Abstract
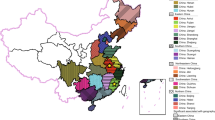
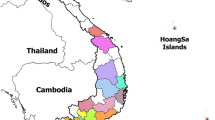
In several parts of China, there have been a large number of pseudorabies (PR) outbreaks which have devastated many swine farms even though the herds had been previously immunized with gE-deleted vaccines (Bartha-K61). The emergence of these outbreak-associated PRV strains might indicate that Bartha-K61 vaccine could not provide effective protection and poses challenges for current serologic diagnostics of anti-PRV antibodies. Here, we performed phylogenetic analyses based on partial gE, gB, and gC genes to provide information about the molecular epidemiology, diagnostics, and immune protection in these outbreak-associated PRV strains. Our results indicated that the maximal nucleotide sequence divergence for gE, gB, and gC genes are 1.7, 0.4, and 2.7 % within the cluster where outbreak-associated PRV strains were located, and are 2.3, 2.7, and 7.6 % with other clusters in the phylogenetic trees, respectively. Phylogenetic analyses revealed that gE, gB, and gC genes of the twelve outbreak-associated PRV strains clustered to a relatively independent branch of the tree, and evolved from the same ancestor with strains Ea-China-1999, Fa-China-2001, and BJ-China-2008. The genetic relationship between these outbreak-associated PRV strains and strain Bartha is not close which may genetically explain the emergence of PR outbreaks in Bartha-K61-vaccinated swine farms. We suggest that these outbreak-associated PRV strains originate from earlier strains in local regions in China.



Similar content being viewed by others
References
T.Q. An, J.M. Peng, Z.J. Tian, H.Y. Zhao, N. Li, Y.M. Liu, J.Z. Chen, C.L. Leng, Y. Sun, D. Chang, G.Z. Tong, Emerg. Infect. Dis. J. 19, 1749–1755 (2013)
Z. Gu, J. Dong, J. Wang, C. Hou, H. Sun, W. Yang, J. Bai, P. Jiang, Virus Res. 195, 57–63 (2015)
A. Marcaccini, M. Lopez Pena, M.I. Quiroga, R. Bermudez, J.M. Nieto, N. Aleman, Vet. Immunol. Immunopathol. 124, 264–273 (2008)
B.G. Klupp, C.J. Hengartner, T.C. Mettenleiter, L.W. Enquist, J. Virol. 78, 424–440 (2004)
L. Jacobs, T.G. Kimman, J. Gen. Virol. 74, 2201–2206 (1993)
L. Jacobs, R.H. Meloen, A.L.J. Gielkens, J.T.V. Oirschot, J. Gen. Virol. 71, 881–887 (1990)
B.T. Ober, A. Summerfield, C. Mattlinger, K.-H. Wiesmüller, G. Jung, E. Pfaff, A. Saalmüller, H.-J. Rziha, J. Virol. 72(6), 4866–4873 (1998)
B.T. Ober, B. Teufel, K.-H. Wiesmüller, G. Jung, E. Pfaff, A.S.H.-J. Rziha, J. Virol. 74(4), 1752–1760 (2000)
T. Ben-Porat, J.M. Demarchi, B. Lomnicai, A.S. Kaplan, Virology 154(2), 325–334 (1986)
A.K. Robbins, D.J. Dorney, M.W. Wathen, M.E. Whealy, C. Gold, R.J. Watson, L.E. Holland, S.D. Weed, M. Levine, J.C. Glorioso, L.W. Enquist, J. Virol. 61(9), 2691–2701 (1987)
C.A. Rue, P. Ryan, Virology 307, 12–21 (2003)
C.A. Rue, P. Ryan, J. Virol. Methods 151, 101–106 (2008)
T.L. Goldberg, R.M. Weigel, E.C. Hahn, G. Scherba, Epidemiol. Infect. 126, 415–424 (2001)
T.A. Hall, Nucleic Acids Sym. Ser. 41, 95–98 (1999)
K. Tamura, J. Dudley, M. Nei, S. Kumar, Mol. Biol. Evol. 24, 1596–1599 (2007)
R. Wu, C. Bai, J. Sun, S. Chang, X. Zhang, J. Vet. Sci. 14, 363 (2013)
X. Yu, Z. Zhou, D. Hu, Q. Zhang, T. Han, X. Li, X. Gu, L. Yuan, S. Zhang, B. Wang, P. Qu, J. Liu, X. Zhai, K. Tian, Emerg. Infect. Dis. J. 20, 102–104 (2014)
Y. Luo, N. Li, X. Cong, C.-H. Wang, M. Du, L. Li, B. Zhao, J. Yuan, D.-D. Liu, S. Li, Y. Li, Y. Sun, H.-J. Qiu, Vet. Microbiol. 174, 107–115 (2014)
P. Jinmei, A. Tongqing, Z. Hongyuan, L. Yimin, C. Jiacheng, L. Chaoliang, S. Yan, C. Dan, T. Zhijun, T. Guangzhi, Chin. J. Prev. Vet. Med. 35, 1–4 (2013)
R. Zeng, Q.Y.D. Torrents, C. Martinez, The University of Minnesota, Leman China Swine Conference Proceedings, 298, (2014)
P.R.C. Ministry of Agriculture, (2012)
Acknowledgments
This work was supported by grants from China Agriculture Research System (No. CARS-36), the Pig Industry Technology System Innovation Team Project of Henan Province (S2012-06) and the National Natural Science Foundation of China (No. 31172348). The authors are grateful to the invaluable comments from Renfeng Li, from Henan Institute of Science and Technology, Xinxiang, P.R. China, and would like to thank Dr Gregson, University College London, for revising the paper.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Edited by William Dundon.
Rights and permissions
About this article
Cite this article
Wang, Y., Qiao, S., Li, X. et al. Molecular epidemiology of outbreak-associated pseudorabies virus (PRV) strains in central China. Virus Genes 50, 401–409 (2015). https://doi.org/10.1007/s11262-015-1190-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-015-1190-0