Abstract
The aim of the present study was to evaluate a new improved multiplex polymerase chain reaction (PCR) hybridisation assay to detect multidrug-resistant tuberculosis. The assay, developed to detect rifampin (rpoB) and isoniazid (katG) gene mutations causing Mycobacterium tuberculosis resistance, was recently extended to include inhA gene mutations that code for low-level isoniazid resistance. Interpretable results were obtained in 115 isolates and in all smear-positive clinical specimens. Rifampin resistance was correctly identified in all specimens and in 20 of 21 (95%) multidrug-resistant isolates compared to BACTEC 460TB. Isoniazid resistance correlated in 18 of 22 (82%) specimens, in 31 of 31 (100%) high-level and 24 of 28 (86%) low-level isoniazid-resistant isolates. The assay was rapid, easy to perform and directly applicable in smear-positive specimens. We predict that the assay may be a useful tool to combat and prevent new cases of multi- and extensively drug-resistant tuberculosis.
Similar content being viewed by others
Introduction
Multidrug-resistant tuberculosis (MDR-TB) is defined as disease caused by Mycobacterium tuberculosis resistant to at least rifampin (RIF) and isoniazid (INH), the two most important first-line anti-TB drugs. MDR-TB represents a serious problem for clinical management, and has become an important public health issue. The rapid detection of drug resistance permits the establishment of an effective treatment regimen, minimises the risk of further resistance development and limits the spread of drug-resistant M. tuberculosis.
Over 400,000 MDR-TB cases emerge every year, 50% amongst new TB cases and 50% in previously treated TB patients [1]. Approximately 5–7% of these cases are expected to have extensively drug-resistant TB (XDR-TB) [2]. MDR-TB and XDR-TB are associated with an extremely high mortality, especially in the human immunodeficiency virus (HIV) coinfected [3, 4]. Therefore, rapid methods to determine drug resistance are urgently in demand.
Resistance to INH is conferred by mutations within the catalase-peroxidase enzyme gene (katG in 50–95% of resistant strains), the enoyl-acyl carrier protein reductase gene (inhA in 15–35% of resistant strains) and in the ahpC–oxyR intergenic region (10–15% of resistant strains) [5–9]. Resistance to RIF is predominantly linked to mutations within the RNA polymerase, B-subunit gene (rpoB in approximately 96% of all RIF-resistant stains tested) [10].
Recently, a combined multiplex polymerase chain reaction (PCR) and DNA strip hybridisation assay, the GenoType MTBDRplus (GTplus), was further developed from the original GenoType MTBDR (GT) version (Hain Lifescience, Nehren, Germany). The technique combines a multiplex PCR followed by the hybridisation to specific membrane-bound probes for the identification of either wild-type (WT) or specific mutations. In addition to the detection of mutations in the genes conferring resistance to RIF (rpoB) and high-level INH (katG), as applied in the original version, probes to detect mutations within the inhA gene causing low-level INH resistance have been included. In the new version of the DNA strip, the inhA gene is covered by two inhA WT (−15, −16 and −8 nucleic acid positions) and four inhA mutation probes (C15T, A16G, T8C and T8A). In addition, three more rpoB WT probes targeting codons 505–533 were included.
The aim of the present study was to evaluate the performance of the new GTplus assay for the rapid detection of mutations causing RIF and high- or low-level INH resistance in M. tuberculosis isolates and directly in clinical specimens. The GTplus assay results were compared to drug susceptibility testing (DST) performed by the proportion method BACTEC 460TB (Becton Dickinson Diagnostic Instruments Systems, Sparks, MD).
Materials and methods
The study was carried out at the International Reference Laboratory of Mycobacteriology, Copenhagen, Statens Serum Institute (SSI), Denmark.
Isolates and clinical specimens
A total of 115 M. tuberculosis isolates, 53 from SSI, Denmark, and 62 from the National TB Reference Laboratory in Lithuania, were included. In order to test the performance of the additional inhA gene mutation, the Danish isolates consisted of 28 low-level (0.1 μg/ml) INH-resistant isolates, five high-level (0.4 μg/ml) INH-resistant and 20 susceptible strains. The Lithuanian isolates included a mixture of multidrug-resistant and susceptible strains representing 20% of the MDR-TB isolates and 10% of the non-MDR-TB isolates from Lithuania in 2006.
