Abstract
Purpose
This study investigates an efficient (nearly real-time) two-stage spine labeling algorithm that removes the need for an external training while being applicable to different types of MRI data and acquisition protocols.
Methods
Based solely on the image being labeled (i.e., we do not use training data), the first stage aims at detecting potential vertebra candidates following the optimization of a functional containing two terms: (i) a distribution-matching term that encodes contextual information about the vertebrae via a density model learned from a very simple user input, which amounts to a point (mouse click) on a predefined vertebra; and (ii) a regularization constraint, which penalizes isolated candidates in the solution. The second stage removes false positives and identifies all vertebrae and discs by optimizing a geometric constraint, which embeds generic anatomical information on the interconnections between neighboring structures. Based on generic knowledge, our geometric constraint does not require external training.
Results
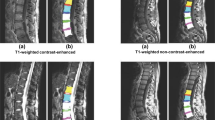
We performed quantitative evaluations of the algorithm over a data set of 90 mid-sagittal MRI images of the lumbar spine acquired from 45 different subjects. To assess the flexibility of the algorithm, we used both T1- and T2-weighted images for each subject. A total of 990 structures were automatically detected/labeled and compared to ground-truth annotations by an expert. On the T2-weighted data, we obtained an accuracy of 91.6% for the vertebrae and 89.2% for the discs. On the T1-weighted data, we obtained an accuracy of 90.7% for the vertebrae and 88.1% for the discs.
Conclusion
Our algorithm removes the need for external training while being applicable to different types of MRI data and acquisition protocols. Based on the current testing data, a subject-specific model density and generic anatomical information, our method can achieve competitive performances when applied to T1- and T2-weighted MRI images.







Similar content being viewed by others
Notes
Typically, spine MRI studies contain more than 100 images.
Our meta-analysis of accuracy is not a direct comparison between the methods because the used data sets are different, and might have various levels of difficulties. For instance, the CT data set in [29] contains several unusual appearances (e.g., abnormal spine curvature), and undergo large variations in the field of view, image noise and resolutions.
References
Fardon DF, Milette PC (2001) Nomenclature and classification of lumbar disc pathology: recommendations of the combined task forces of the North American Spine Society, American Society of Spine Radiology, and American Society of Neuroradiology. Spine 26(5):E93–E113
Wimmer M, Major D, Novikov AA, Buhler K (2016) Local entropy-optimized texture models for semi-automatic spine labeling in various MRI protocols. In: International symposium on biomedical imaging (ISBI), pp 155–159
De Leener B, Cohen-Adad J, Kadoury S (2015) Automatic segmentation of the spinal cord and spinal canal coupled with vertebral labeling. IEEE Trans Med Imaging 34(8):1705–1718
Ullmann E, Paquette JFP, Thong WE, Cohen-Adad J (2014) Automatic labeling of vertebral levels using a robust template-based approach. Int J Biomed Imaging 719520(1–719520):9
Cai Y, Landis M, Laidley DT, Kornecki A, Lum A, Li S (2016) Multi-modal vertebrae recognition using transformed deep convolution network. Comput Med Imaging Graphics 51:11–19
Kelm BM, Wels M, Zhou SK, Seifert S, Sühling M, Zheng Y, Comaniciu D (2013) Spine detection in CT and MR using iterated marginal space learning. Med Image Anal 17(8):1283–1292
Glocker B, Feulner J, Criminisi A, Haynor DR, Konukoglu E (2012) Automatic localization and identification of vertebrae in arbitrary field-of-view CT scans. Med Image Comput Comput Assist Interv MICCAI 3:590–598
Zhan Y, Dewan M, Harder M, Zhou X S (2012) Robust MR spine detection using hierarchical learning and local articulated model. Med Image Comput Comput Assist Interv MICCAI 1:141–148
Roberts MG, Cootes TF, Adams JE (2012) Automatic location of vertebrae on DXA images using random forest regression. Med Image Comput Comput Assist Interv MICCAI 3:361–368
Alomari RS, Corso JJ, Chaudhary V (2011) Labeling of lumbar discs using both pixel- and object-level features with a two-level probabilistic model. IEEE Trans Med Imaging 30(1):1–10
