Abstract
The Rubinstein-Taybi Syndrome (RSTS) is a congenital disease that affects brain development causing severe cognitive deficits. In most cases the disease is associated with dominant mutations in the gene encoding the CREB binding protein (CBP). In this work, we present the first quantitative analysis of brain abnormalities in a mouse model of RSTS using magnetic resonance imaging (MRI) and two novel self-developed automated algorithms for image volumetric analysis. Our results quantitatively confirm key syndromic features observed in RSTS patients, such as reductions in brain size (−16.31%, p < 0.05), white matter volume (−16.00%, p < 0.05) and corpus callosum (−12.40%, p < 0.05). Furthermore, they provide new insight into the developmental origin of the disease. By comparing brain tissues in a region by region basis between cbp+/− and cbp+/+ littermates, we found that cbp haploinsufficiency is specifically associated with significant reductions in prosencephalic tissue, such us in the olfactory bulb and neocortex, whereas regions evolved from the embryonic rhombencephalon were spared. Despite the large volume reductions, the proportion between gray-, white-matter and cerebrospinal fluid were conserved, suggesting a role of CBP in brain size regulation. The commonalities with holoprosencephaly and arhinencephaly conditions suggest the inclusion of RSTS in the family of neuronal migration disorders.
Similar content being viewed by others
Introduction
The Rubinstein-Taybi syndrome (RSTS, MIM 180849)1 is a congenital, autosomal dominant disorder with rare incidence (1:100,000—1:125,000 human newborns)2,3,4. The clinical picture of the syndrome was first described in 19575 and includes mental impairment, growth delay, skeletal abnormalities, hypertelorism and microcephaly6,7. The main cause of RSTS are heterozygous mutations of the CREBBP gene (chromosome 16p13.3)8, which occurs in 40–60% of the diagnoses3,9,10. An alternative cause is a mutated EP300 gene (22q13.2), which happens in 3% of the RSTS diagnoses11. CREBBP and EP300 encode two paralog transcriptional co-activators with intrinsic lysine acetyltransferase (KAT) activity known as CREB binding protein (CBP) and p3007, respectively. Both proteins are highly similar and thought to play a major role in transcriptional regulation. Research in mouse models of RSTS, such as the Crebbp haploinsufficient mouse (cbp+/−)12 has revealed that these animals present similar features to patients, specifically cognitive impairment, growth delay and anatomical abnormalities. Additionally, it has been recently shown that CBP also plays a critical role in the regulation of adult neurogenesis13 and the differentiation of the cortical neural progenitor cells14.
A considerable number of studies and case reports have included brain imaging exams in RSTS patients15,16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38. These studies have consistently shown that microcephalia is a main feature in RSTS, as confirmed by a meta-analysis of 732 patients39. Other abnormalities, such as white matter abnormalities25,26,31, corpus callosum dysgenesis17,24,26,33,35 and abnormalities20,27 and cortical dysmorphologies31,35 including pachygiria21 and malgyration35 have been also reported. All these features may be correlated with the characteristic cognitive impairment. Preliminary brain images studies in mouse models of RSTS have also revealed some gross abnormalities in brain size13,40.
Most of these studies, both in mice and humans, were qualitative and a precise description of the impact of CBP deficiency in brain development and structure is still lacking. To address this situation, we performed a quantitative volumetric study of brain structure in the RSTS mouse model of Crebbp haploinsufficient. Whereas traditional volumetry, also known as stereology, calculates these volumes based on histologic sections of the ex vivo brain, neurocomputational volumetry techniques allow the automatic analysis of brain images acquired in vivo by Magnetic Resonance Imaging (MRI) or Computed Tomography (CT), avoiding morphometric distortions introduced by histological tissue preparation41 and therefore providing a more precise quantification of brain anatomy42. We have used two in-house developed algorithms for automated volumetry analysis based on MRI images acquired at 7T to unveil the selective impact of CBP deficiency in the development of different brain regions. We provide novel evidences suggesting that the brain of RSTS mice present holoprosencephaly, composed of important dysgenesis in the olfactory bulb and cortex and suggestive of an underlying neuronal migration disorder. The latter conclusion is also supported by case reports in RSTS patients reporting cortical abnormalities.