Eighty-one pretreated clinical specimens that consisted of 61 respiratory specimens (sputum, tracheal secretion, bronchioalveolar lavage) and 20 non-respiratory specimens (12 gastric lavage, six lymph node and two tissue biopsies) were included. Specimens were selected based on smear grade, nucleic acid amplification, culture and DST results and were processed by conventional procedures as described elsewhere [11]. Smears were stained with auramine-rhodamine and examined by fluorescent microscopy [12]. Nucleic amplification was performed by strand displacement amplification as previously described [13].
The Lithuanian isolates were sent for reference purposes to SSI and were, therefore, of no consequence to the patients from which they had been obtained. None of the Danish patients had prohibited the use of their tissue specimens for research (the Registry of Human Tissue Utilization, National Board of Health, Denmark).
Drug susceptibility testing
DST for RIF, INH, ethambutol and pyrazinamide was performed in Denmark by using the BACTEC 460TB system, according to the manufacturer’s instructions [14]. BACTEC 460TB was used as the gold standard and the critical concentrations were 2.0 μg/ml for RIF and 0.1 μg/ml for INH. In addition, INH was tested at 0.4 μg/ml, and 2.0 μg/ml in the BACTEC 460TB system. High-level INH resistance was defined as INH resistance at a concentration ≥0.4 μg/ml according to the Clinical and Laboratory Standards Institute (CLSI, formerly the National Committee for Clinical Laboratory Standards [NCCLS]) [15]. The critical concentrations used were 2.5 μg/ml for ethambutol and 100 μg/ml for pyrazinamide. When drug resistance was observed for RIF and/or INH, additional DST was performed at concentrations of 1.0 μg/ml for amikacin and 2.0 μg/ml for ofloxacin.
GenoType MTBDRplus assay
The new DNA strip includes the following probes: conjugate control (CC), amplification control (AC), M. tuberculosis complex control (TUB), specific gene locus controls (rpoB, katG, inhA), WT sequences (rpoB WT1–8, katG WT, inhA WT1–2), the four most frequent rpoB gene mutations (D516V, H526Y, H526D, S531L), the two mutations at codon 315 in the katG gene (S315T1, S315T2) and the four mutations in the inhA promoter region (C15T, A16G, T8C, T8A).
Aliquots of 1,000 μl of M. tuberculosis isolate in Dubos medium (SSI, Copenhagen, Denmark) or a volume of frozen surplus material (−20°C) of 500 μl NaOH-N-acetyl-L-cysteine-pretreated clinical specimen were centrifuged at 10,000×g for 15 min. The pellet was resuspended in 100 μl of distilled water, vortexed, heat inactivated for 20 min at 100°C, lysed by ultrasonification for 15 min at 60°C and centrifuged at 13,000×g for 5 min.
For amplification, a mixture consisting of 35 μl primer nucleotide mix, 5 μl of 10× PCR buffer with 20 mM MgCl2 (isolates) or 5 μl of 10× PCR buffer without MgCl2 (clinical specimens), 1 μl of 25 mM MgCl2 (isolates) or 3 μl of 25 mM MgCl2 (clinical specimens), 2 U of FastStart Taq DNA polymerase (Roche Applied Science, Mannheim, Germany) and 5 μl supernatant in a final volume of 50 μl was prepared.
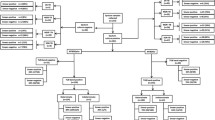
Amplification was performed in a thermocycler (Perkin Elmer 9600 thermal cycler; Perkin Elmer Corporation, Norwalk, CT) using the following amplification profile: initial denaturation of 5 min at 95°C, followed by 10 cycles of 30 s at 95°C and 2 min at 58°C, and 20 cycles (isolates) or 30 cycles (clinical specimens) of 25 s at 95°C, 40 s at 53°C and 40 s at 70°C and a final extension step of 8 min at 70°C. Amplification was visualised by agarose electrophoresis: amplification control (63 bp), M. tuberculosis complex control (115 bp), rpoB (166 bp), katG (120 bp) and inhA (110 bp). Hybridisation was performed using a pre-programmed Auto-LiPA system (InnoGenetics, Zwijnaarde, Belgium) and the GTplus DNA strips were interpreted according to the manufacturer’s instructions, as shown in Fig. 1.
