Abstract
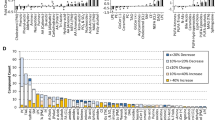
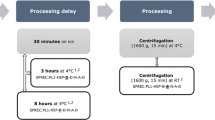
A blood pre-centrifugation delay of 24 h at room temperature influenced the proton NMR spectroscopic profiles of human serum. A blood pre-centrifugation delay of 24 h at 4°C did not influence the spectroscopic profile as compared with 4 h delays at either room temperature or 4°C. Five or ten serum freeze–thaw cycles also influenced the proton NMR spectroscopic profiles. Certain common in vitro preanalytical variations occurring in biobanks may impact the metabolic profile of human serum.




Similar content being viewed by others
Abbreviations
- SST:
-
Serum separation tube
- SPREC:
-
Standard preanalytical code
- TSP:
-
3-(trimethylsilyl) 3,3,3,3-tetradeutero-propionic acid
- CPMG:
-
Carr-Purcell-Meiboom-Gill
- HSQC:
-
Heteronuclear single quantum coherence
References
Barton RH, Nicholson JK, Elliott P, Holmes E (2008) High-throughput 1H NMR-based metabolic analysis of human serum and urine for large-scale epidemiological studies/validation study. Int J Epidemiol 37 (Suppl 1):i31–i40
Bernini P, Bertini I, Luchinat C, Nincheri P, Staderini S, Turano P (2011) Standard operating procedures for pre-analytical handling of blood and urine for metabolomics studies and biobanks. J Biomol NMR 49:231–243
Betsou F, The ISBER Working Group on Biospecimen Science (2010) Standard preanalytical coding for biospecimens: defining the sample PREanalytical code (SPREC). Cancer Epidemiol Biomarkers Prevention 19:1004–1011
Comstock GW, Burke AE, Norkus EP, Gordon GB, Hoffman SC, Helzlsouer KJ (2001) Effects of repeated freeze-thaw cycles on concentrations of cholesterol, micronutrients, and hormones in human plasma and serum. Clin Chem 47:139–142
Deprez S, Sweatman BC, Connor SC, Haselden JN, Waterfield CJ (2002) Optimisation of collection, storage and preparation of rat plasma for 1H NMR spectroscopic analysis in toxicology studies to determine inherent variation in biochemical profiles. J Pharm Biomed Anal 30:1297–1310
Eriksson L, Johansson E, Kettaneh-Wold N, Wold S (2001) Multi- and megavariate data analysis: principles and applications. Umeå, Umetrics
Fujiwara M, Kobayashi T, Jomori T, Maruyama Y, Oka Y, Sekino H, Imai Y, Takeuchi K (2009) Pattern recognition analysis for 1H NMR spectra of plasma from hemodialysis patients. Anal Bioanal Chem 394:1655–1660
Gartland KP, Sanins SM, Nicholson JK, Sweatman BC, Beddell CR, Lindon JC (1990) Pattern recognition analysis of high resolution 1H NMR spectra of urine. A nonlinear mapping approach to the classification of toxicological data. NMR Biomed 3:166–172
Gavaghan CL, Holmes E, Lenz E, Wilson ID, Nicholson JK (2000) An NMR-based metabonomic approach to investigate the biochemical consequences of genetic strain differences: application to the C57BL10J and Alpk:ApfCD mouse. FEBS Lett 484:169–174
Goldsmith P, Raj Prasad K, Ahmad N, Fisher J (2009) 1H NMR spectroscopic study of blood serum for the assessment of liver function in liver transplant patients. J Gastrointestin Liver Dis 18:508–517
Griffin JL, Shockor JP (2004) Metabolic profiles of cancer cells. Nat Rev Cancer 4:551–564
He Q, Ren P, Kong X, Wu Y, Wu G, Li P, Hao F, Tang H, Blachier F, Yin Y (2011) Comparison of serum metabolite compositions between obese and lean growing pigs using an NMR-based metabonomic approach. J Nutrit Biochem (in press)
Holmes E, Nicholls AW, Lindon JC, Ramos S, Spraul M, Neidig P, Connor SC, Connelly J, Damment SJ, Haselden J, Nicholson JK (1998) Development of a model for classification of toxin–induced lesions using 1H NMR spectroscopy of urine combined with pattern recognition. NMR Biomed 11:235–244
