Abstract
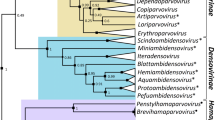
In determining putative recombination events and their evolution rates in the RNAs 1 and 2 of currently the known members of the family Bromoviridae, a detailed study comprising 107 accessions retrieved from the international databases, has been carried out by using RECCO and RDP v3.31β algorithms. These programs allowed the detection of potential recombination sites in all the five virus genera composing the family Bromoviridae with various degrees of consistency. The RNAs 1 and 2 showed inferred phylogenies fully congruent and clearly delineated five clusters representing the five studied virus genera. In this respect, we proposed to classify the Ilarviruses in three distinct subgroups instead of 10 as mentioned in several reports of the International Committee on Taxonomy of Viruses where its suggestions were based on antigenic differences. Moreover, we confirmed that Alfalfa mosaic virus should be considered as a component of the Ilarvirus genus instead of being the unique representative of Alfamovirus genus. In addition, Pelargonium zonate spot and Olive latent 2 viruses fully deserve their affiliation to the family Bromoviridae.

Similar content being viewed by others
References
R. Aaziz, M. Tepfer, J. Gen. Virol. 80, 1339–1346 (1999)
M. Alejska, A. Kurzyniska-Kokorniak, M. Broda, R. Kierzek, M. Figlerowicz, Acta Biochim. Pol. 48, 391–407 (2001)
M. Baroth, M. Orlich, H.J. Thiel, P. Becher, Virology 278, 456–466 (2000). doi:https://doi.org/10.1006/viro.2000.0644
M.F. Boni, D. Posada, M.W. Feldman, Genetics 176, 1035–1047 (2007). doi:https://doi.org/10.1534/genetics.106.068874
M. Boulila, Phytopathol. Mediterr. 46, 285–294 (2007)
M. Boulila, Plant Mol. Biol. Rep. (2008). doi: https://doi.org/10.1007/s11105-008-0071-2
F.M. Codoner, S.F. Elena, Arch. Virol. 151, 299–307 (2006). doi:https://doi.org/10.1007/s00705-005-0628-4
F.M. Codoner, S.F. Elena, J. Gen. Virol. 89, 1739–1747 (2008). doi:https://doi.org/10.1099/vir.0.2008/000166-0
E. Domingo, J. Holland, C. Biebricher, M. Eigen, in Molecular Basis of Virus Evolution, ed. by A.J. Gibbs, C.H. Calisher, F. Garcia-Arenal (Cambridge University Press, Cambridge, 1995), pp. 181–191
M. Eigen, Trends Microbiol. 4, 216–218 (1996). doi:https://doi.org/10.1016/0966-842X(96)20011-3
C.M. Fauquet, M.A. Mayo, J. Maniloff, U. Desselberger, L.A. Ball, Eighth Report of the International Committee on Taxonomy of Viruses (Elsevier/Academic Press, London, 2005)
F. Garcia-Arenal, B.A. McDonald, Phytopathology 93, 941–952 (2003). doi:https://doi.org/10.1094/PHYTO.2003.93.8.941
D. Gallitelli, M. Finetti-Sialer, G.P. Martelli, Arch. Virol. 150, 407–411 (2005). doi:https://doi.org/10.1007/s00705-004-0450-4
M.J. Gibbs, J.S. Armstrong, A.J. Gibbs, Bioinformatics 16, 573–582 (2000). doi:https://doi.org/10.1093/bioinformatics/16.7.573
A.E. Greene, R.F. Allison, Science 263, 1423–1425 (1994)
J.J. Holland, J.C. DeLaTorre, D.A. Steinhauer, in Genetic Diversity of RNA Viruses, ed. by J.J. Holland (Springer-Verlag, Berlin, 1992), pp. 1–20
