Abstract
The sequencing of the zebrafish genome should be completed by the end of 2002. Direct assignment of function on the basis of this information would be facilitated by the development of a rapid, targeted ‘knockdown’ technology in this model vertebrate. We show here that antisense, morpholino-modified oligonucleotides1 (morpholinos) are effective and specific translational inhibitors in zebrafish. We generated phenocopies of mutations of the genes no tail (ref. 2), chordin (ref. 3), one-eyed-pinhead (ref. 4), nacre (ref. 5) and sparse (ref. 6), removing gene function from maternal through post-segmentation and organogenesis developmental stages. We blocked expression from a ubiquitous green fluorescent protein (GFP) transgene, showing that, unlike tissue-restricted limitations found with RNA-based interference in the nematode7, all zebrafish cells readily respond to this technique. We also developed also morpholino-based zebrafish models of human disease. Morpholinos targeted to the uroporphyrinogen decarboxylase gene8 result in embryos with hepatoerythropoietic porphyria. We also used morpholinos for the determination of new gene functions. We showed that embryos with reduced sonic hedgehog (ref. 9) signalling and reduced tiggy-winkle hedgehog (ref. 10) function exhibit partial cyclopia and other specific midline abnormalities, providing a zebrafish genetic model for the common human disorder holoprosencephaly. Conserved vertebrate processes and diseases are now amenable to a systematic, in vivo, reverse-genetic paradigm using zebrafish embryos.
Similar content being viewed by others
Main
Morpholinos are chemically modified oligonucleotides with base-stacking abilities similar to those of natural genetic material11. Morpholinos have been shown to bind to and block translation of mRNA in vitro1,11, in tissue culture cells1,11 and, recently, in specific and restricted applications in vivo12,13,14. For example, morpholinos targeting β-catenin can block translation through early developmental stages of the Xenopus laevis embryo, but are apparently ineffective at the end of gastrulation and beyond14. Morpholinos function through an RNase-H–independent mechanism by hindering translational initiation1,11. This approach makes in vivo targeting highly predictable and reduces non-specific effects. For example, we show here that all nine genes studied were functionally depleted using the first morpholino designed against the leader sequence of each transcript.
We selected a non-essential ubiquitous GFP transgene (E-line; Fig. 1e,f) to test for the applicability of antisense morpholinos as a general strategy for gene ‘knockdown’ in zebrafish. Microinjection is a very effective delivery system for morpholinos in zebrafish (Fig. 1c,d). We injected a GFP-targeted morpholino (GFP-MO) and efficiently reduced GFP transgene expression (Fig. 1k,l), as determined using fluorescence and western-blot assays. The level of GFP depletion resulted in some embryos with no visible GFP fluorescence, indicating a nearly complete loss of GFP protein (Fig. 1k,l). From these studies on the GFP transgene, we demonstrated the specific depletion of the targeted protein in all cells of the 28-hour zebrafish embryo using morpholinos.
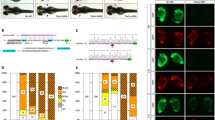
Uninjected, sphere-stage (a) and 28-h (b) embryos are shown under FITC illumination. Injection resulted in an even distribution of fluorescein-tagged control morpholinos, as assayed using FITC; sphere (c) and 28-h (d) stages are shown. e–q, GFP expression inhibition in 28-h-stage E-line embryos by injected GFP antisense morpholino (GFP-MO). All panels show 28-h embryos under FITC illumination. e,f, GFP fluorescence in uninjected E-line embryos is ubiquitous. g,h, E-line embryos injected with 4.5 ng of control morpholino. i,j, E-line embryos injected with 4.5 ng of a 4-base mismatch GFP antisense morpholino (GFPD4-MO). Note that fluorescence is near control levels in these embryos. k,l, E-line embryos injected with 4.5 ng of GFP morpholino. GFP expression is inhibited in all cells. m,n, Wild-type embryos. o, GFP fluorescence inhibition graph. Blue, data from control-MO–injected embryos; red, GFPD4-MO–injected embryos; black, GFP-MO–injected embryos. Translational inhibition induced by GFP-MO is sequence-specific and dose-dependent. We analysed 15 embryos each from independent experiments for 4.5-ng and 9-ng data points, and 15 embryos for each other data point shown. p, Detection of GFP protein using western-blot analysis in GFP-MO injected embryos. Protein from 28-h stage E-line embryos injected with the indicated amounts of GFP-MO were assayed as described. Note the dose-dependent loss of GFP protein in these embryos. q, Translational inhibition by GFP-MO occurs through a post-transcriptional mechanism. Sibling embryos to those used in (p) were analysed for GFP transcripts using a northern blot. Scale bars, 0.2 mm.
