Abstract
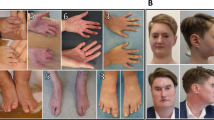
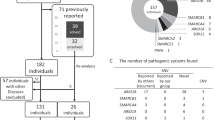
ABL1 is a proto-oncogene well known as part of the fusion gene BCR-ABL1 in the Philadelphia chromosome of leukemia cancer cells1. Inherited germline ABL1 changes have not been associated with genetic disorders. Here we report ABL1 germline variants cosegregating with an autosomal dominant disorder characterized by congenital heart disease, skeletal abnormalities, and failure to thrive. The variant c.734A>G (p.Tyr245Cys) was found to occur de novo or cosegregate with disease in five individuals (families 1–3). Additionally, a de novo c.1066G>A (p.Ala356Thr) variant was identified in a sixth individual (family 4). We overexpressed the mutant constructs in HEK 293T cells and observed increased tyrosine phosphorylation, suggesting increased ABL1 kinase activities associated with both the p.Tyr245Cys and p.Ala356Thr substitutions. Our clinical and experimental findings, together with previously reported teratogenic effects of selective BCR-ABL inhibitors in humans2,3,4,5 and developmental defects in Abl1 knockout mice6,7, suggest that ABL1 has an important role during organismal development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
de Klein, A. et al. A cellular oncogene is translocated to the Philadelphia chromosome in chronic myelocytic leukaemia. Nature 300, 765–767 (1982).
Pye, S.M. et al. The effects of imatinib on pregnancy outcome. Blood 111, 5505–5508 (2008).
Apperley, J. Issues of imatinib and pregnancy outcome. J. Natl. Compr. Canc. Netw. 7, 1050–1058 (2009).
Jain, N., Sharma, D., Agrawal, R. & Jain, A. A newborn with teratogenic effect of imatinib mesylate: a very rare case report. Med. Princ. Pract. 24, 291–293 (2015).
Ault, P. et al. Pregnancy among patients with chronic myeloid leukemia treated with imatinib. J. Clin. Oncol. 24, 1204–1208 (2006).
Li, B. et al. Mice deficient in Abl are osteoporotic and have defects in osteoblast maturation. Nat. Genet. 24, 304–308 (2000).
Qiu, Z., Cang, Y. & Goff, S.P. c-Abl tyrosine kinase regulates cardiac growth and development. Proc. Natl. Acad. Sci. USA 107, 1136–1141 (2010).
Tartaglia, M. et al. Mutations in PTPN11, encoding the protein tyrosine phosphatase SHP-2, cause Noonan syndrome. Nat. Genet. 29, 465–468 (2001).
Yang, W. et al. Ptpn11 deletion in a novel progenitor causes metachondromatosis by inducing hedgehog signalling. Nature 499, 491–495 (2013).
Hafner, C., Toll, A. & Real, F.X. HRAS mutation mosaicism causing urothelial cancer and epidermal nevus. N. Engl. J. Med. 365, 1940–1942 (2011).
Aoki, Y. et al. Germline mutations in HRAS proto-oncogene cause Costello syndrome. Nat. Genet. 37, 1038–1040 (2005).
Lahiry, P., Torkamani, A., Schork, N.J. & Hegele, R.A. Kinase mutations in human disease: interpreting genotype–phenotype relationships. Nat. Rev. Genet. 11, 60–74 (2010).
Yang, Y. et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N. Engl. J. Med. 369, 1502–1511 (2013).
Yang, Y. et al. Molecular findings among patients referred for clinical whole-exome sequencing. J. Am. Med. Assoc. 312, 1870–1879 (2014).
Brasher, B.B. & Van Etten, R.A. c-Abl has high intrinsic tyrosine kinase activity that is stimulated by mutation of the Src homology 3 domain and by autophosphorylation at two distinct regulatory tyrosines. J. Biol. Chem. 275, 35631–35637 (2000).
Hantschel, O. et al. A myristoyl/phosphotyrosine switch regulates c-Abl. Cell 112, 845–857 (2003).
Hardin, J.D., Boast, S., Mendelsohn, M., de los Santos, K. & Goff, S.P. Transgenes encoding both type I and type IV c-abl proteins rescue the lethality of c-abl mutant mice. Oncogene 12, 2669–2677 (1996).
Nagar, B. et al. Structural basis for the autoinhibition of c-Abl tyrosine kinase. Cell 112, 859–871 (2003).
