Abstract
High numbers of melanocytic naevi (moles), and mutations in the p16 gene (CDKN2A), are two strong risk factors for cutaneous malignant melanoma. We have previously reported linkage of mole count to the CDKN2A locus. Here, we report genome-wide scans for mole counts (differentiated into flat, raised and atypical subtypes) with a total of 796 microsatellite markers for 424 families with 1024 twins and siblings, plus genotypes for 690 parents. Inclusion of 221 pairs of MZ twins enabled separation of shared environmental and polygenic influences, so placing an upper limit to estimates of QTL variance. Maximum likelihood multipoint variance component methods were used to assess linkage of naevus count. Sex, age, body surface area, skin colour, hair colour, sunburn and facial freckles were included as covariates. Peak linkage of flat mole count was to regions on chromosomes 2, 9, 8 and 17 with lod scores 2.95, 2.95, 2.50 and 2.15, respectively. The support for linkage to the CDKN2A gene region (9p21) increased to 3.42 when additional fine mapping markers were added. For raised mole count, there was suggestive evidence of linkage in our sample to chromosome 16 (lod=1.87), and for atypical mole count on chromosomes 1, 6 and X with lod scores of 2.20, 2.00 and 2.00, respectively. The multivariate linkage peaks generally match those from individual trait analyses, with the exception of a new peak on chromosome 4 (point-wise empirical P-value=0.001). We replicate our earlier finding of linkage to CDKN2A and discovering linkage to several novel regions that may also influence risk of the development of malignant melanoma.
Similar content being viewed by others
Introduction
Cutaneous malignant melanoma (CMM; MIM 15560) is common in the Australian state of Queensland, where a fair skinned population lives in tropical and subtropical latitude.1 Although fewer than 1% of cases in this population are strongly familial, the overall heritability of CMM is approximately 45%.2, 3, 4
The genes so far identified as involved in the aetiology of CMM have been either discovered in high-risk families – rare highly penetrant mutations such as those in the cell cycle regulators CDKN2A, CDK4 and INK4, or are common alleles at pigmentation loci such as MC1R, underlying well known phenotypic risk factors such as hair, skin and eye colour.5, 6, 7, 8
One of the greatest phenotypic risk factors for CMM is the number of acquired benign melanocytic naevi (moles) present on an individual's skin9 – presence of more than 4 moles (over 2 mm in diameter) on the arms increases CMM risk four-fold in pale-skinned individuals.10, 11 The exact nature of moles is still not completely clear. They are benign tumours of melanocytes and naevus cells that appear on the skin in childhood, increasing in number until early adulthood. Each mole is thought to represent the clonal expansion (10–23 doublings) of a single melanocyte12, 13, 14 harbouring a somatic mutation in a cell cycle regulatory or senescence gene.15, 16, 17 Melanocytes from up to 80% of moles carry BRAF or NRAS somatic mutations,18, 19, 20, 21 and transgenic fish expressing the commonest BRAFV600E mutant spontaneously develop naevus-like lesions.22
Twin analyses have shown naevus count to be more heritable than CMM risk per se.23, 24, 25, 26, 27 In carriers of the rare functional mutations of CDKN2A, nevogenesis, and thus total mole count, is generally increased.28 These naevi are often macroscopically unusual in appearance as well as exhibiting dysplastic histopathology. In a study of unselected adolescent twins,25 we showed a linkage of mole count and mole density to the region around CDKN2A on chromosome 9, though this finding was not replicated by a smaller study of older twins from the UK.27
In the present paper, we extend our previous report of linkage to markers near CDKN2A (chromosome 9p) and present results from the first whole genome scan of naevus count using a considerably expanded sample of adolescent twins and their families ascertained through Brisbane primary schools.
Results
Measured covariates and naevus counts
Box-Cox regressions, either including only an intercept or the covariates, found that a cube-root transformation of raised and flat naevus counts was superior to the more commonly used log transformation (Shapiro–Wilks test of normality of residuals, W=0. 9977, P=0.1232). A negative binomial generalised additive mixed model for the covariates also fitted well, giving a very similar distribution of residuals to that obtained from the cube-root transformed Gaussian model. In addition, the transformed values output by the SQTL program were also virtually identical to those from the regression model for the cube-root transformed data. Naevus counts were higher in males than females, increased with age up to 16 years old, were lower in redheads, and higher in lighter skinned individuals. Parental and self assessed sun exposure weakly increased naevus counts (see Table 1 and Figure 1).