Materials and Methods
Animal preparation
Cbp+/− mice12 (CBP group) and control littermates (WT group) were generated as previously described13. Adult mice (95 to 118 days old, WT group: 104,17 ± 7,70 days old, CBP group: 106,00 ± 7,00 days old) were weighted 14.5 to 28 g (WT group: 24.6 ± 2.7 g, CBP group: 17.8 ± 2.3 g, total: 21.5 ± 4.3 g). These mice are maintained on a DBA and C57BL/6J mixed background because cbp+/− mice are not viable in a pure C57BL/6J background40,43. Mice were maintained according to animal care standards established by the European Union and all the protocols were approved by the Institutional Animal Care and Use Committee and carried-out in accordance with the approved guidelines.
Image acquisition
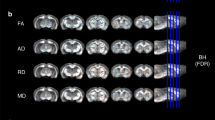
Before the image acquisition, the animals were anesthetized with isoflurane. During the acquisition, the head of the animal was situated in a stereotactic frame while temperature, cardiac frequency and oxygen saturation (SpO2) were monitored. The images were acquired with a 7-T MRI scanner (Bruker BioSpin, Ettlingen, Germany) under coronal 3D T1-weighted configuration, using an EPI sequence with the following parameters: TR = 360 ms, TE = 10 ms, 35 averages, in plane resolution 86 × 86 μm, slice thickness 0.35 mm, with a matrix size of 256 × 256, obtaining 17 DICOM images (Fig. 1a).
Block diagram and intermediate results of the methods.
The preparation of the 11 animals (6 WT, 5 CBP) is followed by the acquisition of magnetic resonance images (Panel (a)). The common image preprocessing stage is composed by format conversion, reorientation (Panel (b)) and brain segmentation, which produces an extracted brain image (Panel (c)). At this point, the procedure splits into two image processing algorithms. Brain tissues volumetry further segments the previously extracted brain into three substances (grey matter, white matter and cerebrospinal fluid, Panels (d–f)) and obtains their volumes for all the subjects. On the other hand, brain regions volumetry implies a combination of normalizations to fit the atlas to each of the brain images in order to get the regional volumes of the subjects (Panel (g)). Progressive overlappings (1–3) of segmented regions over brain axial slices from selected CBP (Panel (g), left column) and WT (Panel (g), right column) subjects are shown. All the resulting volumes were compared by means of t-tests.
Common image preprocessing
The common image preprocessing stage consisted in three tasks. Firstly, the DICOM images were converted to the NIfTI format. Afterwards, in order to accommodate them to the software that implements the two image processing algorithms (further detailed below), the images were reoriented by means of the FMRIB Software Library (FSL)44 (Oxford Centre for Functional MRI of the Brain, Oxford, United Kingdom) (Fig. 1b). Finally, brain segmentation was performed by means of the Brain Extraction Tool (BET)45, which is also included in FSL. This operation basically consists in skull-stripping by fitting a deformable model to the brain surface, thus producing the clean intracranial volume used for brain tissues and brain regions volumetry (Fig. 1c).
Brain Tissues Volumetry
For the volumetric analysis of the brain tissues, the already extracted brain images were segmented in their three constituent tissues, i.e., grey matter (GM), white matter (WM) and cerebrospinal fluid (CSF) (Fig. 1d–f). This process was carried out with the Unified Segmentation algorithm46, included in the Statistical Parametric Mapping (SPM) software (Wellcome Trust Centre for Neuroimaging, London, United Kingdom). This algorithm also spatially normalizes the images with the assistance of mouse brain tissue probability maps (TPM)47. Each one of the three resulting images is composed by voxels whose intensity indicates the probability of belonging to the correspondent tissue. In order to render the voxels that most probably belong to each of the three tissues, the three segments were binarized. Lastly, a self-developed algorithm was run to compute the volume of each tissue by summing all the belonging voxels and multiplying by the voxel volume. Two-tailed Student’s t-tests were applied in order to show significant differences in the volume values between CBP and WT groups.