We also analysed several well-characterized genetic loci. We injected a chordin antisense morpholino (chordin-MO) and observed a highly specific series of phenotypes dependent on dose (Fig. 2). Embryos injected with higher doses of chordin-MO are phenocopies of chordin null-mutant embryos15,16 (Fig. 2c). At low doses, the equivalent of a reduced chordin loss-of-function phenotype was seen, with embryos displaying only a partially expanded blood island, u-shaped somites and an abnormal tail fin with multiple folds (Fig. 2b). In a single experiment, we saw the equivalent of an allelic series for the loss of chordin by the use of different doses of the chordin morpholino.
a–h,w, chordin morpholino (chd-MO) injection data. i–l, one-eyed pinhead morpholino (oep-MO) injection data. m,o,r,t, no tail morpholino (ntl-MO) injection data. n–q,s–v, oep-MO and ntl-MO co-injection data. a,d,f,k,n,s, Wild-type embryos. a–c,i,j,m, 28-h-old embryos. r, Three-day-old embryos. b,c,e,g,h, Embryos injected with 4.5 ng of chd-MO. b, Weak/moderate chd-MO injection phenotype. Abnormal u-shaped somites, abnormal tail fin with multiple folds and expanded blood island (filled arrowhead) are observed. c, Strong chd-MO injection phenotype, observed at high frequency in high-dose injections (≥75% at 4.5-ng injection, n=423). Note abnormal u-shaped somites, extremely expanded blood island (filled arrowhead), abnormal tail fin and reduced head (open arrow). w, chd phenotype frequency and strength depends on the injected chd-MO dose. Black, weak/moderate phenotype frequency; blue, strong phenotype frequency. Embryos were examined as a function of dose: 0.09 ng, n=169; 0.9 ng, n=97; 1.5 ng, n=399; 4.5 ng, n=423; 9 ng, n=224. d,e, In situ hybridization analysis of otx-2 in 80% epiboly embryos. Reduction of otx-2 in (e) is consistent with head reduction in (c) (otx-2 reduction frequency is 78%, n=39). f,g, In situ hybridization analysis of gata-2 in a 65% epiboly embryo. The gata-2 expression domain is enlarged and shifted anteriorly (filled arrows) in (g), indicating ventralization of the mesoderm (gata-2 is expanded in 100% of the injected embryos, n=30). We tested the ability of exogenously-supplied chordin function to reverse the effects of chordin-MO to check for the specificity of gene targeting by chordin-MO. Synthetic Xenopus chordin mRNA (ref. 30) rescued the phenotypic consequences of the chordin-MO (76% showed strong chordin phenotype with 4.5 ng chordin-MO (n=423); 10% displayed the strong chordin phenotype when injected with 4.5 ng chordin-MO and 300 pg Xchordin mRNA (n=30)). i,j,l, Embryos injected with 9 ng oep-MO. i, Phenocopy of zygotic oep mutant. Cyclopia and ventral curvature are observed. The phenotype frequency is 30±5% (n=291). j, Phenocopy of maternal-zygotic oep mutant. Severe cyclopia, somite absence in the trunk, misshapen tail somites and reduced notochord are observed. The phenotype frequency is 13±5% (n=291). These effects are rescued by the injection of synthetic oep mRNA (43% of embryos displayed oep phenotypes upon injection of 9 ng oep-MO (n=291) and 0% of embryos displayed oep phenotypes on injection with 9 ng oep-MO and 50 pg oep mRNA (n=33)). k,l, In situ hybridization analysis of pax2.1 (asterisk) and axial in tailbud-stage embryos. Note the