Relationship between age and flat and raised mole counts from cross-sectional data (XS) on male and female adolescent twins and their siblings, including longitudinal data (Long) from two visits two years apart. Dense lines represent smoothed effects of age on mole count based on cross-sectional data from the first visit (these are fitted via locally weighted regression). The dotted lines represent changes in mole count from the first to the second visit for individuals who lie outside 1.5 times the interquartile range of counts.
Polygenic additive effects estimation
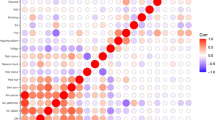
Multivariate genetic variance components for the three different type of naevus count were estimated by combining data from monozygotic and dizygotic twins and sibs in a classical twin family analysis. The proportion of variance due to additive polygenic effects for flat, raised and atypical mole counts were 57, 67 and 42%, respectively. There were moderate genetic correlations (0.40 between flat and atypical, 0.49 for flat with raised and 0.58 between raised and atypical) among the counts of the different types of naevus, as well as effects of family environment on flat and raised naevus counts (see Figure 2).
Multivariate genetic analysis (Cholesky decomposition) for flat, raised and atypical naevus count. A=additive polygenic effects, C=Common environmental effects and E=unique environmental effects. Standardized variance components contributions (proportion of total variance) are given beside paths, which are all positive.
Univariate genome scan results
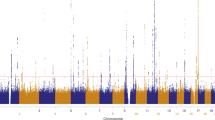
In Figure 3 each plot displays lod scores for three different types of naevus counts, as well as a multivariate lod combining all three counts. The strongest evidence for linkage was obtained for flat mole count at 2p and in the region of the CDKN2A gene on chromosome 9 (for both, lod=2.95, genome-wide P=0.08). Inclusion of fine mapping markers increased the lod for 9p to 3.43, and that obtained from SQTL to 3.27 (Figure 4). The highest lod score for raised mole count was 1.87 (genome-wide P=0.34) on chromosome 16 near D16S3068. For atypical mole count, there were three hits with lods of 2.20 (genome-wide P=0.39), 2.00 (P=0.48) and 2.00 (P=0.49) on chromosomes 1, 8 and X, respectively. These and lesser peaks of interest are listed in Table 2.
Variance components linkage analysis lod curves (MERLIN) for flat mole count around peak lods on four best chromosomes. For chromosome 9 (panel a), the maximum likelihood and semiparametric (SQTL) linkage lod scores are shown: black solid line, VC using 5 cM resolution marker set; red solid line, VC using scan and fine mapping markers; red broken line, semiparametric analysis using scan and fine mapping markers. Each x-axis tick represents a marker.
Multivariate genome scan results
The multivariate linkage results (see Figure 3, black curve) are consistent with the univariate results, notably for chromosome 9, but do show a chromosome 4 peak that is not seen for any of the individual phenotypes. For the chromosome 9 peak, the trait specific QTL heritabilities were 22, 7 and 5% for flat, raised and atypical naevus counts, respectively, at Visit 1, and 20, 0 and 18% at Visit 2. A model holding the heritabilities of the three types equal was rejected. Ideally, we would obtain the empirical genome-wide P-value but for the multivariate case this would take many months of computational time so we are restricted to the empirical pointwise P-value for the peak marker D9S925 which was 0.005, equivalent to a lod of about 2.0.
The peak markers on chromosome 4 were D4S406 and D4S402, with an empirical pointwise P-value of 0.001, equivalent to lod∼2.6. The trait specific heritabilities were 10, 0 and 2% for flat, raised and atypical naevus counts at Visit 1 (1, 1 and 3% at Visit 2).
Discussion
Regions linked to macular naevus count
Our phenotypic twin analyses of these data show that 60% of the variance in flat mole count is due to polygenic additive genetic factors, 30% to common family environment and 10% due to unique environmental variance, including errors of measurement (which would contribute half of this29). The genetic correlation between the subcounts of different mole types was only moderate, suggesting one might expect linkage to different regions, as well as some loci in common to all three mole types.