Representative DNA strip patterns obtained with the GenoType MTBDRplus (GTplus) assay. The positions of the oligonucleotide probes are given on the left. The target genes and specific probe lines are shown from the top to bottom as follows: conjugate control; amplification control (23S rRNA); Mycobacterium tuberculosis complex-specific control (23S rRNA); locus control of rpoB amplification; eight rpoB wild-type (WT) probes; four rpoB mutant probes with mutations in codons 516, 526 or 531; locus control of katG amplification; one katG codon 315 WT probe; two katG probes with mutations in codon 315; locus control of inhA amplification; two inhA WT probes; and four mutation probes in the inhA promoter region (−15C/T, −16A/G, −8T/C and −8T/A). Samples with different strip susceptibility patterns are shown on the right. Lane 1, drug-susceptible (rpoB, katG, inhA WT); lane 2, rifampin (RIF)-susceptible, low-level isoniazid (INH)-resistant (rpoB, katG WT, inhA C15T mutation); lane 3, multidrug-resistant tuberculosis (MDR-TB) (rpoB S531L, katG S315T1, inhA C15T mutation); lane 4, MDR-TB (rpoB H526D, katG S315T1, inhA C15T mutation); lane 5, RIF susceptible, high-level INH-resistant (rpoB, inhA WT and absent katG locus control and katG WT is lacking)
Sequencing
The DNA sequence analysis of rpoB, katG, inhA and oxyR–ahpC was performed for samples having discrepant results between DST and the assay. For the one isolate which had a discrepant result by genotype that was susceptible to RIF, the rpoB gene was subjected to sequencing by a previously described method [16]. Sequencing of the four discrepant isolates and the four isolates obtained from the four discrepant clinical samples was performed in the following regions: katG (codons 315 and 463), fabG1–inhA and the oxyR–ahpC intergenic region using primers previously described [17–19]. The PCR conditions used for the katG codon 315, the fabG–inhA and the oxyR–ahpC intergenic region were as follows: primers katg1F and katg4F (katG), 519 and 520 (fabG1–inhA) and tomap1 and tomap2 (oxyR–ahpC) using AmpliTaq Gold (Applied Biosystems, Foster City, CA) and 1.5 mM MgCl2 (2.5 mM for oxyR–ahpC), with 30 cycles of 1 min at 95°C, 40 s at 62°C and 1 min at 72°C. PCR conditions for the katG codon 463 analysis were as follows: primers 542 and 543 were used with the GC-RICH PCR system (Roche Applied Science, Mannheim, Germany) using 0.3-mM GC-RICH resolution and 40 cycles of 1 min at 95°C, 30 s at 50°C and 90 s at 72°C. The products were sequenced using the BigDye Terminator v1.1 Cycle Sequencing Kit (Applied Biosystems, Foster City, CA).
Results
Drug susceptibility results
Among the 62 Lithuanian isolates analysed by genotype, 21 were resistant to RIF and INH (MDR-TB), five were high-level INH mono-resistant and 36 were susceptible. Only one of the 21 (5%) MDR isolates also had XDR M. tuberculosis. The 53 Danish isolates included 28 low-level INH mono-resistant isolates, five high-level INH mono-resistant and 20 susceptible strains. DST performed on the 61 culture-positive clinical specimens identified one isolate which was MDR M. tuberculosis and 21 were monodrug-resistant to INH, of which, seven were low-level INH-resistant and 39 were found to be fully susceptible.
GTplus performance in isolates
Sufficient amplification and interpretable genotype results was obtained in all isolates. The genotype results were concordant with BACTEC results in 61 of 62 (98%) Lithuanian isolates and in 49 of 53 (93%) Danish isolates.