Oktay AB, Akgul YS (2011) Localization of the lumbar discs using machine learning and exact probabilistic inference. Med Image Comput Comput Assist Interv MICCAI 3:158–165
Ma J, Lu L, Zhan Y, Zhou XS, Salganicoff M, Krishnan A (2010) Hierarchical segmentation and identification of thoracic vertebra using learning-based edge detection and coarse-to-fine deformable model. Med Image Comput Comput Assist Interv MICCAI 1:19–27
Klinder T, Ostermann J, Ehm M, Franz A, Kneser R, Lorenz C (2009) Automated model-based vertebra detection, identification, and segmentation in ct images. Med Image Anal 13(3):471–482
Huang S, Chu Y, Lai S, Novak C (2009) Learning-based vertebra detection and iterative normalized-cut segmentation for spinal MRI. IEEE Trans Med Imaging 28(10):1595–1605
Miles B, Ayed I, Law MWK, Garvin GJ, Fenster A, Li S (2013) Spine image fusion via graph cuts. IEEE Trans Biomed Eng 60(7):1841–1850
Wang Z, Zhen X, Tay K, Osman S, Romano W, Li S (2015) Regression segmentation for \(\text{ m }^{3}\) spinal images. IEEE Trans Med Imaging 34(8):1640–1648
Schmidt S, Kappes JH, Bergtholdt M, Pekar V, Dries SPM, Bystrov D, Schnörr C (2007) Spine detection and labeling using a parts-based graphical model. In: Information processing in medical imaging (IPMI), pp 122–133
Nikolova M, Esedoglu S, Chan TF (2006) Algorithms for finding global minimizers of image segmentation and denoising models. SIAM J Appl Math 66(5):1632–1648
Bertsekas DP (1999) Nonlinear programming. Athena Scientific, Belmont
Yuan J, Bae E, Tai X-C, Boykov Y (2010) A study on continuous max-flow and min-cut approaches. Part I: binary labeling, technical report CAM-10-61, UCLA
Yuan J, Bae E, Tai X-C (2010) A study on continuous max-flow and min-cut approaches. In: IEEE conference on computer vision and pattern recognition (CVPR)
Tang M, Ben Ayed I, Boykov Y (2014) Pseudo-bound optimization for binary energies. Eur Conf Comput Vis ECCV 5:691–707
Taniai T, Matsushita Y, Naemura T (2015) Superdifferential cuts for binary energies. In: IEEE conference on computer vision and pattern recognition (CVPR), pp 2030–2038
Ben Ayed I, Punithakumar K, Li S (2015) Distribution matching with the bhattacharyya similarity: a bound optimization framework. IEEE Trans Pattern Anal Mach Intell 37(9):1777–1791
Punithakumar K, Yuan J, Ayed IB, Li S, Boykov Y (2012) A convex max-flow approach to distribution-based figure-ground separation. SIAM J Imaging Sci 5(4):1333–1354
Salah MB, Ben Ayed I, Yuan J, Zhang H (2014) Convex-relaxed kernel mapping for image segmentation. IEEE Trans Image Process 23(3):1143–1153
Chambolle A (2004) An algorithm for total variation minimization and applications. J Math Imaging Vis 20(1–2):89–97
Lootus M, Kadir T, Zisserman A (2013) Computational methods and clinical applications for spine imaging. In: Jianhua Y, Tobias K, Shuo L (eds) Vertebrae detection and labelling in lumbar MR images. Springer, Cham, pp 219–230
Yang D, Xiong T, Xu D, Huang Q, Liu D, Zhou SK, Xu Z, Park J, Chen M, Tran TD, Chin SP, Metaxas D, Comaniciu D (2017) Automatic vertebra labeling in large-scale 3D CT using deep image-to-image network with message passing and sparsity regularization. In: Information processing in medical imaging (IPMI), pp 633–644
Glocker B, Zikic D, Konukoglu E, Haynor DR, Criminisi A (2013) Vertebrae localization in pathological spine CT via dense classification from sparse annotations. Med Image Comput Comput Assist Interv MICCAI 2:262–270
Chen H, Shen C, Qin J, Ni D, Shi L, Cheng JCY, Heng P-A (2015) Automatic localization and identification of vertebrae in spine CT via a joint learning model with deep neural networks. Med Image Comput Comput Assist Interv MICCAI 1:515–522
Acknowledgements
This work was supported in part by the Natural Sciences and Engineering Research Council of Canada (NSERC).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declare that they have no competing interests.
Ethical approval
The study was approved by the University of Western Ontario Research Ethics Board.
Rights and permissions
About this article
Cite this article
Hojjat, SP., Ayed, I., Garvin, G.J. et al. Spine labeling in MRI via regularized distribution matching. Int J CARS 12, 1911–1922 (2017). https://doi.org/10.1007/s11548-017-1651-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11548-017-1651-0