Likely candidate genes under linkage peaks for these phenotypes would include traditional oncogenes and tumour suppressors. As we have controlled for sun exposure, ancestry, facial freckling and skin and hair colouring, we would not expect any of these peaks to represent pigmentation genes such as MC1R and OCA2. A simple one-hit model for nevogenesis, following Blewitt,15 is that all common naevi represent effects of a single somatic mutation (such as BRAFV600E) in a melanocyte. In the absence of DNA repair and surveillance mechanisms, we might then expect the appearance of approximately 2000 naevi per individual (given a final melanocyte population of 2 × 109 and spontaneous somatic mutation rate of 1 × 10−6). The far lower observed mean mole count probably reflects the usually high efficiency of DNA damage surveillance and repair, essential in melanocytes given their role in photoprotection and known resistance to apoptotic signals following UV exposure.30 Environmental exposures that affect the somatic mutation rate (such as UV exposure) and genetic variation in known and yet to be characterised tumour suppressor genes will explain both interindividual difference and the considerable familial correlations in mole count.
The most significantly linked regions for flat mole count were: the telomeric region of chromosome 2p; chromosome 9p in the vicinity of CDKN2A; around D8S373 on chromosome 8q24.3; and chromosome 17p11, in the vicinity of the CMT1A duplication.
In an earlier study using a subset of these families, we detected linkage of mole count to the microsatellite marker D9S942 close to CDKN2A.25 In the present superset of families, using the 5 cM resolution marker set, evidence for linkage to chromosome 9p21 using only the genome scan markers peaked at a lod of 2.95. However, replicating the original single-point variance components analysis using the present set of families and the marker D9S942 obtained a lod score of 3.44, similar to that from multipoint analysis including additional fine-mapping markers in and around CDKN2A (lod=3.42; with SQTL lod=3.23). This lod score is of the same magnitude as that reported by Zhu et al25 despite a doubling in sample size. However, detailed analysis of the mole count data demonstrates that a cube root transformation is more appropriate than the log transform we had used earlier and produces more conservative lod-scores.
The drop-1 lod interval on the tip of chromosome 2p spans approximately 6 Mbp and contains numerous genes of unknown function. The region does contain the p53-responsive gene 2 (PRG2), one alias for which is ‘melanoma associated gene 50’,31 but tissue expression data and the known functions of the Drosophila homologue, peroxidasin, do not support this as a likely naevus gene.
There are 105 recognised genes within the confidence region on chromosome 8 in SWISS-PROT, TREMBL, mRNA and/or RefSeq. These include some members of the mitogen-activated protein kinase (MAPK) pathway, PTK2 (protein tyrosine kinase 2 and ERK8.
Among the known genes underlying the central region (11.6–14.1 Mbp) of the chromosome 17 peak, there are two good candidates for moliness, MAP2K4 and ELAC2. Mitogen-activated protein kinase 4 (MAP2K4/MKK4/SEK1) lies at 11.86 Mb and is a potent physiologic activator of the stress-activated protein kinases implicated in breast and pancreatic cancer.32, 33 It is in the JUN kinase pathway which is related to the RAS/RAF/MAPK pathway which in turn is heavily implicated in melanoma.18, 19, 34, 35 The recent work of Solit et al36 suggests cells carrying a BRAF mutation usually retain sensitivity to growth inhibition via blocking of the RAS/RAF/MEK/ERK pathway. ELAC2 (at 12.83 Mb, in the middle of the peak), has been identified as an hereditary prostate cancer gene.37 Meta analyses have found significant association between variants in ELAC2 and prostate cancer risk,38, 39 although the risk genotypes may cause only 2% of the population risk. Significantly, there is epidemiologic evidence of a link between familial prostate cancer and melanoma.40, 41 The other obvious chromosome 17 candidate, TP53, is approximately 6 Mbp distal to the most telomeric marker D17S299 of that interval.
Finally, the peak observed on chromosome 4 in the multivariate analysis was unusual, in that there were no corresponding peaks in the univariate analyses. As in the case of other peaks, the multivariate signal was being driven largely by flat mole count (with a specific QTL heritability of 10%). A gene of possible interest in this region is TIFA (TRAF-interacting protein with a forkhead-associated domain), in that it lies almost directly under the peak, and modulates TRAF6, involved in NF-κB activation; NF-κB is upregulated in dysplastic naevi and malignant melanomas.42
Higher heritability not associated with higher linkage peaks
By contrast with the results for flat mole count, and despite a slightly higher heritability, the genome scan showed only weakly suggestive linkage peaks for raised mole count on chromosomes 16p pericentromerically, and on distal 7p and 2q, none of which overlap with those for flat naevi. These peaks do not overlie obvious candidate genes. It is not clear whether this represents a genuine difference between the genetic architectures of the development of macular versus papular naevi. There is surprisingly little known about the natural history of flat and raised naevi. It is believed that raised naevi represent a later stage of development of flat naevi, but based on relatively small studies. Even though the sample size used in this study was larger than in the majority of genome scans of any trait to date, it is still well short of those required to reliably detect loci of even moderate effect.