prechordal mesoderm reduction in the injected embryo (l) compared with wild type (k) (filled arrowheads; frequency 45%, n=24). m,r, Embryos injected with 9 ng of ntl-MO. ntl-MO injection phenocopies a ntl null mutation with 98% frequency (n=118). Normal head, abnormal somites and reduced tail are prominent. As a further test of specificity, ntl-MO was injected into ntl mutant embryos and their normal siblings. No additional defects were noted in the ntl mutant embryos due to the injection of ntl-MO (n=72, data not shown). n–q,s–v, oep-MO and ntl-MO interaction. n,s, Uninjected embryos. o,t, Embryos injected with 9 ng ntl-MO. p,u, Embryos injected with 9 ng oep-MO. q,v, Embryos co-injected with 9 ng ntl-MO and 9 ng oep-MO. n–q, In situ hybridization analysis of myod in 10–12-somite embryos. Note adaxial mesoderm reduction and posterior somite fusion in ntl-MO–injected embryo (92% frequency, n=25; o). oep-MO–injected embryos (p) have posterior fusion of the adaxial mesoderm (frequency is 41%, n=22). ntl-MO and oep-MO co-injected embryos (q) show the absence of adaxial mesoderm and reduced somitic mesoderm, with 52% frequency (n=24). s–v, Whole-mount in situ hybridization for shh in 10–12-somite stage embryos. t, ntl-MO injected embryos show a weak expansion of shh expression (80%, n=25). u, oep-MO injection results in a strong expansion of the shh expression domain (50%, n=22). v, ntl- and oep-MO co-injection results in a severe reduction in shh expression in 52% of the co-injected embryos (n=31). x, Western-blot analysis of 28-h-stage E-line embryos injected with the indicated amounts of ntl-MO. Ntl protein is depleted at 4.5 ng and 9 ng doses and is severely reduced at 3 ng dose. The lower signal from uninjected embryos is due to reduced protein load in this lane, as determined by Ponceau S staining of the membrane (data not shown). Scale bars, 0.2 mm.
We selected the one-eyed pinhead gene4 (oep) to test for depletion of maternal gene activity. Embryos deficient in oep function are defective in signalling through the nodal pathway, and embryonic oep function is due to both maternal and zygotic genetic contributions that are distinguishable on the basis of specific criteria17. We injected oep-MO and noted phenotypes consistent with loss of zygotic oep function (Fig. 2i,l,p,u). Injections with higher concentrations of oep-MO resulted in a phenocopy of the loss of both zygotic and maternal oep function (Fig. 2j). Morpholinos are thus capable of targeting maternal gene function, albeit at reduced levels compared with zygotic gene targeting.
We chose no tail (ntl; ref. 2) to identify genetic interactors using morpholinos. We injected ntl-MO and obtained embryos indistinguishable from those caused by a null mutation18, using molecular (Fig. 2o,t) and phenotypic (Fig. 2m,r) criteria. Ntl protein is specifically and quantitatively reduced in embryos injected with ntl-MO (Fig. 2x). Activation of the somitic mesodermal marker myod requires input from both oep and ntl pathways19. Embryos reduced in either ntl (Fig. 2o) or oep (Fig. 2p) function display an altered but robust expression of myod; embryos reduced in both functions express myod in only a few cells (Fig. 2q). Morpholinos are thus effective tools for the testing of genetic interactions in vivo.
