Abstract
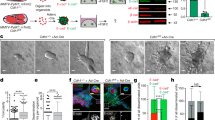
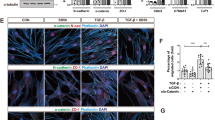
Using a validated tetracycline (tet)-regulated MCF7-founder (MCF7F) expression system to modulate expression of CD44 standard form (CD44s), we report the functional importance of CD44s and that of a novel transcriptional target of hyaluronan (HA)/CD44s signaling, EMS1/cortactin, in underpinning breast cancer metastasis. In functional experiments, tet-regulated induction of CD44s potentiated the migration and invasion of MCF7F cells through HA-supplemented Matrigel. EMS1/cortactin was identified by expression profiling as a novel transcriptional target of HA/CD44 signaling, an association validated by quantitative PCR and immunoblotting experiments in a range of breast cancer cell lines. The mechanistic basis underpinning CD44-promoted transcription of EMS1/cortactin was shown to be dependent upon a NFκB mechanism, since pharmacological inhibition of IκKinase-2 or suppression of p65 Rel A expression attenuated CD44-induced increases in cortactin mRNA transcript levels. Overexpression of a c-myc tagged murine cortactin construct in the weakly invasive, CD44-deficient MCF7F and T47D cells potentiated their invasion. Furthermore, the functional importance of cortactin to CD44s-promoted metastasis was demonstrated by selective suppression of cortactin in CD44-expressing MCF7F-B5 and MDA-MB-231 breast cancer cells using RNAi, which was shown to result in attenuated CD44-promoted invasion and CD44-promoted adhesion to bone marrow endothelial cells (BMECs).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Al-Hajj M, Wicha MS, Benito-Hernandez A, Morrison SJ, Clarke MF . (2003). Proc Natl Acad Sci U S A 100: 3983–3988.
Arderiu G, Diaz-Ricart M, Buckley B, Escolar G, Ordinas A . (2002). Biochem J 364: 65–71.
Auvinen P, Tammi R, Parkinnen J, Tammi M, Agren U, Johansson R et al. (2000). Am J Pathol 156: 529–536.
Berner HS, Nesland JM . (2001). Breast Cancer Res Treat 65: 23–29.
Bourguignon LYW, Gunja-Smith N, Iida N, Zhu HB, Young LJ, Muller WJ et al. (1998). J Cell Physiol 176: 206–215.
Bourguignon LYW, Singleton PA, Diedrich F . (2004). J Biol Chem 279: 29654–29669.
Bourguignon LYW, Zhu H, Shao L, Chen YW . (2000). J Biol Chem 275: 1829–1838.
Bourguignon LYW, Zhu H, Shao L, Chen YW . (2001). J Biol Chem 276: 7327–7336.
Bourguignon LYW, Zhu H, Shao L, Zhu D, Chen YW . (1999). Cell Motil Cytoskeleton 43: 269–287.
Bowden ET, Barth M, Thomas D, Glazer RI, Mueller SC . (1999). Oncogene 18: 4440–4449.
Bryce NS, Clark ES, Leysath JL, Currie JD, Webb DJ, Weaver AM . (2005). Curr Biol 15: 1276–1285.
Draffin JE, McFarlane S, Hill A, Johnston PG, Waugh DJJ . (2004). Cancer Res 64: 5702–5711.
Fitzgerald KA, Bowie AG, Sheehy-Skeffington B, O'Neill LAJ . (2000). J Immunol 164: 2053–2063.
Herrera-Gayol A, Jothy S . (2001). Int J Exp Pathol 82: 193–200.
Hirao M, Sato N, Kondo T, Yonemura S, Monden M, Sasaki T et al. (1996). J Cell Biol 135: 37–51.
Hui R, Campbell DH, Lee CS, McCaul K, Horsfall DJ, Musgrove EA et al. (1997). Oncogene 15: 1617–1623.
Isacke CM, Yarwood H . (2002). Int J Biochem & Cell Biol 34: 718–721.
Kinoshita J, Haga S, Shimizu T, Imamura H, Watanabe O, Kajiwara T . (2000). Breast Cancer Res Treat 53: 177–183.
Kozlow W, Guise TA . (2005). J Mammary Gland Biol Neoplasia 10: 169–180.
Lee JY, Spicer AP . (2000). Curr O pin Cell Biol 12: 581–586.
Lester BR, McCarthy JB . (1992). Cancer Metastasis Rev 11: 31–44.