As this paper was in process of revision, Falchi et al62 published a similar genome scan study of naevus count in 865 adult twin families. Using 194 families (141 DZ) where the twins were under the age of 35, they found evidence of linkage of log total naevus count to chromosomes 9p (lod=2.54 at D9S157) and 9q (lod=2.55 at D9S167) and to chromosome 5q (lod=3.47 at D5S638). These peaks did not appear in the older families, where the best linkage was to chromosome 2p25 (lod=2.75 at WIAF-933).
In conclusion, ours is a large study using a complete genome scan in an attempt to map genes responsible for three different types of naevus in adolescent twins. The peak on 9p21 lies directly over CDKN2A, which we have previously implicated in control of variation in flat mole count,25 and which is now confirmed by Falchi et al.62 We also find linkage of naevus count to multiple other regions, some not previously linked to melanoma, which may contain novel melanoma risk loci. Continuing expansion of our sample along with a planned genome-wide association scan will clarify the identity and importance of some of these loci.
Materials and methods
Twin sample
Twins were recruited in the context of an ongoing study of melanoma risk factors including benign melanocytic naevi (moles), sun exposure time and pigmentation related variables. The clinical protocol has been described in detail elsewhere.25, 43, 44 Briefly, twins were enlisted by contacting the principals of primary schools (first 7 years of education) in the greater Brisbane area, media appeals and by word of mouth. It is estimated that approximately 50 percent of the eligible birth cohort were recruited into the study. Twins were examined at age 12 years, and siblings at the same occasion if under 20 years of age, as described below. At the same time, twins and their parents completed questionnaires measuring risk factors, which we supplemented by sun diaries recorded at age 13 years. The examination of the twins was repeated at age 14 years, but not in the siblings.
The sample appears representative with respect to naevus count25 and IQ45 and it seems reasonable, therefore, to suppose it also representative of the Queensland population. Informed consent was obtained from all participants and parents before testing. This report is based on phenotype data collected from the study inception in May 1992 to February 2004.
Phenotype information
A nurse examined each individual and counted all naevi on the entire body surface, excluding buttocks, chest and abdomen. The naevi were counted and classified into three types: flat (macular), raised (papular) or atypical. The flat and raised naevus counts have a distribution with a long tail to the right, so for most purposes they are cube-root transformed to stabilise variances and improve normality. As atypical naevi were infrequent, counts of these have been binned into four categories (nil, 1–2, 3–4, five or more) and analysed as an ordinal variable.
The covariates used in the QTL model included sex, age, body surface area, hours of sun exposure, year and season examined and ancestry. Pigmentation variables such as skin colour, hair colour and freckling on the face were also used as covariates in the model, as detailed in Zhu et al.25
Naevus count and genotyping were available for 424 twin families (355 DZ, 69 MZ). Genotypes were available for both parents in 308 of these families, for one parent only in 74 families, and for neither parent in 42 families. One or more extra siblings were both phenotyped and genotyped in 133 DZ families and, of necessity for linkage analysis, all 69 MZ families. The number of offspring with unique genotypes (ie omitting MZ co-twins) was 1024 (874 from DZ families, 150 from MZ families). An additional 221 pairs of MZ twins have phenotypic (but no genotypic) information available, and inclusion of this group in the variance components model allows us to partition and stabilise the estimation of the proportion of variance due to common environmental effects.46
Zygosity and genotype information
Zygosity of same-sex twins was determined by typing nine independent DNA microsatellite polymorphisms plus the sex marker amelogenin at QIMR using the Profiler multiplex marker set (AmpFLSTRR Profiler PlusT, Applied Biosystems, Foster City, CA). The probability of dizygosity given concordance for all markers in our panel was <10−4. All twins and available parents were also typed for ABO, Rh and MNS blood groups by the Red Cross Blood Service, Brisbane.