Howe FA, Opstad KS (2003) 1H MR spectroscopy of brain tumours and masses. NMR Biomed 16:123–131
Kaartinen J, Hiltunen Y, Kovanen PT, Ala-Korpela M (1998) Application of self-organizing maps for the detection and classification of human blood plasma lipoprotein lipid profiles on the basis of 1H NMR spectroscopy data. NMR Biomed 11:168–176
Kriat M, Confort-Gouny S, Vion-Dury J, Sciaky M, Viout P, Cozzone PJ (1992) Quantitation of metabolites in human blood serum by proton magnetic resonance spectroscopy. A comparative study of the use of formate and tsP as concentration standards. NMR Biomed 5:179–184
Le Moyec L, Valensi P, Charniot JC, Hantz E, Albertini JP (2005) Serum 1H-nuclear magnetic spectroscopy followed by principal component analysis and hierarchical cluster analysis to demonstrate effects of statins on hyperlipidemic patients. NMR Biomed 18:421–429
Lindon JC, Holmes E, Nicholson JK (2001) Pattern recognition methods and applications in biomedical magnetic resonance. Prog Nucl Magn Reson Spectrosc 39:1–40
Lindon JC, Nicholson JK, Holmes E, Keun HC, Craig A, Pearce JTM, Bruce SJ, Hardy N, Sansone SA, Antti H, Jonsson P, Daykin C, Navarange M, Beger RD, Verheij ER, Amberg A, Baunsgaard D, Cantor GH, Lehman-McKeeman L, Earll M, Wold S, Johansson E, Haselden JN, Kramer K, Thomas C, Lindberg J, Schuppe-Koistinen I, Wilson ID, Reily Schaefer H, Spraul M (2005) Summary recommendations for standardization and reporting of metabolic analyses. Nat Biotechnol 23:833–838
Piotto M, Moussallieh FM, Dillmann B, Imperiale A, Neuville A, Brigand C, Bellocq JP, Elbayed K, Namer IJ (2008) Metabolic characterization of primary human colorectal cancers using high resolution magic angle spinning 1H magnetic resonance spectroscopy. Metabolomics 5:292–301
Psychogios N, Hau DD, Peng J, Guo AC, Mandal R, Bouatra S, Sinelnikov I, Krishnamurthy R, Eisner R, Gautam B, Young N, Xia J, Knox C, Dong E, Huang P, Hollander Z, Pedersen TL, Smith SR, Bamforth F, Greiner R, McManus B, Newman JW, Goodfriend T, Wishart DS (2011) The human serum metabolome. PLoS One 6(2):e16957
Sukumaran DK, Garcia E, Hua J, Tabaczynski W, Odunsi K, Andrews C, Szyperski T (2009) Standard operating procedure for metabonomic studies of blood serum and plasma samples using a 1H-NMR micro-flow probe. Magn Reson Chem 47:S81–S85
Teahan O, Gamble S, Holmes E, Waxman J, Nicholson JK, Bevan C, Keun HC (2006) Impact of analytical bias in metabonomic studies of human blood serum and plasma. Anal Chem 78:4307–4318
Tenenhaus M, Gauchi JP, Menardo C (1995) Regression PLS et applications. Revue Statistique Appliquée XLIII(1):7–63
Walsh MC, Brennan L, Malthouse JPG, Roche HM, Gibney MJ (2006) Effect of acute dietary standardization on the urinary, plasma, and salivary metabolomic profiles of healthy humans. Am J Clin Nutr 84:531–539
Wishart DS, Knox C, Guo AC et al. (2009) HMDB: a knowledgebase for the human metabolome. Nucleic Acids Res 37(Database issue):D603–D610
Acknowledgments
This work was supported by the Conseil Régional de Picardie. We are grateful to Brian de Witt and Karsten Hiller for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Fliniaux, O., Gaillard, G., Lion, A. et al. Influence of common preanalytical variations on the metabolic profile of serum samples in biobanks. J Biomol NMR 51, 457–465 (2011). https://doi.org/10.1007/s10858-011-9574-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-011-9574-5