The oep-MO illustrates limitations of this technology in zebrafish. One-half of oep-MO–injected embryos failed to elicit the oep loss-of-function phenotype. In addition, embryos from a wild-type strain in Japan fail to respond to oep-MO, but they do respond to chordin-MO and ntl-MO (M. Mieda, pers. comm.). Polymorphism in 5′ UTR sequences at the oep locus might explain these non-responding embryos, a hypothesis we are currently testing.
Morpholino hybridization specificity can sometimes be relaxed at higher concentrations. Injection of more than 9 ng of oep-MO will result in the specific loss of eye development due to the apparent targeting of a second gene. A morpholino designed to target the bozozok (boz)/dharma/nieuwkoid (refs 20,21) locus results in a hypomorphic boz (ref. 20) phenocopy at low doses, whereas a neural degeneration phenotype is also noted at concentrations above 4.5 ng (data not shown). Central nervous system degeneration was a common phenotypic classification observed in chemically induced mutant screens22, and consequently we expect this to be one major class of phenotype observed due to lax morpholino targeting. Conclusive determination of any phenotype is made through either the targeting of the same gene by a second, non-overlapping sequence oligo or RNA rescue. Reduced-specificity targeting is not common and was not noted in high-concentration injections for any of the remaining five genes.
We studied later-acting genes to determine the persistence of morpholino effects. We injected nacre-MO and noted a characteristic and nearly complete loss of body pigmentation through the first 50 hours of development (Fig. 3b,d), a phenotype indistinguishable from that observed in the nacre mutant5. At later time points, pigmentation returns at a variable rate (data not shown). Injection of sparse-MO also duplicated a known zebrafish pigment mutation, sparse (ref. 6). Dorsal melanocytes were reduced in these embryos at 65 hours (Fig. 3f) and at 10 days (Fig. 3h) of development. Morpholino-based gene targeting is thus completely penetrant throughout the first two days of development, which include the critical vertebrate processes of somitogenesis and organogenesis in the zebrafish embryo.
a–d, nacre-MO injection phenotype. Note extreme pigment reduction in 2-day embryo injected with 9 ng nacre-MO (98% penetrance; n=112; b,d) compared with wild type (a,c). e–h, sparse phenocopy of 65-hour (e,f) and 10-day (g,h) embryos. f,h, Embryos injected with 9 ng sparse-MO. Note the reduced number of pigmented cells (melanocytes) in 95% of these fish (n=159); sparse is required for melanocyte survival throughout the period from 2 to 10 days of embryonic development6. i–l, Morpholino-based modelling of HEP. Morpholino-based inhibition of urod results in embryos with fluorescent (j) and photosensitive (l) blood cells (100%; n=118). Embryos in all panels were exposed to light before analysis. i, Sibling control morpholino-injected embryo, analysed using a rhodamine filter set. j, Embryo injected with urod-MO, analysed using a rhodamine filter set. Note the intense auto-fluorescence from the likely accumulation of photosensitive porphyrins in the circulating blood cells. k,l, Photosensitivity of urod-MO–injected blood cells. Exposure to light effectively depletes red blood cells in urod-MO–injected embryos (l), as visibly assayed analysing the circulatory system in the heart and surrounding blood vessels after embryonic exposure to light. k, Sibling, control-injected embryo. Arrows in (k) and (l) mark an area of concentration of visible red blood cells.
We have begun the systematic development of morpholino-based models of human disease. Hepatoerythropoietic porphyria (HEP) is caused by a defect in haem biosynthesis through the loss of the uroporphyrinogen decarboxylase (encoded by urod) enzyme23. The manifestations of this syndrome include fluorescent and photosensitive red blood cells, and embryos injected with urod-MO display both phenotypes (Fig. 3j,l), as noted in a hypomorphic urod mutation8. The complete phenotypic penetrance of embryos injected with urod-MO demonstrates the usefulness of the morpholino-based animal models of human disease for applications such as drug development and testing.