Guex, N. & Peitsch, M.C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18, 2714–2723 (1997).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Schwarz, J.M., Cooper, D.N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat. Methods 11, 361–362 (2014).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Shen, Y. et al. A SNP discovery method to assess variant allele probability from next-generation resequencing data. Genome Res. 20, 273–280 (2010).
Acknowledgements
We thank the families for their participation and collaboration. This work was funded in part by US National Human Genome Research Institute (NHGRI)/National Heart, Lung, and Blood Institute (NHLBI) grant U54HG006542 to the Baylor-Hopkins Center for Mendelian Genomics (BH-CMG) and the NIH Common Fund, through the Office of Strategic Coordination/Office of the NIH Director under award U01HG007709. W.-L.C. was supported by Cancer Prevention Research Institute of Texas (CPRIT) training Program RP140102. The work of N.A.M. is supported by the Rashid Family Fund.
Author information
Authors and Affiliations
Contributions
X.W., Y.Y., W.-L.C., C.P.S., C.-A.C., and J.R.L. designed the studies. X.W., Y.Y., C.P.S., W.-L.C., J.R.L., S.E.P., J.A.R., and F.X. participated in the writing of the manuscript. X.W., C.Q., Y.D., D.M.M., R.A.G., C.M.E., M.W., F.X., and Y.Y. performed sequencing data analysis. W.-L.C. and C.-A.C. performed the kinase activity assays. X.W., J.A.R., A.A.S., L.A.-G., M.M., N.A.M., G.A., C.P.S., and Y.Y. recruited patients and gathered detailed clinical information for the study. Y.Y., C.P.S., and J.R.L. supervised the studies.
Corresponding authors
Ethics declarations
Competing interests
J.R.L. is a paid consultant for Regeneron Pharmaceuticals, holds stock ownership in 23andMe and Lasergen, Inc., is on the scientific advisory board of Baylor Genetics, and is a co-inventor on US and European patents related to molecular diagnostics. S.E.P. is on the scientific advisory board of Baylor Genetics. The Department of Molecular and Human Genetics at Baylor College of Medicine derives revenue from molecular genetic testing offered at Baylor Genetics.
Integrated supplementary information
Supplementary Figure 1 The growth chart of the probands.
(a) Subject 1, height curve. (b) Subject 1, weight curve. (c) Subject 3, height curve. (d) Subject 3, weight curve. (e) Subject 3, head circumference curve. (f) Subject 4, height curve. (g) Subject 4, weight curve. (h) Subject 4, head circumference curve. (i) Subject 6, height curve. (j) Subject 6, weight curve. (k) Subject 6, head circumference curve. A chart for either 2–20 years old or 0–3 years old is shown for each patient. 5th, 25th, 50th, 75th, and 100th percentile curves are provided as references (http://www.cdc.gov/growthcharts). Figures were generated using an Excel sheet developed by Ernest M. Post, MD, and Zimi Medical Technologies, LLC.
Supplementary Figure 2 Sanger sequencing of the two variants in affected families.
The c.734A>G (p.Tyr245Cys) variant was identified in families 1–3, and the c.1066G>A (p.Ala356Thr) variant was identified in family 4.
Supplementary Figure 4 Full-length gels and blots with molecular weight standards showing the effect of the ABL1 variants (isoform 1b) on phosphorylation.
The multiple non-specific bands seen in full blots of p-ABL1 and p-STAT5 from cells expressing ABL1-1bTyr245Cys and ABL1-1bAla356Thr (but not in wild-type cells) may represent non-specific proteins resulting from the increased activity of both ABL1-1bTyr245Cys and ABL1-1bAla356Thr.
Supplementary Figure 5 The effect of ABL1 variants (isoform 1a) on phosphorylation.
(a) Overall phosphotyrosine levels and phosphorylation of specific ABL1 substrates were analyzed by transiently expressing the wild-type and mutant constructs in HEK 293T cells and immunoblotting. Cells expressing both variants showed increased overall phosphotyrosine levels and phosphorylation of STAT5 when compared with wild-type cells. Increased levels of phosphorylated ABL1 were observed for Ala337Thr but not Tyr226Cys owing to the substitution of the Tyr226 residue, which is recognized by the anti-phospho-ABL1 antibody. No significant difference in the phosphorylation levels of CrKL, SMAD2, and SMAD3 was observed between cells expressing the mutants and wild-type protein. Antibodies used in detection included anti-phosphotyrosine (p-Tyr; 4G10) antibody for the overall phosphotyrosine level and anti-phospho-ABL1 (p-ABL1), anti-phospho-STAT5 (p-STAT5), anti-phospho-CrkL (p-CrkL), and anti-phospho-SMAD2 and SMAD3 (p-Smad2/3) antibodies to determine the phosphorylation levels of specific ABL1 substrates in whole-cell lysates. The level of GAPDH is used as an internal loading control. Experiments for each construct were performed in triplicate. (b) Quantification of the immunoblot results. Data are normalized to GAPDH protein levels, with the wild-type protein set at 1.0. **P ≤ 0.01, ****P ≤ 0.0001; n.s., P > 0.05.