Brain Regions Volumetry
For the brain regions volumetric analysis, the already extracted brain images were segmented in their 43 constituent regions, according to the Kovačević atlas of the mouse brain48. This process was carried out by superposing the atlas to each individual brain image, which was done by a combination of spatial transformations. Firstly, the atlas was spatially normalized from its native space to the space of the TPM and then the atlas in the space of the TPM was spatially normalized in order to adapt to each of the 11 image spaces, each one from one individual (Fig. 1g). Finally, in a similar way to the brain tissue volumetry, the volume of each region was computed by summing all the belonging voxels and multiplying by the voxel volume prior to two-tailed Student’s t-tests that show significant differences among both groups.
Results
While there have been few in vivo volumetry studies in mouse models of intellectual disability disorders42,49, to the best of our knowledge we have developed and applied the first automated volumetry analysis of the mouse brain conducted in vivo.
Brain tissue volumetry
We report the volumes of gray matter (GM), white matter (WM) and cerebrospinal fluid (CSF) from 11 subjects (5 cbp+/− mice and 6 wild type littermates). Results reveal that the brains of CBP group are smaller than those of the WT group (Fig. 2). Specifically, GM, WM and CSF volumes are significantly lower in CBP subjects (−16.43% [p < 0.0001], −16.00% [p = 0.005] and −34.55% [p = 0.044], respectively). Similarly, brain volume, composed by GM and WM, significantly changed by −16.31% (p < 0.0001) whereas total intracranial volume (TICV), formed by the brain and CSF volumes, significantly changed by −17.20% (p < 0.0001). In contrast to the absolute volume values, the proportions between the three tissue classes are kept constant in both groups. Regarding brain tissue ratios, the GM- and WM-to-brain-volume ratios along with GM-, WM-, CSF- and brain-to-TICV ratios did not significantly change between groups (p > 0.05). In other words, the brains are reduced in volume but the ratios are unaltered, which leads us to preliminary conclude that, CBP hemydeficiency provokes a down-scaling effect in the brain.
Results of the brain tissues volumetry analysis.
Volumes (mean ± standard deviation) of the three brain tissues (Grey Matter, White Matter and Cerebrospinal Fluid) and their ratios are represented for Wildtype (6 subjects) and CBP group (5 subjects). Both groups show similar proportions of the three matters, which allows us to conclude that Rubinstein-Taybi Syndrome has no effect in terms of brain tissues proportion.
Brain regions volumetry
We report the volumes of 43 brain regions (according to the Kovačević atlas of the mouse brain48) for each of the 11 subjects. In 41 of the 43 regions of the CBP group, a volume reduction was observed (obtaining a volume change of −10.5 ± 4.6% [mean ± standard deviation]), being significant (p < 0.05) in 25 of the 41 regions (mean volume change of −13.34 ± 1.41%).
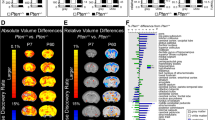
When the changes in volume per region were mapped on the mouse brain atlas (Fig. 3), we identified three zones of different developmental origin showing differential behaviors. First, the posterior part of the brain (cerebellum and brain stem), which is derived from the embryonic rhombencephalon, showed no significant reduction in volume (−0.68 ± 0.21%). Second, regions that are developmentally derived from the embryonic prosencephalon, such as the telencephalon and diencephalon, showed the largest reduction in volume (−12.51 ± 0.36%). Finally, the anterior part of the brain (olfactory bulb) showed a lower but significant volume reduction (−6.28%). Interestingly, our quantification reveals that cortical and corpus callosum volumes are equally reduced (−12.79% and −12.40% respectively).