Holoprosencephaly (HPE) occurs at high frequency in the human embryo (1:250) and in live births (1:16,000; ref. 24), and in extreme cases the phenotype is cyclopia. The gene sonic hedgehog (shh) is thought to have a critical role in the development of this disease in humans25,26. Zebrafish shh mutations, however, result in no anterior midline signalling defects27. A second shh orthologue expressed in the anterior midline, tiggy-winkle hedgehog (twhh; ref. 10), may explain this lack of phenotype in a highly conserved vertebrate developmental process. We targeted both shh and twhh using morpholinos to test for redundancy and to develop zebrafish as a genetic model for HPE. Embryos injected with shh-MO display phenotypes characteristic of a shh mutation27: ‘u’-shaped somites, lack of the horizontal myoseptum (Fig. 4g) and small pectoral fins (Fig. 4c). Embryos injected with twhh-MO are indistinguishable from controls (Fig. 4b,f). Injection of both twhh-MO and shh-MO, however, resulted in embryos with synergistic defects in somitic patterning in the trunk (Fig. 4m), a new phenotype in the forebrain, partial cyclopia (Fig. 4d), and loss of transcription of the hedgehog target gene, patched (ptc; Fig. 4j; ref. 28). Recent confirmation of these results in embryos deficient for sonic hedgehog indicates a 100% penetrance (n=52) of partial cyclopia due to injection of 9 ng twhh-MO; twhh-MO failed to cause any cyclopia in sibling embryos (0%, n=120; S. Bingham and A. Chandrasekhar, pers. comm.). We conclude that zebrafish embryos contain two functionally redundant orthologues of mammalian SHH with similar roles in anterior midline patterning. This result directly validates the zebrafish as an effective model for the role of sonic hedgehog signalling in HPE. Finally, these experiments demonstrate the usefulness of morpholinos for both single and multiple gene knockdowns for the understanding of vertebrate embryonic development and disease.
Evidence for redundancy in function between two zebrafish hedgehog orthologues, shh and twhh, in midline signalling. a,e,i, Wild-type embryos. b,f, Embryos injected with 18 ng twhh-MO and 9 ng control-MOs. c,g,o, Embryos injected with 18 ng shh-MO and 9 ng control-MO. d,h,j,p, Embryos co-injected with shh-MO and twhh-MO (13.5 ng each). k,q, Embryo injected with shh-MO and twhh-MO (13.5 ng each) and twhh mRNA (100 pg). a–h, Three-day-old embryos. i–k, Dorsal view of two-day embryos analysed for the expression of the hedgehog target gene, ptc. j, Note the strong reduction in ptc expression (n=47) in the midline (arrow) and lens (open arrowhead). Expression of ptc in the pectoral fin is variably reduced (filled arrowhead) in these embryos. In this example, expression is only noted in one fin bud, consistent with the observed asymmetrical defects in fin bud development at the phenotypic level for shh-MO–injected embryos (data not shown) and for shh-mutant embryos27. k, ptc expression is restored in embryos injected with twhh-MO and shh-MO by the addition of twhh mRNA (n=37). l, Loss of ptc expression due to the injection of twhh-MO and shh-MO is reversed in both head and fin bud. b, twhh-MO–injected embryos have a normal head, ‘v’-shaped somites and normal myoseptum (f; open arrow) compared with wild-type embryos (a,e). c, shh-MO–injected embryos have a normal head, somites are ‘u’ shaped and myoseptum is absent (g; open arrow). Also, reduced finbuds are observed in some shh-MO–injected embryos (c; open arrowhead) compared with wild-type or twhh-MO–injected embryos (b; open arrowhead). In shh-MO and twhh-MO co-injected embryos, partial cyclopia (d) and absent myoseptum (h; white arrow) is observed in most injected embryos. n–q, Embryos at the 10-somite stage stained for myod. shh-MO–injected embryos show a weak reduction of adaxial and somitic mesoderm (o; 42%, n=12) compared with a wild-type embryo (n). Shh-MO and twhh-MO co-injected embryos show a reduction of adaxial mesoderm (p; 94%, n=17). This reduction is reversed and adaxial mesoderm is lightly expanded by synthetic zebrafish twhh mRNA (q; 0% reduced, 59% expanded, n=17), confirming the specificity of targeting to the twhh locus using twhh-MO. m, Frequency of the observed phenotypes due to shh-MO and twhh-MO injections: black, cyclopia; blue, ‘u’-shaped somites; red, reduced finbuds. We scored 25 embryos injected with the indicated morpholino combination from at least two independent experiments to generate each data point. Anterior is to the left. Scale bars, 0.2 mm.