Genome scans were carried out at the Australian Genome Research Facility, Melbourne (AGRF) and Center for Inherited Disease Research, Baltimore (CIDR). After integrating both genome scans, the marker set consisted of 762 highly polymorphic autosomal and 34 X-linked microsatellite markers at an average spacing of 4.8 cM. The average heterozygosity of markers was 0.78, and the mean information content was 0.77. Full details of the genome scan are provided elsewhere.47
In the region around CDKN2A, since we had previously observed linkage to mole count25 and because of the plausibility of this gene as a candidate, we genotyped an additional 19 microsatellite (D9S104, D9S126, D9S1604, D9S162, D9S1678, D9S169, D9S1748, D9S1749, D9S1875, D9S2136, D9S319, D9S52, D9S736, D9S942, D9S958, D9S974, D9S975, D9S976, IFNA) and SNP (CDKN2A*-981G>T, rs3731249, rs3731249, rs3814960, rs2069426, rs115115, rs3088440, rs12353062, rs10967819, rs1360589, rs1333039, rs1333050, rs1360590, rs1412829, rs1537370, rs1537371, rs1537375, rs1537378, rs1547704, rs1556516, rs1591136, rs1853186, rs2106119, rs2157719, rs7032979, rs7853090, rs7866783, rs944797, rs944800, rs974336) markers.
Statistical methods
In preliminary analyses, we tested various transformations of the mole count phenotypes, and performed generalized additive mixed model regression analyses versus the measured covariates, including family as a random effect. These analyses were carried out using the R statistical package,48 and specifically the mgcv package.49
The genotype and pedigree data were tested for errors: Mendelian inconsistencies were checked using Sib-pair;50 GRR51 and Relpair52 were used to identify pedigree errors such as those due to sample mix up; potential genotyping errors were detected using MENDEL,53 MERLIN54 and SIBMED.55 The multipoint identity by descent (IBD) probabilities were estimated by using MERLIN for autosomal markers and MINX54, 56 for chromosome X markers. For the fine mapping markers around CDKN2A, we used the new ‘cluster’ approach implemented in MERLIN 1.0 to form haplotypes from markers in linkage disequilibrium before IBD estimation. Structural equation modelling analyses were performed using the Mx software package57 as described elsewhere.25, 47, 58 Both univariate (each mole count variable in turn) and multivariate (flat, raised and atypical mole count) variance components linkage analyses were performed, using MERLIN 1.0 (univariate), and Mx and MENDEL 6.0 (multivariate). The multivariate analyses incorporate measures from both visits by the twins, so there are up to six mole count variables in these analyses. The genetic and environmental covariances between mole counts at the two visits were left unconstrained, to allow for the presence of gene by age interaction. That is, we have fitted repeated measures models with random regression terms for visit, thus incorporating the data from both occasions, but allowing for the testing for genetic heterogeneity (which was absent).
We have also analysed these data using the recently described semi-parametric variance components linkage analysis approach, implemented in the program SQTL.59 In this approach, optimal transformation of the data to fit the underlying hypothesis of multivariate normality of the variance components analysis is included in the semiparametric likelihood evaluated.60
For the univariate genetic analyses, we calculated genome-wide significance levels based on analysis of 1000 gene-dropping simulated datasets generated by the MERLIN computer program. The highest lod score for each chromosome was kept and the number of false positives counted as per Kruglyak and Daly.61
The empirical thresholds for suggestive linkage (one expected false positive per genome scan) were ∼1.7 for flat and atypical mole count, and ∼1.4 for raised mole count. The thresholds for significant linkage (one expected false positive per 20 genome scans) were ∼3.3 for flat, ∼3 for raised and ∼3.65 for atypical mole counts. Production of genome-wide thresholds was not feasible for the multivariate linkage analyses, as they are computationally intensive, but we did estimate a point-wise empirical P-value for the highest peaks and other several comparison regions to test the adequacy of the asymptotic P-values generated assuming the test statistic was χ2 with 6 degrees of freedom. These simulations found the test statistic was better described as a χ2 variate with 8.9 degrees of freedom, and this interpolation has been used to adjust the plotted asymptotic P-values for the multivariate analysis.
References
MacLennan R, Green AC, McLeod GR, Martin NG : Increasing incidence of cutaneous melanoma in Queensland, Australia. J Natl Cancer Inst 1992; 84: 1427–1432.
Aitken JF, Duffy DL, Green A, Youl P, MacLennan R, Martin NG : Heterogeneity of melanoma risk in families of melanoma patients. Am J Epidemiol 1994; 140: 961–973.