Results of the brain regions volumetry analysis.
Colored maps indicating regional volume change (in %) of CBP group mean brain with respect to WT group are shown at axial slices Z = 2 (a), Z = 5 (b), Z = 9 (c) and Z = 14 (d) of the three-dimensional magnetic-resonance image. Brain regions can be grouped in three zones according to their behavior: regions evolved from the embryonic rhombencephalon that experiment no significant volume change (−0.68 ± 0.21%, blue regions), olfactory bulb that shows a low level of volume reduction (−6.28%, green regions) and regions developed from prosencephalon, which show a higher reduction (−12.51 ± 0.36%, orange regions).
Validation of volumetric results
In order to validate the results from the volumetric analysis of the brain tissues and regions using the novel algorithms, we compared our results with those from Kovačević et al.48. The volumes reported for the WT group are coincidental with those presented by them. The small differences in absolute volume values may result from the fact that Kovačević and colleagues used inbred mice whereas we used a mixed C57/DBA background. Besides, Kovačević et al. analyzed images from ex vivo mouse brains while this work has dealt with in vivo brain images.
Discussion
Our results show two novel observations. First, consistent with studies in humans, we quantitatively confirm the existence of microcephaly and WM abnormalities in the brain of cbp+/− mice. We quantitatively confirm a drastic reduction (−16.31%) in brain volume associated to cbp haploinsufficiency, a finding in good agreement with the characteristic microcephaly associated to RSTS39. Our results further show that microcephaly is not only produced by GM hypoplasia, since a concomitant and proportional decrease (−16.00%) of WM volume was also found. Previous studies and case reports support this conclusion showing WM abnormalities25 and volume reductions26,31 in RSTS patients.
Second, our results suggest for the first time that CBP plays a differential role in the control of the development of prosencephalic regions, including the olfactory bulb and neocortex. Interestingly, in these areas tissue reductions kept a constant proportion, which suggests a role for CBP in brain size regulation. These results quantitatively confirm key syndromic features of RSTS, such as microcephaly, as well as anomalies in the WM, corpus callosum and cortex. In particular, our results show that the regions evolved from the embryonic prosencephalon are greatly reduced in cbp+/− subjects (−12.51 ± 0.36%). This could point to a mild form of holoprosencephaly (HPE, MIM 236100), which has been already linked with RSTS57,50. HPE is also associated with Dandy-Walker malformation (OMIM 220219)51, that has been also reported in RSTS patients29,34,35,37,52,53 and facial dysmorphism50, found in this RSTS mouse model13. The volumetric results in the prosencephalon drive two important observations. On the one hand, the volume of the cortex is greatly reduced in cbp+/− animals, pointing to the diagnosis of cortical atrophy. On the other hand, the olfactory bulb is also reduced (although less dramatically), what could be related to arhinencephaly, a diagnose that describes hypoplasia or absent olfactory tracts and bulb and that has been equally linked to HPE and RSTS57,50. Interestingly, HPE and arhinencephaly share an underlying etiology: they are both considered neuronal migration disorders (NMD). A perturbed neuronal migration mechanism could be the product of a failure in the neurogenesis or lack of differentiation of the progenitor cells to produce the three neural lineages (neurons, astrocytes and oligodendrocytes). In fact, such defects have been recently shown to occur in cbp haploinsufficiency during development14,54. Furthermore, CBP deficiency is also associated with a reduction of the number of newborn neurons in response to environmental enrichment in adult mice13.