Methods
Morpholinos.
Morpholinos were obtained from Gene Tools, LLC. Morpholinos work through an RNase-H independent process that blocks translational initiation1. Consequently, all morpholinos were arbitrarily designed to bind to the 5′ UTR or sequences flanking and including the initiating methionine. We selected sequences based on design parameters according to the manufacturer's recommendations, namely 21–25mer antisense oligonucleotides of ∼50% G/C and A/T content with no predicted internal hairpins. Four consecutive G nucleotides were also avoided. In addition, we now test each design sequence for representation elsewhere in the genome. The significance of this prescreening will greatly increase once the zebrafish genome is completely sequenced. Sequences were as follows (sequence complimentary to the predicted start codon is underlined in all cases): chordin-MO, 5′–ATCCACAGCAGCCCCTCCATCATCC–3′ (fluorescein tagged at the 3′ end); GFP-MO, 5′–TCTTCTCCTTTACTCATTTTCTACC–3′; GFPD4-MO, 5′–TCTACTCGTTTACTCATTATCTtCC–3′; negative control MO sequence (untagged or fluorescein tagged at the 3′ end), 5′–CCTCTTACCTCAGTTACAATTTATA–3′; oep-MO, 5′–GCCAATAAACTCCAAAACAACTCGA–3′; ntl-MO, 5′–GACTTGAGGCAGGCATATTTCCGAT–3′; shh-MO, 5′–CAGCACTCTCGTCAAAAGCCGCATT–3′; twhh-MO, 5′–TTCCATGACGTTTGAATTATCTCTT–3′; nacre-MO, 5′–CATGTTCAACTATGTGTTAGCTTCA–3′; sparse-MO, 5′–TATAAGTCCATCTATCTCATGTGTG–3′; urod-MO, 5′–GAATGAAACTGTCCTTATCCATCA–3′.
Zebrafish GFP strain E-line.
The zebrafish E-line transgenic line contains a single copy of the pT-EF1α-GFP-pA transposon at the E-line locus (D. Mohn and P. Hackett, pers. comm.). We obtained heterozygous E-line embryos from an outcross of homozygous E-line adults.
Morpholino injections.
Morpholino oligonucleotides were solubilized in water at the concentration of 8 mM (∼65 mg/ml). The resulting stock solution was diluted to working concentrations of 0.09–3 mg/ml in water or 1×Danieau solution (58 mM NaCl, 0.7 mM KCl, 0.4 mM MgSO4, 0.6 mM Ca (NO3)2, 5 mM HEPES, pH 7.6) before injection into the yolk as described10. The resulting buffered Danieau MO solution injections caused lower mortality rate in the injected embryos compared with water MO solution injections, but did not affect the penetrance of the observed phenotypes (data not shown). The injected volume was 1.5–15 nl, depending on the required injected MO dose. Wild-type or heterozygous E-line embryos of 1–16-cell stages were injected into the yolk. For morpholino distribution analysis, fluorescein-labelled control or fluorescein-labelled chordin-MO (90 pg) was injected into wild-type embryos from 1–16-cell stages of development and analysed using FITC filters on a Zeiss Axioplan2 fluorescence microscope. At this concentration, chordin-MO–injected embryos developed normally. Images were obtained using a Kodak DCS420 Digital Camera.