Lithuanian isolates
The GTplus assay identified the presence of the katG S315T1 mutation in all of the INH-resistant Lithuanian isolates. The data are presented in Table 1. Three of the 26 (12%) isolates with the katG codon 315 gene mutation had an additional mutation in the inhA promoter region C15T. Mutation and WT mismatch for katG and inhA correlated in 25 of 26 (96%) isolates. One isolate carried the katG S315T1 mutation; however, WT was not absent, possibly indicating a mixture of INH-resistant and -susceptible bacteria. All of the four different rpoB type mutations present on the DNA strip were detected in the 21 RIF-resistant isolates. The most prevalent rpoB mutation was S531L, accounting for 18 of 21 (86%) isolates. Only one Lithuanian strain contained both the rpoB S531 and the rpoB D516Y mutations. One isolate that gave a discrepant result compared to BACTEC 460TB was fully susceptible to RIF; however, it showed no rpoB mutations present by genotype, although the rpoB WT 8 probe was lacking. Sequencing of this isolate revealed an rpoB L533M mutation with a nucleotide substitution in codon 533; CTG (Leu) to ATG (Met), as shown in Table 2.
Danish isolates
The GTplus assay detected a mutation in the inhA promoter region C15T in 24 of 28 (86%) low-level INH isolates. Four of the low-level INH-resistant isolates had no mutations detected by the GTplus assay. Sequencing revealed that three of the discrepant isolates by BACTEC 460TB had a G→A mutation at position −G49A in the oxyR–ahpC intergenic region (Table 2). In one of the isolates, the C-terminal region of the katG 463 region was not amplified by PCR and, in two other isolates, only a weak amplification was observed for this genetic region. These findings in the katG 463 region were not further analysed. Three of the five high-level INH-resistant Danish isolates showed the S315T1 katG mutation. In two of the five high-level INH-resistant Danish isolates, the genotype showed that the locus control probe band for the katG gene was absent, as shown in Fig. 1. These findings indicated partial deletion of the katG gene, which was verified by PCR. The genotype correctly identified all 20 Danish INH-susceptible isolates when compared to BACTEC 460TB.
GTplus performance in clinical specimens
Interpretable GTplus assay results were found in 62 of 81 (77%) clinical specimens tested. The GTplus assay detected INH resistance correctly in 18 of 22 (82%) INH-resistant specimens, as shown in Table 3. There were four discrepant results between the drug susceptibility patterns and those obtained by the GTplus assay concerning INH resistance. All had high-level INH resistance with the inhA mutation C15T, which normally confers low-level INH resistance; this mutation was confirmed by sequencing (Table 2). Nine of the 22 (41%) INH-resistant specimens had the katG S315T1 mutation and 11 of 22 (50%) had the inhA C15T mutation. The high-level INH-resistant specimens had the katG S315T1 mutation in 9 of 15 (60%) specimens and the low-level INH-resistant specimens showed the inhA C15T mutation in 7 of 7 (100%) specimens. In two INH high-level resistant specimens, the katG WT was absent; however, no katG mutations were detected. This was most probably due to the katG mutation being located elsewhere in the gene. Only the rpoB S531L mutation was identified in all RIF-resistant specimens, and all of these were MDR-TB specimens. Interpretable DNA strip readings were obtained in 42 of 42 (100%) smear-positive respiratory specimens and in 6 of 6 (100%) smear-positive non-respiratory specimens, regardless of the DST result. Only 5 of 19 (26%) respiratory and 9 of 14 (64%) non-respiratory smear-negative specimens revealed interpretable genotype results. The data are presented in Table 4.
Discussion
The GTplus assay gave interpretable results when applied directly in all of the smear-positive clinical specimens, including both respiratory and non-respiratory samples, regardless of the drug susceptibility result. However, interpretable results were only obtained in 42% of the smear-negative specimens. As the new GTplus version extensively covers the rpoB mutations responsible for RIF resistance, the catalase-peroxidase katG codon 315 mutations and, additionally, the enoyl ACP reductase inhA mutations responsible for INH resistance, the new assay has the potential for the rapid detection of MDR M. tuberculosis in smear-positive specimens.