Aitken JF, Bailey-Wilson J, Green AC, MacLennan R, Martin NG : Segregation analysis of cutaneous melanoma in Queensland. Genet Epidemiol 1998; 15: 391–401.
Do KA, Aitken JF, Green AC, Martin NG : Analysis of melanoma onset: assessing familial aggregation by using estimating equations and fitting variance components via Bayesian random effects models. Twin Res 2004; 7 (1): 98–113.
Palmer JS, Duffy DL, Box NF et al: MC1R polymorphisms and risk of melanoma: is the association explained solely by pigmentation phenotype? Am J Hum Genet 2000; 66: 176–186.
Box NF, Duffy DL, Chen W et al: MC1R genotype modifies risk of melanoma in families segregating CDKN2A mutations. Am J Hum Genet 2001; 69: 765–773.
Goldstein AM, Tucker MA : Genetic epidemiology of cutaneous melanoma: a global perspective. Arch Dermatol 2001; 137: 1493–1496.
Newton Bishop JA, Harland M, Bennett DC et al: Mutation testing in melanoma families: INK4A, CDK4 and INK4D. Br J Cancer 1999; 80: 295–300.
Gandini S, Sera F, Cattaruzza MS et al: Meta-analysis of risk factors for cutaneous melanoma: I. Common and atypical naevi. Eur J Cancer 2005; 41: 28–44.
Swerdlow AJ, Green A : Melanocytic naevi and melanoma: an epidemiological perspective. Br J Dermatol 1987; 117: 137–146.
Grulich AE, Bataille V, Swerdlow AJ et al: Naevi and pigmentary characteristics as risk factors for melanoma in a high-risk population: a case-control study in New South Wales, Australia. Int J Cancer 1996; 67: 485–491.
Robinson WA, Lemon M, Elefanty A, Smith MH, Markham N, Norris D : Human acquired naevi are clonal. Melanoma Res 1998; 8: 499–503.
Indsto JO, Cachia AR, Kefford RF, Mann GJ : X inactivation, DNA deletion, and microsatellite instability in common acquired melanocytic nevi. Clin Cancer Res 2001; 7: 4054–4059.
Hui P, Perkins A, Glusac E : Assessment of clonality in melanocytic nevi. J Cutan Pathol 2001; 28: 140–144.
Blewitt RW : Single genetic mutations can account for melanocytic naevi. Br J Dermatol 2005; 152: 896–902.
Bennett DC : Human melanocyte senescence and melanoma susceptibility genes. Oncogene 2003; 22: 3063–3069.
Michaloglou C, Vredeveld LC, Soengas MS et al: BRAFE600-associated senescence-like cell cycle arrest of human naevi. Nature 2005; 436: 720–724.
Pollock PM, Harper UL, Hansen KS et al: High frequency of BRAF mutations in nevi. Nat Genet 2003; 33: 19–20.
Yazdi AS, Palmedo G, Flaig MJ et al: Mutations of the BRAF gene in benign and malignant melanocytic lesions. J Invest Dermatol 2003; 121: 1160–1162.
Kumar R, Angelini S, Snellman E, Hemminki K : BRAF mutations are common somatic events in melanocytic nevi. J Invest Dermatol 2004; 22: 342–348.
Turner DJ, Zirvi MA, Barany F, Elenitsas R, Seykora J : Detection of the BRAF V600E mutation in melanocytic lesions using the ligase detection reaction. J Cutaneous Pathol 2005; 32 (5): 334–339.
Patton EE, Widlund HR, Kutok JL et al: BRAF mutations are sufficient to promote nevi formation and cooperate with p53 in the genesis of melanoma. Curr Biol 2005; 15: 249–254.
Siemens HW : Der Zwillingspathologie. Berlin: Springer, 1925.
Duffy DL, Macdonald AM, Easton DF, Ponder BAJ, Martin NG : Is the genetics of moliness simply the genetics of sun exposure? A path analysis of nevus counts in British twins, Genetic Analysis Workshop 7: Issues in gene mapping and the detection of major genes. Cytogenet Cell Genet 1992; 59: 194–196.
Zhu G, Duffy DL, Eldridge A et al: A major quantitative-trait locus for mole density is linked to the familial melanoma gene CDKN2A: a maximum-likelihood combined linkage and association analysis in twins and their sibs. Am J Hum Genet 1999; 65: 483–492.