Li Y, Tondravi M, Lui J, Smith E, Haudenschild CC, Kaczmarek M et al. (2001). Cancer Res 61: 6906–6911.
McKee CM, Penno MB, Cowman M, Burdick MD, Strieter RM, Bao C et al. (1996). J Clin Invest 98: 2403–2413.
Monaghan M, Mulligan K, Gillespie H, Trimble A, Winter P, Johnston PG et al. (2000). J Pathol 192: 519–525.
Mullins RD . (2000). Curr Opin Cell Biol 12: 91–96.
Oliferenko S, Kaverina I, Small JV, Huber LA . (2000). J Cell Biol 148: 1159–1164.
Rodrigo JP, Garcia LA, Ramos S, Lazo PS, Suarez C . (2000). Clin Cancer Res 6: 3177–3182.
Sayegh TY, Arora PD, Fan L, Laschinger CA, Greer PA, McCulloch CA et al. (2005). Mol Biol Cell 12: 5514–5527.
Schuuring E, Verhoeven E, Mooi WJ, Michalides RJ . (1992). Oncogene 7: 355–361.
Tilghman RW, Hoover RL . (2002). FASEB J 16: 1257–1259.
Timpson P, Lynch DK, Schramek D, Walker F, Daly RJ . (2005). Cancer Res 65: 3273–3280.
Toole BP . (2000). Glycobiology 12: 37R–42R.
Tsukita S, Oishi K, Sato N, Sagara J, Kawai A, Tsukita S . (1994). J Cell Biol 126: 391–401.
Urono T, Lui J, Zhang P, Fan YX, Egile C, Li R et al. (2001). Nat Cell Biol 3: 259–266.
Van Rossum AG, Gibcus J, van der Wal J, Schuuring E . (2005). Breast Cancer Res 7: 235–237.
Weaver AM, Karginov AV, Kinley AW, Weed SA, Li Y, Parsons JT et al. (2001). Curr Biol 11: 370–374.
Weber GF, Bronson RT, Ilagan J, Cantor H, Schmits R, Mak TW . (2002). Cancer Res 62: 2281–2286.
Yamaguchi H, Wyckoff J, Condeelis J . (2005). Curr Opin Cell Biol 17: 559–564.
Zhu D, Bourguignon LYW . (1998). Cell Motil Cytoskel 39: 209–222.
Acknowledgements
Microarray analysis was conducted by Dr Kevin Robertson (Genomic Technology and Informatics, University of Edinburgh). We thank Dr Stephen Moore and Dr Austin Tanney (Almac-Diagnostics, Craigavon, Northern Ireland) for statistical analysis of the microarray data, Dr George Tzircotis (Institute of Cancer Research, London) for provision of the FITC-labelled HA, Professor Thomas Parsons (University of Virginia, Charlottesville) for the gift of the c-myc-tagged cortactin construct, Dr Babbette Weksler (Cornell University, NY) for the transformed human BMEC cell line and Professor Luke O'Neill (Trinity College, Dublin) for the luciferase reporter constructs. Finally, we extend our thanks to Professor Peter Hamilton (QUB) for assistance with microscopy and imaging studies and Professor Peter Hall (QUB) for access to real-time PCR instrumentation. This work was supported by a research grant from Cancer Research UK (A4106/C11512) to DJJW.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hill, A., McFarlane, S., Mulligan, K. et al. Cortactin underpins CD44-promoted invasion and adhesion of breast cancer cells to bone marrow endothelial cells. Oncogene 25, 6079–6091 (2006). https://doi.org/10.1038/sj.onc.1209628
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209628
Keywords
This article is cited by
-
Effects of CD44 siRNA on inhibition, survival, and apoptosis of breast cancer cell lines (MDA-MB-231 and 4T1)
Molecular Biology Reports (2024)
-
Cav2.2-NFAT2-USP43 axis promotes invadopodia formation and breast cancer metastasis through cortactin stabilization
Cell Death & Disease (2022)
-
Profiling the expression of pro-metastatic genes in association with the clinicopathological features of primary breast cancer
Cancer Cell International (2021)
-
CD146, a novel target of CD44-signaling, suppresses breast tumor cell invasion
Cell Communication and Signaling (2017)
-
WNT5A signaling impairs breast cancer cell migration and invasion via mechanisms independent of the epithelial-mesenchymal transition
Journal of Experimental & Clinical Cancer Research (2016)