Previous studies have demonstrated that the first version of the GT assay, which only included mutations in rpoB and codon 315 of the katG gene, had a high performance in isolates. The GT assay detected 92–100% and 67–89% of the M. tuberculosis strains resistant to RIF and INH, respectively [5, 20, 21]. Two studies compared the GT assay with INNO-LiPA Rif.TB (Innogenetics, Zwijnaarde, Belgium) in 41 and 52 M. tuberculosis isolates, respectively. These studies showed that the genotypic results were concordant with phenotypic DST for INH in 73% and 90% of strains, whereas it was 95% and 98% for RIF, respectively [22, 23]. The advantage of the GT assay compared to INNO-LiPA Rif.TB is the detection of resistance to both RIF and INH. Likewise, we found that RIF resistance was correctly identified by the new assay in 98% of the Lithuanian isolates tested. One strain susceptible by BACTEC 460TB had an absent rpoB WT band in the eighth position with no rpoB mutations present on the DNA strip. Sequencing of this strain revealed an rpoB L533M mutation, which was not developed in the GTplus assay. This mutation, which has only rarely been reported, did not appear to result in a phenotypic change in DST, and it remains to be determined whether the rpoB L533M mutation is associated with RIF resistance [24]. The most prevalent mutation found was the rpoB S531L in 86% of the Lithuanian isolates. The rpoB S531L mutation was identified in all Danish RIF-resistant clinical specimens in concordance with a previous study [25]. Studies have shown that mutations at codon 531 within the rpoB gene are the most frequent worldwide [10]. All of the Lithuanian and Danish high-level INH-resistant isolates were concordant with BACTEC 460TB results. The katG S315T1 mutation accounted for 100% of the INH-resistant Lithuanian isolates. Complete deletion of the katG gene is a rare phenomenon that results in high-level INH resistance [26]. In our study, we found two isolates which showed the partial absence of the katG gene and high-level INH resistance in BACTEC 460TB. The low-level INH-resistant isolates showed a high (86%) correlation between genotypic and phenotypic results, indicating that the new GTplus assay effectively detects low-level INH resistance. Three isolates had a previously undescribed point mutation in position −49 (G→A) in the oxy–ahpC region. It is possible that the identified mutation in position −49 (G49A) of the oxy–ahpC intergenic region is associated with low-level INH resistance, but this remains to be further investigated. Overall, we find that the inclusion of the inhA probes for INH resistance in the new GTplus version assay improves the ability of the assay to detect INH resistance.
The first version of the GT assay was optimised for application directly in pretreated clinical specimens and was found to have a high performance in several studies. In 106 clinical specimens, interpretable GT results were achieved in 95% of the 38 smear-positive respiratory specimens [27]. Another study, which tested the GT assay directly in 42 smear-positive sputum specimens, found 100% concordance with conventional drug susceptibility testing [28]. In a study where the GT assay was directly used in 143 smear-positive specimens, a sensitivity of 94% for katG and 91% for rpoB was achieved and the assay correctly identified INH resistance in 84% of specimens with high-level INH resistance and 96% of specimens with RIF resistance [29]. Recently, studies on the new extended GTplus assay have found, similarly to our findings, that it could be applied directly in smear-positive specimens [30, 31]. One study also included 100 smear-negative M. tuberculosis specimens and found for RIF interpretable results in 16 of the 20 (80%) specimens that were culture-positive [31]. These findings for the smear-negative samples were higher than our findings.
In conclusion, the GTplus assay is rapid (<48 h) and easy to perform for the simultaneous detection of RIF and low- or high-level INH resistance in M. tuberculosis isolates and clinical specimens obtained from patients suspected of having TB. Our study demonstrated a high concordance between GTplus assay resistance patterns and those obtained by the conventional BACTEC 460TB system. In laboratories with well-established M. tuberculosis culture and DST systems, we propose that the GTplus assay should be used as a supplement to the gold standard DST in obtaining a rapid preliminary DST result, as not all clinically relevant INH mutations are covered by the assay. The assay may be applied to smear-positive samples from patients suspected of treatment failure, recurrent TB, drug-resistant TB exposure or cases originating from countries with high levels of drug resistance. However, in countries that have yet to implement DST, the GTplus assay may be a valuable tool for the detection of MDR-TB until conventional DST facilities become available. We predict that the new extended GTplus version assay may be a useful tool for the World Health Organization (WHO) Global Task Force to combat and prevent new cases of MDR-TB and XDR-TB.