Neuronal migration is a fundamental process in the development of the cortex, which in normal conditions is composed by six layers, created in an inside-out manner by means of subsequent waves of radially migrating neurons. NMDs are known to cause an abnormally thin cortex55 and cortical defects like (micro)lissencephaly, (poli)microgyria and pachigyria, observed in RSTS21. Furthemore, malgyration and microscopic abnormalities of the cytoarchytecture of the cerebral cortex35 along with “abnormal cortical infoldings”31 have also been reported in RSTS patients. NMD-caused microlissencephaly is also shown to contribute to microcephaly55, a feature often found in RSTS patients and reported here for the first time in cbp+/− mice.
More research in RSTS needs to be done to clarify the causes of this congenital disorder. Effective clinical diagnosis is fundamental for the early detection of RSTS. While there exist RSTS clinical guidelines3,4, there is a lack of established diagnostic criteria for the disease21 and the inclusion of neuroimaging studies in the clinical workflow has been proposed15,17,18,21,22,23,25,29,37,56. Our results provide strong evidence in support of neuroimaging-assisted diagnosis to reach early detection and treatment of RSTS-associated disorders. Furthermore, they offer neurobiological insight into the role of CBP in normal brain development and the consequences of its haploinsufficiency in RSTS models. The advance in the field will contribute to a better understanding of the disease and will help its diagnosis.
Additional Information
How to cite this article: Ateca-Cabarga, J. C. et al. Brain size regulations by cbp haploinsufficiency evaluated by in-vivo MRI based volumetry. Sci. Rep. 5, 16256; doi: 10.1038/srep16256 (2015).
References
Rubinstein, J. H. & Taybi, H. Broad thumbs and toes and facial abnormalities. A possible mental retardation syndrome. Am J Dis Child 105, 588–608 (1963).
Van Belzen, M., Bartsch, O., Lacombe, D., Peters, D. J. & Hennekam, R. C. Rubinstein-Taybi syndrome (CREBBP, EP300). Eur J Hum Genet. 19, preceeding 118–120 (2011).
Hennekam, R. C. Rubinstein-Taybi syndrome. Eur J Hum Genet. 14, 981–985 (2006).
Wiley, S., Swayne, S., Rubinstein, J. H., Lanphear, N. E. & Stevens, C. A. Rubinstein-Taybi syndrome medical guidelines. Am J Med Genet A. 119A, 101–110 (2003).
Michail, J., Matsoukas, J. & Theodorou, S. Pouce bot arqué en forte abduction-extension et autres symptomes concomitants. Rev Chir Orthop 43, 142–146 (1957).
Barco A. The Rubinstein-Taybi syndrome: modeling mental impairment in the mouse. Genes Brain Behav 6, 32–39 (2007).
Lopez-Atalaya, J. P., Valor, L. M. & Barco, A. Epigenetic factors in intellectual disability: the Rubinstein-Taybi syndrome as a paradigm of neurodevelopmental disorder with epigenetic origin. Prog Mol Biol Transl Sci. 128, 139–176 (2014).
Petrij, F., Giles, R. H., Dauwerse, H. G., Saris, J. J., Hennekam, R. C. M., Masuno, M., Tommerup, N., Van Ommen, G. J. B., Goodman, R. H., Peters, D. J. M. & Breuning, M. H. Rubinstein-Taybi syndrome caused by mutations in the transcriptional co-activator CBP. Nature 376, 348–351 (1995).
Zimmermann, N., Acosta, A. M., Kohlhase, J. & Bartsch, O. Confirmation of EP300 gene mutations as a rare cause of Rubinstein-Taybi syndrome. Eur J Hum Genet. 15, 837–842 (2007).
Bartholdi, D. et al. Genetic heterogeneity in Rubinstein-Taybi syndrome: delineation of the phenotype of the first patients carrying mutations in EP300. J Med Genet. 44, 327–333 (2007).
Roelfsema, J. H. et al. Genetic heterogeneity in Rubinstein-Taybi syndrome: mutations in both the CBP and EP300 genes cause disease. Am J Hum Genet. 76, 572–580 (2005).