E.M.J. Jaspars, L. Bos, in: CMI/AAB. Descriptions of Plant viruses, no 229 (1980)
E.V. Koonin, A.E. Gorbalenya, J. Mol. Evol. 28, 524–527 (1989). doi:https://doi.org/10.1007/BF02602932
S.L. Kosakovsky Pond, D. Posada, M.B. Gravenor, C.H. Woelk, S.D.W. Frost, Mol. Biol. Evol. 23(10), 1891–1901 (2006)
S. Kumar, M. Nei, J. Dudley, K. Tamura, Brief. Bioinform. 9(4), 299–306 (2008). doi:https://doi.org/10.1093/bib/bbn017
M.M.C. Lai, Microbiol. Rev. 56, 61–79 (1992)
J.P. Legg, J.M. Thresh, Virus Res. 71, 135–149 (2000). doi:https://doi.org/10.1016/S0168-1702(00)00194-5
M.A. Larkin, G. Blackshileds, N.P. Brown, R. Chenna, P.A. McGettigan, H. McWilliam, F. Valentin, I.M. Wallace, A. Wilm, R. Lopez, J.D. Thompson, T.J. Gibson, D.G. Higgins, Bioinformatics 23(21), 2947–2948 (2007). doi:https://doi.org/10.1093/bioinformatics/btm404
D. Martin, E. Rybicki, Bioinformatics 16, 562–563 (2000). doi:https://doi.org/10.1093/bioinformatics/16.6.562
D.P. Martin, D. Posada, K.A. Crandall, C. Williamson, AIDS Res. Hum. Retroviruses 21, 98–102 (2005). doi:https://doi.org/10.1089/aid.2005.21.98
D.P. Martin, C. Williamson, D. Posada, Bioinformatics 21, 260–262 (2005). doi:https://doi.org/10.1093/bioinformatics/bth490
J. Maydt, T. Lengauer, Bioinformatics 22(9), 1064–1071 (2006). doi:https://doi.org/10.1093/bioinformatics/btl057
F. Monci, S. Sanchez-Campos, J. Navas-Castillo, E. Moriones, Virology 303, 317–326 (2002). doi:https://doi.org/10.1006/viro.2002.1633
M. Nagai, Y. Sakoda, M. Mori, M. Hayashi, H. Kida, H. Akashi, J. Gen. Virol. 84(Pt 2), 447–452 (2003). doi:https://doi.org/10.1099/vir.0.18773-0
M. Padidam, S. Sawyer, C.M. Fauquet, Virology 265, 218–225 (1999). doi:https://doi.org/10.1006/viro.1999.0056
D. Posada, K. Crandall, Proc. Natl. Acad. Sci. USA 98, 13757–13762 (2001). doi:https://doi.org/10.1073/pnas.241370698
D. Posada, K.A. Crandall, J. Mol. Evol. 54, 396–402 (2002)
C. Rampitsch, K.C. Eastwell, Arch. Virol. 142, 1911–1918 (1997). doi:https://doi.org/10.1007/s007050050210
P.J. Schiel, P.H. Berger, J. Gen. Virol. 81, 273–278 (2000)
M. Schierup, J. Hein, Genetics 156, 879–891 (2000)
M. Schierup, J. Hein, Mol. Biol. Evol. 17, 1578–1579 (2000)
S.W. Scott, M.T. Zimmerman, X. Ge, Arch. Virol. 143, 1187–1198 (1998). doi:https://doi.org/10.1007/s007050050366
J.M. Smith, J. Mol. Evol. 34, 126–129 (1992)
I.E. Tzanetakis, R.R. Martin, Virus Res. 112, 32–37 (2005). doi:https://doi.org/10.1016/j.virusres.2005.02.010
G.F. Weiller, Mol. Biol. Evol. 15, 326–335 (1998)
M. Worobey, E.C. Holmes, J. Gen. Virol. 80, 2535–2543 (1999)
D. Zimmern, in RNA Genetics, ed. by J.J. Holland, E.R. Domingo, P. Ahlquist (CRC Press, Boca Raton, 1988), pp. 211–240
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Boulila, M. Recombination structure and genetic relatedness among members of the family Bromoviridae based on their RNAs 1 and 2 sequence analyses. Virus Genes 38, 435–444 (2009). https://doi.org/10.1007/s11262-009-0340-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-009-0340-7