Abstract
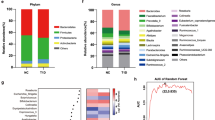
Intestinal flora changes were found in patients and animals with type 1 diabetes (T1D). However, few studies have provided any explicit clues of changes in highly disease related commensal microbiota before disease onset and their relationships with disordered peripheral immune cells. We conducted 16S rRNA microbiota analysis of non-obese diabetic (NOD) mice from weaning to diabetes onset to identify highly disease related microbes and performed Spearman correlation analysis between anomalous flora and peripheral immune cells. We found NOD mice had increased exclusive bacteria and decreased community richness or diversity, besides, with the features of decreased abundance of Bacteroidetes and increased abundance of Firmicutes, Proteobacteria or Deferribacteres and remarkable fluctuations of genus relative abundance. Furthermore, kinds of highly T1D related genus and their strong correlations with peripheral immune cells, especially neutrophils, were discovered. Microbial changes in NOD mice differed from that of ICR mice and highly disease associated microbes have strong correlations with the peripheral neutrophil ratio, which provide evidence that neutrophils are possibly involved in the pathogenesis of T1D.





Similar content being viewed by others
Availability of data and material
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- GALT:
-
Gut associated lymphoid tissues
- IBD:
-
Inflammatory bowel disease
- NETs:
-
Neutrophil extracellular trap
- NOD:
-
Non-obese diabetic
- NOR:
-
Nonobese diabetes-resistant
- PLN:
-
Pancreatic lymph node
- SCFA:
-
Short chain fatty acid
- SPF:
-
Specific pathogen-free
- T1D:
-
Type 1 diabetes
References
Alkanani AK, Hara N, Gottlieb PA et al (2015) Alterations in intestinal microbiota correlate with susceptibility to type 1 diabetes. Diabetes 64:3510–3520. https://doi.org/10.2337/db14-1847
Atarashi K, Tanoue T, Oshima K et al (2013) Treg induction by a rationally selected mixture of Clostridia strains from the human microbiota. Nature 500:232–236. https://doi.org/10.1038/nature12331
Atarashi K, Tanoue T, Ando M et al (2015) Th17 cell induction by adhesion of microbes to intestinal epithelial cells. Cell 163:367–380. https://doi.org/10.1016/j.cell.2015.08.058
Brown CT, Davis-Richardson AG, Giongo A et al (2011) Gut microbiome metagenomics analysis suggests a functional model for the development of autoimmunity for type 1 diabetes. PLoS ONE 6:e25792. https://doi.org/10.1371/journal.pone.0025792
Brown EM, Kenny DJ, Xavier RJ (2019) Gut microbiota regulation of T cells during inflammation and autoimmunity. Annu Rev Immunol 37:599–624. https://doi.org/10.1146/annurev-immunol-042718-041841
Canfora EE, Meex RCR, Venema K, Blaak EE (2019) Gut microbial metabolites in obesity, NAFLD and T2DM. Nat Rev Endocrinol 15:261–273. https://doi.org/10.1038/s41574-019-0156-z
Davis-Richardson AG, Triplett EW (2015) A model for the role of gut bacteria in the development of autoimmunity for type 1 diabetes. Diabetologia 58:1386–1393. https://doi.org/10.1007/s00125-015-3614-8
Davis-Richardson AG, Ardissone AN, Dias R et al (2014) Bacteroides dorei dominates gut microbiome prior to autoimmunity in Finnish children at high risk for type 1 diabetes. Front Microbiol 5:678. https://doi.org/10.3389/fmicb.2014.00678
de Goffau MC, Luopajarvi K, Knip M et al (2013) Fecal microbiota composition differs between children with beta-cell autoimmunity and those without. Diabetes 62:1238–1244. https://doi.org/10.2337/db12-0526
de Goffau MC, Fuentes S, van den Bogert B et al (2014) Aberrant gut microbiota composition at the onset of type 1 diabetes in young children. Diabetologia 57:1569–1577. https://doi.org/10.1007/s00125-014-3274-0
Dobranowski PA, Tang C, Sauve JP et al (2019) Compositional changes to the ileal microbiome precede the onset of spontaneous ileitis in SHIP deficient mice. Gut Microbes 10:578–598. https://doi.org/10.1080/19490976.2018.1560767
Du Teil Espina M, Gabarrini G, Harmsen HJM et al (2019) Talk to your gut: the oral-gut microbiome axis and its immunomodulatory role in the etiology of rheumatoid arthritis. FEMS Microbiol Rev 43:1–18. https://doi.org/10.1093/femsre/fuy035
Geng S, Yang L, Cheng F et al (2019) Gut microbiota are associated with psychological stress-induced defections in intestinal and blood-brain barriers. Front Microbiol 10:3067. https://doi.org/10.3389/fmicb.2019.03067