References
World Health Organization (WHO) (2006) Global tuberculosis control: surveillance, planning, financing. WHO, Geneva, Switzerland
Centers for Disease Control and Prevention (CDC) (2006) Emergence of Mycobacterium tuberculosis with extensive resistance to second-line drugs—worldwide, 2000–2004. Morb Mortal Wkly Rep 55:301–305
Espinal MA, Kim SJ, Suarez PG, Kam KM, Khomenko AG, Migliori GB, Baéz J, Kochi A, Dye C, Raviglione MC (2000) Standard short-course chemotherapy for drug-resistant tuberculosis: treatment outcomes in 6 countries. JAMA 283:2537–2545
Gandhi NR, Moll A, Sturm AW, Pawinski R, Govender T, Lalloo U, Zeller K, Andrews J, Friedland G (2006) Extensively drug-resistant tuberculosis as a cause of death in patients co-infected with tuberculosis and HIV in a rural area of South Africa. Lancet 368:1575–1580
Hillemann D, Weizenegger M, Kubica T, Richter E, Niemann S (2005) Use of the genotype MTBDR assay for rapid detection of rifampin and isoniazid resistance in Mycobacterium tuberculosis complex isolates. J Clin Microbiol 43:3699–3703
Mokrousov I, Narvskaya O, Otten T, Limeschenko E, Steklova L, Vyshnevskiy B (2002) High prevalence of KatG Ser315Thr substitution among isoniazid-resistant Mycobacterium tuberculosis clinical isolates from northwestern Russia, 1996 to 2001. Antimicrob Agents Chemother 46:1417–1424
Musser JM, Kapur V, Williams DL, Kreiswirth BN, van Soolingen D, van Embden JD (1996) Characterization of the catalase-peroxidase gene (katG) and inhA locus in isoniazid-resistant and -susceptible strains of Mycobacterium tuberculosis by automated DNA sequencing: restricted array of mutations associated with drug resistance. J Infect Dis 173:196–202
Zhang M, Yue J, Yang YP, Zhang HM, Lei JQ, Jin RL, Zhang XL, Wang HH (2005) Detection of mutations associated with isoniazid resistance in Mycobacterium tuberculosis isolates from China. J Clin Microbiol 43:5477–5482
Telenti A, Honoré N, Bernasconi C, March J, Ortega A, Heym B, Takiff HE, Cole ST (1997) Genotypic assessment of isoniazid and rifampin resistance in Mycobacterium tuberculosis: a blind study at reference laboratory level. J Clin Microbiol 35:719–723
Telenti A, Imboden P, Marchesi F, Lowrie D, Cole S, Colston MJ, Matter L, Schopfer K, Bodmer T (1993) Detection of rifampicin-resistance mutations in Mycobacterium tuberculosis. Lancet 341:647–650
Johansen IS, Lundgren B, Thyssen JP, Thomsen VØ (2002) Rapid differentiation between clinically relevant mycobacteria in microscopy positive clinical specimens and mycobacterial isolates by line probe assay. Diagn Microbiol Infect Dis 43:297–302
Smithwick RW (1976) Laboratory manual for acid-fast microscopy, 2nd edn. Centers for Disease Control, Atlanta
Johansen IS, Thomsen VØ, Johansen A, Andersen P, Lundgren B (2002) Evaluation of a new commercial assay for diagnosis of pulmonary and nonpulmonary tuberculosis. Eur J Clin Microbiol Infect Dis 21:455–460
Siddiqi SH (1996) BACTEC 460TB system. Product and procedure manual. Becton Dickinson Microbiology Systems, Sparks, Maryland
Clinical and Laboratory Standards Institute (CLSI) (2003) Susceptibility testing of mycobacteria, nocardiae, and other aerobic actinomycetes. Approved standard M24-A. National Committee for Clinical Laboratory Standards (NCCLS), Wayne, Pennsylvania
Bártfai Z, Somoskövi A, Ködmön C, Szabó N, Puskás E, Kosztolányi L, Faragó E, Mester J, Parsons LM, Salfinger M (2001) Molecular characterization of rifampin-resistant isolates of Mycobacterium tuberculosis from Hungary by DNA sequencing and the line probe assay. J Clin Microbiol 39:3736–3739
Mokrousov I, Otten T, Filipenko M, Vyazovaya A, Chrapov E, Limeschenko E, Steklova L, Vyshnevskiy B, Narvskaya O (2002) Detection of isoniazid-resistant Mycobacterium tuberculosis strains by a multiplex allele-specific PCR assay targeting katG codon 315 variation. J Clin Microbiol 40:2509–2512