Supplementary Figure 6 The effect of the ABL1 Tyr245Phe variant (isoform 1b) on phosphorylation.
Overall phosphotyrosine levels and phosphorylation of specific ABL1 substrates were analyzed by transiently expressing the wild-type and mutant constructs in HEK 293T cells and immunoblotting. Cells expressing the Tyr245Phe variant showed decreased overall phosphotyrosine levels when compared with cells expressing the wild-type protein. Phosphorylated ABL1 was not observed for Tyr245Cys and Tyr245Phe owing to the substitution of the Tyr245 residue, which is recognized by the anti-phospho-ABL1 antibody. Antibodies used in detection included anti-phosphotyrosine (p-Tyr; 4G10) antibody for the overall phosphotyrosine level and anti-phospho-ABL1 (p-ABL1), anti-phospho-STAT5 (p-STAT5), and anti-phospho-CrkL (p-CrkL) antibodies to determine the phosphorylation levels of specific ABL1 substrates in whole-cell lysates. The level of GAPDH is used as an internal loading control. Longer exposure of the blot is also included to show the overall phosphotyrosine patterns more clearly.
Supplementary Figure 7 The effect of the ABL1 Tyr226Phe variant (isoform 1a) on phosphorylation.
Overall phosphotyrosine levels and phosphorylation of specific ABL1 substrates were analyzed by transiently expressing the wild-type and mutant constructs in HEK 293T cells and immunoblotting. Cells expressing the Tyr226Phe variant showed decreased overall phosphotyrosine levels when compared with cells expressing the wild-type protein. Phosphorylated ABL1 was not observed for Tyr226Cys and Tyr226Phe owing to the substitution of the Tyr226 residue, which is recognized by the anti-phospho-ABL1 antibody. Antibodies used in detection included anti-phosphotyrosine (p-Tyr; 4G10) antibody for the overall phosphotyrosine level and anti-phospho-ABL1 (p-ABL1), anti-phospho-STAT5 (p-STAT5), and anti-phospho-CrkL (p-CrkL) antibodies to determine the phosphorylation levels of specific ABL1 substrates in whole-cell lysates. The level of GAPDH is used as an internal loading control. Longer exposure of the blot is also included to show the overall phosphotyrosine patterns more clearly.
Supplementary Figure 8 Semiquantitative RT–PCR/Sanger sequencing analyses of the c.734A>G (p.Tyr245Cys) mutation in subjects 1 and 2.
Affected subjects 1 (F1:II1) and 2 (F1:III1) have similar levels of mutant and wild-type ABL1 transcripts.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–5 and Supplementary Note (PDF 2615 kb)
Supplementary Data 1
The exact numbers used for statistical calculation. (XLSX 18 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Charng, WL., Chen, CA. et al. Germline mutations in ABL1 cause an autosomal dominant syndrome characterized by congenital heart defects and skeletal malformations. Nat Genet 49, 613–617 (2017). https://doi.org/10.1038/ng.3815
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3815
This article is cited by
-
Human ABL1 deficiency syndrome (HADS) is a recognizable syndrome distinct from ABL1-related congenital heart defects and skeletal malformations syndrome
Human Genetics (2024)
-
Pathogenic variants causing ABL1 malformation syndrome cluster in a myristoyl-binding pocket and increase tyrosine kinase activity
European Journal of Human Genetics (2021)
-
Long non-coding RNA SAP30-2:1 is downregulated in congenital heart disease and regulates cell proliferation by targeting HAND2
Frontiers of Medicine (2021)
-
ISL1 loss-of-function mutation contributes to congenital heart defects
Heart and Vessels (2019)
-
A Novel MEF2C Loss-of-Function Mutation Associated with Congenital Double Outlet Right Ventricle
Pediatric Cardiology (2018)