Hanninen A, Toivonen R, Poysti S et al (2018) Akkermansia muciniphila induces gut microbiota remodelling and controls islet autoimmunity in NOD mice. Gut 67:1445–1453. https://doi.org/10.1136/gutjnl-2017-314508
Honkanen J, Vuorela A, Muthas D et al (2020) Fungal dysbiosis and intestinal inflammation in children with beta-cell autoimmunity. Front Immunol 11:468. https://doi.org/10.3389/fimmu.2020.00468
Hu C, Wong FS, Wen L (2015) Type 1 diabetes and gut microbiota: friend or foe? Pharmacol Res 98:9–15. https://doi.org/10.1016/j.phrs.2015.02.006
Hu Y, Peng J, Li F et al (2018) Evaluation of different mucosal microbiota leads to gut microbiota-based prediction of type 1 diabetes in NOD mice. Sci Rep 8:15451. https://doi.org/10.1038/s41598-018-33571-z
Ilonen J, Lempainen J, Veijola R (2019) The heterogeneous pathogenesis of type 1 diabetes mellitus. Nat Rev Endocrinol 15:635–650. https://doi.org/10.1038/s41574-019-0254-y
Ivanov II, Frutos Rde L, Manel N et al (2008) Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4:337–349. https://doi.org/10.1016/j.chom.2008.09.009
Jaakkola I, Jalkanen S, Hanninen A (2003) Diabetogenic T cells are primed both in pancreatic and gut-associated lymph nodes in NOD mice. Eur J Immunol 33:3255–3264. https://doi.org/10.1002/eji.200324405
Klocperk A, Petruzelkova L, Pavlikova M, et al (2020) Changes in innate and adaptive immunity over the first year after the onset of type 1 diabetes. Acta Diabetol 57:297–307. https://doi.org/10.1007/s00592-019-01427-1
Knip M, Siljander H (2016) The role of the intestinal microbiota in type 1 diabetes mellitus. Nat Rev Endocrinol 12:154–167. https://doi.org/10.1038/nrendo.2015.218
Krych L, Nielsen DS, Hansen AK, Hansen CH (2015) Gut microbial markers are associated with diabetes onset, regulatory imbalance, and IFN-gamma level in NOD mice. Gut Microbes 6:101–109. https://doi.org/10.1080/19490976.2015.1011876
Lee C, Hong SN, Paik NY et al (2019) CD1d modulates colonic inflammation in NOD2-/- mice by altering the intestinal microbial composition comprising acetatifactor muris. J Crohns Colitis 13:1081–1091. https://doi.org/10.1093/ecco-jcc/jjz025
Lehuen A, Diana J, Zaccone P, Cooke A (2010) Immune cell crosstalk in type 1 diabetes. Nat Rev Immunol 10:501–513. https://doi.org/10.1038/nri2787
Lei M, Menon R, Manteiga S, et al (2019) Environmental chemical diethylhexyl phthalate alters intestinal microbiota community structure and metabolite profile in mice. mSystems 4:e00724–19. https://doi.org/10.1128/mSystems.00724-19
Liang Y, Wang X, He D et al (2019) Ameliorating gut microenvironment through staphylococcal nuclease-mediated intestinal NETs degradation for prevention of type 1 diabetes in NOD mice. Life Sci 221:301–310. https://doi.org/10.1016/j.lfs.2019.02.034
Liu Y, Wang X, Chen Q et al (2020) Camellia sinensis and litsea coreana ameliorate intestinal inflammation and modulate gut microbiota in dextran sulfate sodium-induced colitis mice. Mol Nutr Food Res 64:e1900943. https://doi.org/10.1002/mnfr.201900943
Menghini P, Corridoni D, Buttó LF, et al (2019) Neutralization of IL-1α ameliorates Crohn’s disease-like ileitis by functional alterations of the gut microbiome. Proc Natl Acad Sci USA 116:26717–26726. https://doi.org/10.1073/pnas.1915043116
Miranda MCG, Oliveira RP, Torres L et al (2019) Frontline science: abnormalities in the gut mucosa of non-obese diabetic mice precede the onset of type 1 diabetes. J Leukoc Biol 106:513–529. https://doi.org/10.1002/jlb.3hi0119-024rr
Murri M, Leiva I, Gomez-Zumaquero JM, et al (2013) Gut microbiota in children with type 1 diabetes differs from that in healthy children: a case-control study. BMC Med 11:46. https://doi.org/10.1186/1741-7015-11-46
Nekoua MP, Fachinan R, Fagninou A, et al (2019) Does control of glycemia regulate immunological parameters in insulin-treated persons with type 1 diabetes? Diabetes Res Clin Pract 157:107868. https://doi.org/10.1016/j.diabres.2019.107868
Parackova Z, Zentsova I, Vrabcova P et al (2020) Neutrophil extracellular trap induced dendritic cell activation leads to Th1 polarization in type 1 diabetes. Front Immunol 11:661. https://doi.org/10.3389/fimmu.2020.00661
Pearson JA, Wong FS, Wen L (2016) The importance of the Non Obese Diabetic (NOD) mouse model in autoimmune diabetes. J Autoimmun 66:76–88. https://doi.org/10.1016/j.jaut.2015.08.019