Tanaka, Y., Naruse, I., Maekawa, T., Masuya, H., Shiroishi, T. & Ishii, S. Abnormal skeletal patterning in embryos lacking a single Cbp allele: a partial similarity with Rubinstein-Taybi syndrome. Proc Natl Acad Sci USA 94, 10215–10220 (1997).
López-Atalaya, J. P. et al. CBP is required for environmental enrichment-induced neurogenesis and cognitive enhancement. EMBO J 30, 4287–4298 (2011).
Wang, J. et al. CBP histone acetyltransferase activity regulates embryonic neural differentiation in the normal and Rubinstein-Taybi syndrome brain. Dev Cell. 18, 114–125 (2010).
Marzuillo, P. et al. Brain magnetic resonance in the routine management of Rubinstein-Taybi syndrome (RTS) can prevent life-threatening events and neurological deficits. Am J Med Genet A. 164A, 2129–2132 (2014).
de Kort, E., Conneman, N. & Diderich, K. A case of Rubinstein-Taybi syndrome and congenital neuroblastoma. Am J Med Genet A. 164A, 1332–1333 (2014).
Lee, J. S. et al. Clinical and mutational spectrum in Korean patients with Rubinstein-Taybi syndrome: the spectrum of brain MRI abnormalities. Brain Dev. 37, 402–408 (2015).
Marzuillo, P. et al. Novel cAMP binding protein-BP (CREBBP) mutation in a girl with Rubinstein-Taybi syndrome, GH deficiency, Arnold Chiari malformation and pituitary hypoplasia. BMC Med Genet. 14, 28 (2013). 10.1186/1471-2350-14-28.
Li, Z. et al. Phenotypic expansion of the interstitial 16p13.3 duplication: a case report and review of the literature. Gene. 531, 502–505 (2013).
Demeer, B. et al. Duplication 16p13.3 and the CREBBP gene: confirmation of the phenotype. Eur J Med Genet. 56, 26–31 (2013).
Kumar, S., Suthar, R., Panigrahi, I. & Marwaha, R. K. Rubinstein-Taybi syndrome: Clinical profile of 11 patients and review of literature. Indian J Hum Genet. 18, 161–166 (2012).
Giussani, C. et al. The association of neural axis and craniovertebral junction anomalies with scoliosis in Rubinstein-Taybi syndrome. Childs Nerv Syst. 28, 2163–2168 (2012).
Parsley, L., Bellus, G., Handler, M. & Tsai, A. C. Identical twin sisters with Rubinstein-Taybi syndrome associated with Chiari malformations and syrinx. Am J Med Genet A. 155A, 2766–2770 (2011).
Thienpont, B. et al. Duplications of the critical Rubinstein-Taybi deletion region on chromosome 16p13.3 cause a novel recognisable syndrome. J Med Genet. 47, 155–161 (2010).
Kim, S. H., Lim, B. C., Chae, J. H., Kim, K. J. & Hwang, Y. S. A case of Rubinstein-Taybi Syndrome with a CREB-binding protein gene mutation. Korean J Pediatr. 53, 718–721 (2010).
Wójcik, C. et al. Rubinstein-Taybi syndrome associated with Chiari type I malformation caused by a large 16p13.3 microdeletion: a contiguous gene syndrome? Am J Med Genet A. 152A, 479–483 (2010).
Wachter-Giner, T., Bieber, I., Warmuth-Metz, M., Bröcker, E. B. & Hamm, H. Multiple pilomatricomas and gliomatosis cerebri--a new association? Pediatr Dermatol. 26, 75–78 (2009).
Verstegen, M. J., van den Munckhof, P., Troost, D. & Bouma, G. J. Multiple meningiomas in a patient with Rubinstein-Taybi syndrome. Case report. J Neurosurg. 102, 167–168 (2005).
Agarwal, R., Aggarwal, R., Kabra, M. & Deorari, A. K. Dandy-Walker malformation in Rubinstein-Taybi syndrome: a rare association. Clin Dysmorphol. 11, 223–224 (2002).
