Abstract
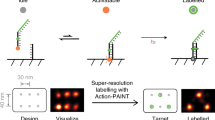
Recent advances in fluorescence super-resolution microscopy have allowed subcellular features and synthetic nanostructures down to 10–20 nm in size to be imaged. However, the direct optical observation of individual molecular targets (∼5 nm) in a densely packed biomolecular cluster remains a challenge. Here, we show that such discrete molecular imaging is possible using DNA-PAINT (points accumulation for imaging in nanoscale topography)—a super-resolution fluorescence microscopy technique that exploits programmable transient oligonucleotide hybridization—on synthetic DNA nanostructures. We examined the effects of a high photon count, high blinking statistics and an appropriate blinking duty cycle on imaging quality, and developed a software-based drift correction method that achieves <1 nm residual drift (root mean squared) over hours. This allowed us to image a densely packed triangular lattice pattern with ∼5 nm point-to-point distance and to analyse the DNA origami structural offset with ångström-level precision (2 Å) from single-molecule studies. By combining the approach with multiplexed exchange-PAINT imaging, we further demonstrated an optical nanodisplay with 5 × 5 nm pixel size and three distinct colours with <1 nm cross-channel registration accuracy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Klar, T. A. & Hell, S. W. Subdiffraction resolution in far-field fluorescence microscopy. Opt. Lett. 24, 954–956 (1999).
Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Gustafsson, M. G. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl Acad. Sci. USA 102, 13081–13086 (2005).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nature Methods 3, 793–795 (2006).
Heilemann, M. et al. Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew. Chem. Int. Ed. 47, 6172–6176 (2008).
Sharonov, A. & Hochstrasser, R. M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl Acad. Sci. USA 103, 18911–18916 (2006).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Hell, S. W. Microscopy and its focal switch. Nat. Methods 6, 24–32 (2009).
Huang, B., Bates, M. & Zhuang, X. Super-resolution fluorescence microscopy. Ann. Rev. Biochem. 78, 993–1016 (2009).
Sahl, S. J. & Moerner, W. E. Super-resolution fluorescence imaging with single molecules. Curr. Opin. Struct. Biol. 23, 778–787 (2013).
Moerner, W. E. & Kador, L. Optical detection and spectroscopy of single molecules in a solid. Phys. Rev. Lett. 62, 2535–2538 (1989).
Russell, E. T., Daniel, R. L. & Watt, W. W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Löschberger, A. et al. Super-resolution imaging visualizes the eightfold symmetry of gp210 proteins around the nuclear pore complex and resolves the central channel with nanometer resolution. J. Cell Sci. 125, 570–575 (2012).
Szymborska, A. et al. Nuclear pore scaffold structure analyzed by super-resolution microscopy and particle averaging. Science 341, 655–658 (2013).
Yildiz, A. et al. Myosin V walks hand-over-hand: single fluorophore imaging with 1.5-nm localization. Science 300, 2061–2065 (2003).
Rittweger, E., Han, K., Irvine, S. E., Eggeling, C. & Hell, S. W. STED microscopy reveals crystal colour centres with nanometric resolution. Nature Photon. 3, 144–147 (2009).
Shtengel, G. et al. Interferometric fluorescent super-resolution microscopy resolves 3D cellular ultrastructure. Proc. Natl Acad. Sci. USA 106, 3125–3130 (2009).
Pertsinidis, A., Zhang, Y. & Chu, S. Subnanometre single-molecule localization, registration and distance measurements. Nature 466, 647–651 (2010).
Vaughan, J. C., Jia, S. & Zhuang, X. Ultrabright photoactivatable fluorophores created by reductive caging. Nat. Methods 9, 1181–1184 (2012).
Raab, M., Schmied, J. J., Jusuk, I., Forthmann, C. & Tinnefeld, P. Fluorescence microscopy with 6 nm resolution on DNA origami. Chemphyschem 15, 2431–2435 (2014).
Kaplan, C. et al. Absolute arrangement of subunits in cytoskeletal septin filaments in cells measured by fluorescence microscopy. Nano Lett. 15, 3859–3864 (2015).
Fernández-Suárez, M., Chen, T. S. & Ting, A. Y. Protein-protein interaction detection in vitro and in cells by proximity biotinylation. J. Am. Chem. Soc. 130, 9251–9253 (2008).
Shroff, H., Galbraith, C. G., Galbraith, J. A. & Betzig, E. Live-cell photoactivated localization microscopy of nanoscale adhesion dynamics. Nat. Methods 5, 417–423 (2008).