Bataille V, Snieder H, MacGregor AJ, Sasieni P, Spector TD : Genetics of risk factors for melanoma: an adult twin study of nevi and freckles. J Natl Cancer Inst 2000; 92: 457–463.
Barrett JH, Gaut R, Wachsmuth R, Bishop JA, Bishop DT : Linkage and association analysis of nevus density and the region containing the melanoma gene CDKN2A in UK twins. Br J Cancer 2003; 88: 1920–1924.
Florell SR, Meyer LJ, Boucher KM et al: Longitudinal assessment of the nevus phenotype in a melanoma kindred. J Invest Dermatol 2004; 123: 576–582.
Aitken JF, Green A, Eldridge A et al: Comparability of naevus counts between and within examiners, and comparison with computer image analysis. Br J Cancer 1994; 69: 487–491.
Kadekaro AL, Kavanagh R, Kanto H et al: Alpha-melanocortin and endothelin-1 activate antiapoptotic pathways and reduce DNA damage in human melanocytes. Cancer Res 2005; 65: 4292–4299.
Weiler SR, Taylor SM, Deans RJ, Kan-Mitchell J, Mitchell MS, Trent JM : Assignment of a human melanoma associated gene MG50 (D2S448) to chromosome 2p25.3 by fluorescence in situ hybridization. Genomics 1994; 22: 243–244.
Xin W, Yun KJ, Ricci F et al: MAP2K4/MKK4 expression in pancreatic cancer: genetic validation of immunohistochemistry and relationship to disease course. Clin Cancer Res 2004; 10: 8516–8520.
Wang L, Pan Y, Dai JL : Evidence of MKK4 pro-oncogenic activity in breast and pancreatic tumors. Oncogene 2004; 23: 5978–5985.
Davies H, Bignell GR, Cox C et al: Mutations of the BRAF gene in human cancer. Nature 2002; 417: 949–954.
Dong J, Phelps RG, Qiao R et al: BRAF oncogenic mutations correlate with progression ratherthan initiation of human melanoma. Cancer Res 2003; 63: 3883–3885.
Solit DB, Garraway LA, Pratilas CA et al: BRAF mutation predicts sensitivity to MEK inhibition. Nature 2006; 439: 358–362.
Tavtigian SV, Simard J, Teng DH et al: A candidate prostate cancer susceptibility gene at chromosome 17p. Nat Genet 2001; 27: 172–180.
Camp NJ, Tavtigian SV : Meta-analysis of associations of the Ser217Leu and Ala541Thr variants in ELAC2 (HPC2) and prostate cancer. Am J Hum Genet 2002; 71: 1475–1478.
Rennert H, Zeigler-Johnson CM, Addya K et al: Association of susceptibility alleles in ELAC2/HPC2, RNASEL/HPC1, and MSR1 with prostate cancer severity in European American and African American men. Cancer Epidemiol Biomarkers Prev 2005; 14: 457–949.
Albright LA, Schwab A, Camp NJ, Farnham JS, Thomas A : Population-based risk assessment for other cancers in relatives of hereditary prostate cancer (HPC) cases. Prostate 2005, DOI:10.1002/pros.20248.
Hemminki K, Chen B : Familial association of prostate cancer with other cancers in the Swedish Family-Cancer Database. Prostate 2005, DOI:10.1002/pros.20284.
Ueda Y, Richmond A : NF-κB activation in melanoma. Pigment Cell Res 2006; 19: 112–124.
Aitken JF, Green AC, MacLennan R, Youl P, Martin NG : The Queensland Familial Melanoma Project: study design and characteristics of participants. Melanoma Res 1996; 6 (2): 155–165.
McGregor B, Pfitzner J, Zhu G et al: Genetic and environmental contributions to size, colour, shape and other characteristics of melanocytic naevi in a sample of adolescent twins. Genet Epidemiol 1999; 16: 40–53.
Wright M, De Geus E, Ando J et al: Genetics of cognition: outline of a collaborative twin study. Twin Res 2001; 4: 48–56.
Evans DM, Medland SE : A note on including phenotypic information from monozygotic twins in variance components QTL linkage analysis. Ann Hum Genet 2004; 67: 613–617.
Zhu G, Evans DM, Duffy DL et al: A genome scan for eye color in 502 twin families: most variation is due to a QTL on chromosome 15q. Twin Res 2004; 7: 197–210.