Ihara, K., Kuromaru, R., Takemoto, M. & Hara, T. Rubinstein-Taybi syndrome: a girl with a history of neuroblastoma and premature thelarche. Am J Med Genet. 83, 365–366 (1999).
Sener, R. N. Rubinstein-Taybi syndrome: cranial MR imaging findings. Comput Med Imaging Graph 19, 417–418 (1995).
Robinson, T. W., Stewart, D. L. & Hersh, J. H. Monozygotic twins concordant for Rubinstein-Taybi syndrome and implications for genetic counseling. Am J Med Genet. 45, 671–673 (1993).
Guion-Almeida, M. L. & Richieri-Costa, A. Callosal agenesis, iris coloboma and megacolon in a Brazilian boy with Rubinstein-Taybi syndrome. Am J Med Genet. 43, 929–931 (1992).
Albanese, A. et al. [Role of diagnostic imaging in Rubinstein-Taybi syndrome. personal experience with 8 cases]. Radiol Med. 81, 253–261 (1991).
Rubinstein, J. H. Broad thumb-hallux (Rubinstein-Taybi) syndrome 1957-1988. Am J Med Genet Suppl. 6, 3–16 (1990).
Hennekam, R. C., Stevens, C. A. & Van de Kamp, J. J. Etiology and recurrence risk in Rubinstein-Taybi syndrome. Am J Med Genet Suppl. 6, 56–64 (1990).
Bonioli, E., Bellini, C. & Di Stefano, A. Unusual association: Dandy-Walker-like malformation in the Rubinstein-Taybi syndrome. Am J Med Genet. 33, 420–421 (1989).
Beluffi, G., Pazzaglia, U. E., Fiori, P., Pricca, P. & Poznanski, A. K. [Oto-palato-digital syndrome. Clinico-radiological study]. Radiol Med. 74, 176–184 (1987).
Cantani, A. & Gagliesi, D. Rubinstein-Taybi syndrome. Review of 732 cases and analysis of the typical traits. Eur Rev Med Pharmacol Sci. 2, 81–87 (1998).
Viosca, J., Lopez-Atalaya, J. P., Olivares, R., Eckner, R. & Barco, A. Syndromic features and mild cognitive impairment in mice with genetic reduction on p300 activity: Differential contribution of p300 and CBP to Rubinstein-Taybi syndrome etiology. Neurobiol Dis. 37, 186–194 (2010).
Martínez-Martínez, M. A., Pacheco-Torres, J., Borrell, V. & Canals, S. Phenotyping the central nervous system of the embryonic mouse by magnetic resonance microscopy. Neuroimage. 97, 95–106 (2014).
Heikkinen, T. et al. Characterization of neurophysiological and behavioral changes, MRI brain volumetry and 1H MRS in zQ175 knock-in mouse model of Huntington’s disease. PLoS One. 7, e50717 (2012), 10.1371/journal.pone.0050717.
Alarcón, J. M. et al. Chromatin acetylation, memory and LTP are impaired in CBP+/− mice: a model for the cognitive deficit in Rubinstein-Taybi syndrome and its amelioration. Neuron. 42, 947–959 (2004).
Smith, S. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage 23 Supp 1, S208–19 (2004).
Smith, S. M. Fast robust automated brain extraction. Hum Brain Mapp 17, 143–155 (2002).
Ashburner, J. & Friston, K. J. Unified segmentation. Neuroimage 26, 839–851 (2005).
Sawiak, S. J., Wood, N. I., Williams, G. B., Morton, A. J. & Carpenter, T. A. Voxel-based morphometry in the R6/2 transgenic mouse reveals differences between genotypes not seen with manual 2D morphometry. Neurobiol Dis 33, 20–27 (2009).
Kovačević, N. et al. A three-dimensional MRI atlas of the mouse brain with estimates of the average and variability. Cereb Cortex 15, 639–645 (2005).