Lavender C, Globan M, Sievers A, Billman-Jacobe H, Fyfe J (2005) Molecular characterization of isoniazid-resistant Mycobacterium tuberculosis isolates collected in Australia. Antimicrob Agents Chemother 49:4068–4074
Baker LV, Brown TJ, Maxwell O, Gibson AL, Fang Z, Yates MD, Drobniewski FA (2005) Molecular analysis of isoniazid-resistant Mycobacterium tuberculosis isolates from England and Wales reveals the phylogenetic significance of the ahpC −46A polymorphism. Antimicrob Agents Chemother 49:1455–1464
Brossier F, Veziris N, Truffot-Pernot C, Jarlier V, Sougakoff W (2006) Performance of the genotype MTBDR line probe assay for detection of resistance to rifampin and isoniazid in strains of Mycobacterium tuberculosis with low- and high-level resistance. J Clin Microbiol 44:3659–3664
Miotto P, Piana F, Penati V, Canducci F, Migliori GB, Cirillo DM (2006) Use of genotype MTBDR assay for molecular detection of rifampin and isoniazid resistance in Mycobacterium tuberculosis clinical strains isolated in Italy. J Clin Microbiol 44:2485–2491
Cavusoglu C, Turhan A, Akinci P, Soyler I (2006) Evaluation of the genotype MTBDR assay for rapid detection of rifampin and isoniazid resistance in Mycobacterium tuberculosis isolates. J Clin Microbiol 44:2338–2342
Mäkinen J, Marttila HJ, Marjamäki M, Viljanen MK, Soini H (2006) Comparison of two commercially available DNA line probe assays for detection of multidrug-resistant Mycobacterium tuberculosis. J Clin Microbiol 44:350–352
Ma X, Wang H, Deng Y, Liu Z, Xu Y, Pan X, Musser JM, Graviss EA (2006) rpoB gene mutations and molecular characterization of rifampin-resistant Mycobacterium tuberculosis isolates from Shandong Province, China. J Clin Microbiol 44:3409–3412
Johansen IS, Lundgren B, Sosnovskaja A, Thomsen VØ (2003) Direct detection of multidrug-resistant Mycobacterium tuberculosis in clinical specimens in low- and high-incidence countries by line probe assay. J Clin Microbiol 41:4454–4456
Ramaswamy S, Musser JM (1998) Molecular genetic basis of antimicrobial agent resistance in Mycobacterium tuberculosis: 1998 update. Tuber Lung Dis 79:3–29
Bang D, Andersen ÅB, Thomsen VØ (2006) Rapid genotypic detection of rifampin- and isoniazid-resistant Mycobacterium tuberculosis directly in clinical specimens. J Clin Microbiol 44:2605–2608
Hillemann D, Rüsch-Gerdes S, Richter E (2006) Application of the Genotype MTBDR assay directly on sputum specimens. Int J Tuberc Lung Dis 10:1057–1059
Somoskövi A, Dormandy J, Mitsani D, Rivenburg J, Salfinger M (2006) Use of smear-positive samples to assess the PCR-based genotype MTBDR assay for rapid, direct detection of the Mycobacterium tuberculosis complex as well as its resistance to isoniazid and rifampin. J Clin Microbiol 44:4459–4463
Hillemann D, Rüsch-Gerdes S, Richter E (2007) Evaluation of the GenoType MTBDRplus assay for rifampin and isoniazid susceptibility testing of Mycobacterium tuberculosis strains and clinical specimens. J Clin Microbiol 45:2635–2640
Barnard M, Albert H, Coetzee G, O’Brien R, Bosman ME (2008) Rapid molecular screening for multidrug-resistant tuberculosis in a high-volume public health laboratory in South Africa. Am J Respir Crit Care Med 177:787–792
Acknowledgement
We thank Hain Lifescience, Nehren, Germany, for providing the free kits.
Conflicts of interest statement
We declare no conflicts of interest.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Vijdea, R., Stegger, M., Sosnovskaja, A. et al. Multidrug-resistant tuberculosis: rapid detection of resistance to rifampin and high or low levels of isoniazid in clinical specimens and isolates. Eur J Clin Microbiol Infect Dis 27, 1079–1086 (2008). https://doi.org/10.1007/s10096-008-0548-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10096-008-0548-9