Round JL, Mazmanian SK (2010) Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc Natl Acad Sci USA 107:12204–12209. https://doi.org/10.1073/pnas.0909122107
Sorini C, Cosorich I, Lo Conte M et al (2019) Loss of gut barrier integrity triggers activation of islet-reactive T cells and autoimmune diabetes. Proc Natl Acad Sci USA 116:15140–15149. https://doi.org/10.1073/pnas.1814558116
Telieps T, Köhler M, Treise I, et al (2016) Longitudinal frequencies of blood leukocyte subpopulations differ between NOD and NOR mice but do not predict diabetes in NOD mice. J Diabetes Res 2016:4208156. https://doi.org/10.1155/2016/4208156
Turley SJ, Lee JW, Dutton-Swain N et al (2005) Endocrine self and gut non-self intersect in the pancreatic lymph nodes. Proc Natl Acad Sci USA 102:17729–17733. https://doi.org/10.1073/pnas.0509006102
Wang Y, Xiao Y, Zhong L, et al (2014) Increased neutrophil elastase and proteinase 3 and augmented NETosis are closely associated with β-cell autoimmunity in patients with type 1 diabetes. Diabetes 63:4239–4248. https://doi.org/10.2337/db14-0480
Wang K, Liao M, Zhou N et al (2019) Parabacteroides distasonis alleviates obesity and metabolic dysfunctions via production of succinate and secondary bile acids. Cell Rep 26:222–235. https://doi.org/10.1016/j.celrep.2018.12.028
Yilmaz B, Juillerat P, Oyas O et al (2019) Microbial network disturbances in relapsing refractory Crohn’s disease. Nat Med 25:323–336. https://doi.org/10.1038/s41591-018-0308-z
You Q, He DM, Shu GF et al (2019) Increased formation of neutrophil extracellular traps is associated with gut leakage in patients with type 1 but not type 2 diabetes. J Diabetes 11:665–673. https://doi.org/10.1111/1753-0407.12892
Yue X, Zhifeng G, Biyu S et al (2013) Roles of Wnt/β-catenin signaling in retinal neuron-like differentiation of bone marrow mesenchymal stem cells from nonobese diabetic mice. J Mol Neurosci 49:250–261. https://doi.org/10.1007/s12031-012-9917-z
Zagato E, Pozzi C, Bertocchi A et al (2020) Endogenous murine microbiota member Faecalibaculum rodentium and its human homologue protect from intestinal tumour growth. Nat Microbiol 5:511–524. https://doi.org/10.1038/s41564-019-0649-5
Zhou H, Sun L, Zhang S et al (2020) Evaluating the causal role of gut microbiota in type 1 diabetes and its possible pathogenic mechanisms. Front Endocrinol (lausanne) 11:125. https://doi.org/10.3389/fendo.2020.00125
Acknowledgements
We appreciate the suggestion in writing from Professor Gu f. and the technical support from Biozeron Biotech. The English language of this article was edited by American Journal Experts.
Funding
National Natural Science Foundation of China [No. 81673340, No. 81973224]; Double first-class innovation team of China Pharmaceutical University [CPU2018GF/GY16, CPU2018GF/GY17].
Author information
Authors and Affiliations
Contributions
WJ designed whole project and revised the manuscript; WY and YQ designed and performed experiments, and prepared the manuscript draft. YQ, WY and FJ analyzed data.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Consent for publication
All co-authors are consent for publication.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10482_2021_1632_MOESM1_ESM.tif
Supplementary Fig. 1: Differences in genera between ICR and NOD mice at different weeks of age. a-w: Comparisons of different genera at different weeks of age in NOD and ICR mice. a: Acetatifactor, b: Coprococcus 2, c: Lachnoclostridium 5, d: Lachnospiraceae FCS020 group, e: Lachnospiraceae NK4A136 group, f: Lachnospiraceae UCG-001, g: Lachnospiraceae UCG-010, h: Oscillibacter, i: Ruminiclostridium 6, j: Ruminococcaceae UCG-013, k: Lactobacillus, l: Faecalibaculum, m: Bacteroides, n: Muribaculum, o: Parabacteroides, p: Caulobacteraceae, q: Bradyrhizobium, r: Parasutterella, s: Rhodococcus, t: Gastranaerophilales, u: Chloroflexi, v: Sulfurovum, w: Mucispirillum. *P < 0.05, **P < 0.01, and ***P < 0.001
Rights and permissions
About this article
Cite this article
Wu, Y., You, Q., Fei, J. et al. Changes in the gut microbiota: a possible factor influencing peripheral blood immune indexes in non-obese diabetic mice. Antonie van Leeuwenhoek 114, 1669–1682 (2021). https://doi.org/10.1007/s10482-021-01632-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01632-5