Zacharoff, L. et al. Cortical metabolites as biomarkers in the R6/2 model of Huntington’s disease. J Cereb Blood Flow Metab. 32, 502–514 (2012).
Petryk, A., Graf, D. & Marcucio, R. Holoprosencephaly: signaling interactions between the brain and the face, the environment and the genes and the phenotypic variability in animal models and humans. Wiley Interdiscip Rev Dev Biol. 4, 17–32 (2015).
Solomon, B. D., Gropman, A. & Muenke, M. Holoprosencephaly Overview. In: GeneReviews (eds Pagon, R. A. et al.), Seattle (WA): University of Washington, Seattle; 1993-2014, 2000 Dec 27 [Updated 2013 Aug 29]. Available from: http://www.ncbi.nlm.nih.gov/books/NBK1530/ [Date of access: September 4, 2015].
Mazzone, D., Milana, A., Praticò, G. & Reitano, G. Rubinstein-Taybi syndrome associated with Dandy-Walker cyst. Case report in a newborn. J Perinat Med. 17, 381–384 (1989).
Barson, A. J. Proceedings: Rubinstein-Taybi syndrome. Arch Dis Child. 49, 495 (1974).
Tsui, D. et al. CBP regulates the differentiation of interneurons from ventral forebrain neural precursors during murine development. Dev Biol. 385, 230–241 (2014).
Ross, M. E. & Walsh, C. A. Human brain malformations and their lessons for neuronal migration. Annu Rev Neurosci. 24, 1041–1070 (2001).
Tanaka, T., Ling, B. C., Rubinstein, J. H. & Crone, K. R. Rubinstein-Taybi syndrome in children with tethered spinal cord. J Neurosurg. 105, 261–264 (2006).
Dubourg, C. et al. Holoprosencephaly. Orphanet J Rare Dis. 2, 2–8 (2007).
Acknowledgements
We are grateful to Begoña Fernández for her excellent technical assistance. We would like to thank S. Sawiak (Wolfson Imaging Centre, University of Cambridge, Cambridge, United Kingdom) for the mouse brain tissue probability maps and the SPMmouse plug-in and to N. Kovačević (Mouse Imaging Centre, Hospital for Sick Children, Toronto, Ontario, Canada) for the atlas of the mouse brain. Supported by grants from the Spanish MINECO to S.C. (BFU 2012–39958) and MINECO and FEDER to D.M. (TEC 2012–33778) and from MINECO (SAF2011-22855) and Generalitat Valenciana (Prometeo/2012/005) to A.B. The Instituto de Neurociencias is “Centre of Excellence Severo Ochoa”.
Author information
Authors and Affiliations
Contributions
J.L.-A., A.B., S.C. and D.M. contributed to the study concept and design. S.C. and D.M. supervised the studies described in the paper. J.C.A.-C. analyzed the data. A.C. and V.P. helped with data analysis. J.C.A.-C., J.L.-A., A.B., S.C. and D.M. wrote the paper. J.C.A.-C., A.C., S.C. and D.M. participated in the development of methodologies. J.C.A.-C., J.L.-A., A.B., S.C. and D.M. interpreted data. All authors discussed the results and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ateca-Cabarga, J., Cosa, A., Pallarés, V. et al. Brain size regulations by cbp haploinsufficiency evaluated by in-vivo MRI based volumetry. Sci Rep 5, 16256 (2015). https://doi.org/10.1038/srep16256
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep16256
This article is cited by
-
A novel CREBBP mutation and its phenotype in a case of Rubinstein–Taybi syndrome
BMC Medical Genomics (2022)
-
The emerging role of chromatin remodelers in neurodevelopmental disorders: a developmental perspective
Cellular and Molecular Life Sciences (2021)
-
The transcriptional coactivator and histone acetyltransferase CBP regulates neural precursor cell development and migration
Acta Neuropathologica Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.