R Development Core Team: R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, 2005. ISBN 3–900051–07–0, URL http://www.R-project.org.
Wood SN : Stable and efficient multiple smoothing parameter estimation for generalized additive models. J Am Statist Assoc 2004; 99: 673–686.
Duffy DL : Sib-pair computer program. Version 0.99. Brisbane: Queensland Institute of Medical Research, 2004.
Abecasis GR, Cherny SS, Cookson WO, Cardon LR : GRR: graphical representation of relationship errors. Bioinformatics 2001; 17 (8): 742–743.
Epstein MP, Duren WL, Boehnke M : Improved inference of relationship for pairs of individuals. Am J Hum Genet 2000; 67: 1219–1231.
Lange K, Cantor R, Horvath S et al: MENDEL Version 4.0: A complete package for the exact genetic analysis of discrete traits in pedigrees and population data sets. Am J Hum Genet 2001; 69 (Suppl): A1886.
Abecasis GR, Cherny SS, Cookson WOC, Cardon LR : MERLIN – Rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002; 30: 97–101.
Douglas JA, Boehnke M, Lange K : A multipoint method for detecting genotyping errors and mutations in sibling-pair linkage data. Am J Hum Genet 2000; 66: 1287–1297.
Abecasis GR, Wigginton JE : Handling marker-marker linkage disequilibrium: pedigree analysis with clustered markers. Am J Hum Genet 2005; 77: 754–767.
Neale MC, Boker SM, Xie G, Maes HH : Mx: Statistical Modeling. VCU Box 900126, 6th Edition. Richmond, VA 23298: Department of Psychiatry, 2003.
Zhu G, Duffy DL, Turner DR, Ewen KR, Montgomery GW, Martin NG : Linkage and association analysis of radiation damage repair genes XRCC3 and XRCC5 with nevus density in adolescent twins. Twin Res 2003; 6: 315–321.
Diao G, Lin DY : A powerful and robust method for mapping quantitative trait loci in general pedigrees. Am J Hum Genet 2005; 77: 97–111.
Bickel PJ, Klassen CAJ, Ritov Y, Wellner JA : Efficient and adaptive estimation in semiparametric models. Baltimore: Johns Hopkins University Press, 1993.
Kruglyak L, Daly MJ : Linkage thresholds for two-stage genome scans. Am J Hum Genet 1998; 62: 994–997.
Falchi F, Spector TD, Perks U, Kato BS, Bataille V : Genome-wide search for nevus density shows linkage to two melanoma loci on chromosome 9 and identifies a new QTL on 5q31 in an adult twin cohort. Human Molecular Genetics, Advance Access published on August 23, 2006. doi:10.1093/hmg/ddl227.
Acknowledgements
Collection of phenotypes and DNA samples was supported by grants from the Queensland Cancer Fund (NGM, NKH), the Australian National Health and Medical Research Council (950998, 981339 and 241944; NGM), and the US National Cancer Institute (CA88363; NKH, NGM, DLD, GWM). The genome scans were supported by the Australian NHMRC's Program in Medical Genomics (NHMRC 219178; NGM, GWM, DLD) and the Center for Inherited Disease Research (CIDR; Director, Dr Jerry Roberts) at The Johns Hopkins University (JMT, NGM). CIDR is fully funded through a federal contract from the National Institutes of Health to The Johns Hopkins University (Contract Number N01-HG-65403). We thank Ann Eldridge, Marlene Grace for phenotype collection, Megan Campbell and Anjali Henders for managing sample processing and preparation; and the twins, their siblings, and their parents for their cooperation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhu, G., Montgomery, G., James, M. et al. A genome-wide scan for naevus count: linkage to CDKN2A and to other chromosome regions. Eur J Hum Genet 15, 94–102 (2007). https://doi.org/10.1038/sj.ejhg.5201729
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5201729
Keywords
This article is cited by
-
Half the Genetic Variance in Vitamin D Concentration is Shared with Skin Colour and Sun Exposure Genes
Behavior Genetics (2019)
-
The continuing value of twin studies in the omics era
Nature Reviews Genetics (2012)
-
Melanoma Genetics: Recent Findings Take Us Beyond Well-Traveled Pathways
Journal of Investigative Dermatology (2012)
-
Genome-wide association studies and genetic architecture of common human diseases
BMC Proceedings (2011)
-
A novel recurrent mutation in MITF predisposes to familial and sporadic melanoma
Nature (2011)