Dempsey, G. T., Vaughan, J. C., Chen, K. H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Xu, K., Babcock, H. P. & Zhuang, X. Dual-objective STORM reveals three-dimensional filament organization in the actin cytoskeleton. Nat. Methods 9, 185–188 (2012).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Schoen, I., Ries, J., Klotzsch, E., Ewers, H. & Vogel, V. Binding-activated localization microscopy of DNA structures. Nano Lett. 11, 4008–4011 (2011).
Jungmann, R. et al. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 10, 4756–4761 (2010).
Johnson-Buck, A. et al. Super-resolution fingerprinting detects chemical reactions and idiosyncrasies of single DNA pegboards. Nano Lett. 13, 728–733 (2013).
Iinuma, R. et al. Polyhedra self-assembled from DNA tripods and characterized with 3D DNA-PAINT. Science 344, 65–69 (2014).
Nieuwenhuizen, R. P. et al. Measuring image resolution in optical nanoscopy. Nature Methods 10, 557–562 (2013).
Cordes, T. et al. Resolving single-molecule assembled patterns with superresolution blink-microscopy. Nano Lett. 10, 645–651 (2010).
Thompson, M. A., Lew, M. D. & Moerner, W. E. Extending microscopic resolution with single-molecule imaging and active control. Annu. Rev. Biophys. 41, 321–342 (2012).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Schmied, J. et al. Fluorescence and super-resolution standards based on DNA origami. Nature Methods 9, 1133–1134 (2012).
Alushin, G. M. et al. High-resolution microtubule structures reveal the structural transitions in αβ-Tubulin upon GTP hydrolysis. Cell 157, 1117–1129 (2014).
Pertsinidis, A. et al. Ultrahigh-resolution imaging reveals formation of neuronal SNARE/Munc18 complexes in situ. Proc. Natl Acad. Sci. USA 110, E2812–E2820 (2013).
Chmyrov, A. et al. Nanoscopy with more than 100,000 ‘doughnuts’. Nat. Methods 10, 737–740 (2013).
Xu, K., Zhong, G. & Zhuang, X. Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons. Science 339, 452–456 (2013).
Dong, L. et al. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science 349, aab3500 (2015).
Smith, C. S., Joseph, N., Rieger, B. & Lidke, K. A. Fast, single-molecule localization that achieves theoretically minimum uncertainty. Nature Methods 7, 373–375 (2010).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucl. Acids Res. 37, 5001–5006 (2009).
Acknowledgements
We thank M. Avendaño, J. Wöhrstein, F. Schüder, G. Church, W. Shih, D. Zhang, S. Agasti, E. Winfree and P. Rothemund for helpful discussions, and C. Myhrvold, N. Liu, W. Sun, X. Chen, F. Vigneault and H. Tang for helpful comments on the manuscript. This work is supported by a National Institutes of Health (NIH) Director's New Innovator Award (1DP2OD007292), an NIH Transformative Research Award (1R01EB018659), an NIH grant (5R21HD072481), an Office of Naval Research (ONR) Young Investigator Program Award (N000141110914), ONR grants (N000141010827 and N000141310593), a National Science Foundation (NSF) Faculty Early Career Development Award (CCF1054898), an NSF grant (CCF1162459) and a Wyss Institute for Biologically Engineering Faculty Startup Fund to P.Y. M.D. acknowledges support from the HHMI International Predoctoral Fellowship. R.J. acknowledges support from the Alexander von Humboldt-Foundation through a Feodor-Lynen Fellowship.
Author information
Authors and Affiliations
Contributions
M.D. conceived of and designed the study, designed and performed the experiments, developed the software, analysed the data and wrote the manuscript. R.J. conceived of and designed the study, interpreted the data and critiqued the manuscript. P.Y. conceived of, designed and supervised the study, interpreted the data and wrote the manuscript. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have filed a patent application. P.Y. and R.J. are co-founders of Ultivue, Inc., a start-up company with interests in commercializing the reported technology.
Supplementary information
Supplementary information
Supplementary information (PDF 9594 kb)
Supplementary information
Supplementary Tables (PDF 393 kb)
Rights and permissions
About this article
Cite this article
Dai, M., Jungmann, R. & Yin, P. Optical imaging of individual biomolecules in densely packed clusters. Nature Nanotech 11, 798–807 (2016). https://doi.org/10.1038/nnano.2016.95
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2016.95
This article is cited by
-
Advances in the multimodal analysis of the 3D chromatin structure and gene regulation
Experimental & Molecular Medicine (2024)
-
Ångström-resolution fluorescence microscopy
Nature (2023)
-
Biotemplated precise assembly approach toward ultra-scaled high-performance electronics
Nature Protocols (2023)
-
Multi-micron crisscross structures grown from DNA-origami slats
Nature Nanotechnology (2023)
-
Design and Simulation of an Autonomous Molecular Mechanism Using Spatially Localized DNA Computation
Interdisciplinary Sciences: Computational Life Sciences (2023)