cells; and (5) creating a second list of genes that showed similar expression behavior to ANK3 under these culture conditions.
Molecular profiling of HGPS fibroblasts containing the most common disease mutation, LMNA c.2036C>T, revealed that ankyrin G is a candidate protein whose mRNA and protein expression was markedly increased in vitro in HGPS fibroblasts compared to normal fibroblasts. Ankyrins are a family of proteins that are believed to link the integral membrane proteins to the underlying spectrin–actin cytoskeleton. They play several key roles in activities such as cell motility, activation, proliferation, contact and the maintenance of specialized membrane domains. ANK3 is expressed primarily in the central nervous system, but is also expressed in other tissues (Kordeli et al. 1995). While no human mutations in ANK3 have been reported, a mutation in the SCN5A gene (MIM 600163.0033) interferes with ankyrin G in cardiomyocytes resulting in Brugada syndrome (MIM 601144), a dominantly inherited cardiac arrhythmia (Mohler et al. 2004).
The marked increase in ankyrin G expression in cultured HGPS fibroblasts compared with extremely low expression in normal fibroblasts might be a fundamental pathway that contributes to the progression to early senescence. Alternatively, high ankyrin G in HGPS cells might be compensatory and secondary to the profound changes in cellular phenotype related to accelerated senescence. Finally, the ankyrin G overexpression might be a non-specific epiphenomenon that has no direct relationship with the abnormal cell biology of HGPS fibroblasts. It would be of interest to determine whether interfering with the over-expression of ankyrin G might affect the altered morphology or dysregulated growth kinetics of cultured HGPS cells. There may be other genes from Table 2, apparently co-ordinately regulated with ANK3 that may have more a direct mechanistic relationship with the abnormal phenotypes of HGPS cells.
In summary, we report an approach with which to filter the mRNA expression differences between HGPS and normal fibroblasts in culture. The filtering strategy hinged on identification of differentially expressed transcripts that were consistent across three distinct stages of culture based on cell passage number. This filtering strategy produced a list of 66 unique differentially expression genes, of which ∼40% were upregulated (and ∼60% were down-regulated) in HGPS cells (Table 1). We validated the marked increase in mRNA expression in HGPS cells that was seen for one gene from this list—ANK3—using Q-RT-PCR, Western analysis of cultured cell lysates and immunofluorescence microscopy. We then created a second list of genes that behaved similar to ANK3 in the cell culture experiments. The findings identify ANK3/ankyrin G as a possible biomarker of senescence in cultured HGPS cells at all stages. Other genes from Table 2 might also be investigated as possible candidates related to progeria and aging.
References
Bridger JM, Kill IR (2004) Aging of Hutchinson-Gilford progeria syndrome fibroblasts is characterised by hyperproliferation and increased apoptosis. Exp Gerontol 39:717–724
Carter DE, Robinson JF, Allister EM, Huff MW, Hegele RA (2005) Quality assessment of microarray experiments. Clin Biochem 38:639–642
Cao H, Hegele RA (2003) LMNA is mutated in Hutchinson-Gilford progeria (MIM 176670) but not in Wiedemann-Rautenstrauch progeroid syndrome (MIM 264090). J Hum Genet 48: 271–274
Csoka AB, Cao H, Sammak PJ, Constantinescu D, Schatten GP, Hegele RA (2004a) Novel lamin A/C gene (LMNA) mutations in atypical progeroid syndromes. J Med Genet 41:304–308
Csoka AB, English SB, Simkevich CP, Ginzinger DG, Butte AJ, Schatten GP, Rothman FG, Sedivy JM (2004b) Genome-scale expression profiling of Hutchinson-Gilford progeria syndrome reveals widespread transcriptional misregulation leading to mesodermal/mesenchymal defects and accelerated atherosclerosis. Aging Cell 3:235–243
De Sandre-Giovannoli A, Bernard R, Cau P, Navarro C, Amiel J, Boccaccio I, Lyonnet S, Stewart CL, Munnich A, Le Merrer M, Levy N (2003) Lamin A truncation in Hutchinson-Gilford progeria. Science 300:2055
Eriksson M, Brown WT, Gordon LB, Glynn MW, Singer J, Scott L, Erdos MR, Robbins CM, Moses TY, Berglund P, Dutra A, Pak E, Durkin S, Csoka AB, Boehnke M, Glover TW, Collins FS (2003) Recurrent de novo point mutations in lamin A cause Hutchinson-Gilford progeria syndrome. Nature 423:293–298
Heyer LJ, Kruglyak S, Yooseph S (1999) Exploring expression data: identification and analysis of coexpressed genes. Genome Res 9:1106–1115
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Kill IR (1996) Localisation of the Ki-67 antigen within the nucleolus evidence for a fibrillarin-deficient region of the dense fibrillar component. J Cell Sci 109:1253–1263
Kordeli E, Lambert S, Bennett V (1995) Ankyrin-G: a new ankyrin gene with neural-specific isoforms localized at the axonal initial segment and node of Ranvier. J Biol Chem 270:2352–2359
Li C, Wong WH (2001) Model-based analysis of oligonucleotide arrays: expression index computation and outlier detection. Proc Natl Acad Sci USA 98:31–36
Mohler PJ, Rivolta I, Napolitano C, LeMaillet G, Lambert S, Priori SG, Bennett V (2004) Na(v)1.5 E1053K mutation causing Brugada syndrome blocks binding to ankyrin-G and expression of Na(v)1.5 on the surface of cardiomyocytes. Proc Natl Acad Sci USA 101:17533–17538
Pollex RL, Hegele RA (2004) Hutchinson-Gilford progeria syndrome. Clin Genet 66:375–381
Acknowledgments
R.A.H. is supported by the Jacob J. Wolfe Distinguished Medical Research Chair, the Edith Schulich Vinet Canada Research Chair (Tier I) in Human Genetics, a Career Investigator award from the Heart and Stroke Foundation of Ontario. J.G.P. is supported by a Career Investigator award from the Heart and Stroke Foundation of Ontario. Operating support came from the Canadian Institutes for Health Research, the Heart and Stroke Foundation of Ontario (PRG4854), the Ontario Research and Development Challenge Fund (Project #0507) and by Genome Canada. C.M.W. was the recipient of a summer student award from the Ontario Genomics Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, J., Robinson, J.F., O’Neil, C.H. et al. Ankyrin G overexpression in Hutchinson-Gilford progeria syndrome fibroblasts identified through biological filtering of expression profiles. J Hum Genet 51, 934–942 (2006). https://doi.org/10.1007/s10038-006-0042-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-006-0042-0
Keywords
This article is cited by
-
Global gene expression in granulosa cells of growing, plateau and atretic dominant follicles in cattle
Reproductive Biology and Endocrinology (2015)
-
Screening of candidate genes in fibroblasts derived from patients with Dupuytren’s contracture using bioinformatics analysis
Rheumatology International (2015)