Effective doses were determined separately for each morpholino. For example, at the effective dose of 4.5 ng and lower, ≥90% of the GFP-MO–injected embryos developed normally as assayed using standard morphological criteria. Higher doses of the GFP-MO resulted in a larger average reduction of GFP protein, but also caused some detectable detrimental effects on development (data not shown). These higher-dose effects were not pursued further for this morpholino. In all cases shown, the dose used for analysis resulted in embryos of two classes, those displaying a specific phenotype or those that were normal using morphological criteria. A small fraction (typically ≤5%) of embryos developed abnormally due to mechanical damage following microinjection.
Penetrance numbers using morphological criteria were independently verified in each example using a specific concentration based on an initial dose response assessment. A minimum sample size of 25 was used in each case.
mRNA injections.
Synthetic mRNA was injected into the yolk of embryos previously injected with the indicated morpholino as described10. Siblings from the same pool of morpholino-injected embryos served as the internal control for these experiments. The sequence of the oep mRNA did not contain any overlap with the oep-MO. Similarly, the twhh mRNA used contained only a six-base overlap with twhh-MO, a level of overlap previously shown to be insufficient for morpholino targeting in vitro and in tissue culture studies1,11. We used chordin mRNA from X. laevis to avoid any sequence homology with the zebrafish chordin-MO and because previous studies showed X. laevis and zebrafish chordin genes encode equivalent specific activities in X. laevis embryos29.
GFP fluorescence analysis.
We divided 15 randomly selected embryos into groups of 3 for FITC analysis, resulting in the determination of 5 values for each experimental data point. Fluorescence pictures of the same exposure were taken of each group using a MICROIMAGE I30B Low Light Integrating Camera and captured using a DC30+ analogue to digital video capture board (Pinnacle Systems) at maximum resolution (640×480) settings. Signal intensities were set to sub-saturation levels (determined using uninjected E-line embryos) for maximal information capture, and fidelity of capture was confirmed by simultaneously imaging both analogue and digital video shots. The resulting images were imported into Adobe Photoshop 5.0 for quantitative analysis. The background for each image was established using the ‘selective colour’ algorithm with the same settings in all images. The ‘mean’ value of the ‘histogram’ algorithm was used to measure the green channel signal (the other channels were removed to minimize non-specific background). All scores were normalized to the values obtained from uninjected E-line (100%) and wild-type (0%) data points. The specific loss of GFP signal using 4.5 ng GFP-MO was noted in 9 separate experiments, with at least 30 embryos assayed in each experiment.
Western-blot analysis.
We carried out standard western-blot analyses using anti-GFP (Clontech) or anti-Ntl antibodies on protein isolated from pools of injected embryos. The protein equivalent from five embryos was loaded per lane.
Northern-blot analysis.
Northern-blot assay was performed according to standard procedures. A 700-bp fragment from a PstI digest corresponding to the GFP-coding region was used as probe. Each lane represents 5 μg total RNA isolated from a pool of 30 embryos. Two independent analyses were performed.
References
Summerton, J. Morpholino antisense oligomers: the case for an RNase H-independent structural type. Biochim. Biophys. Acta 1489, 141–158 (1999).
Schulte-Merker, S., van Eeden, F.J., Halpern, M.E., Kimmel, C.B. & Nusslein-Volhard, C. no tail (ntl) is the zebrafish homologue of the mouse T (Brachyury) gene. Development 120, 1009–1015 (1994).
Schulte-Merker, S., Lee, K.J., McMahon, A.P. & Hammerschmidt, M. The zebrafish organizer requires chordino. Nature 387, 862–863 (1997).
Zhang, J., Talbot, W.S. & Schier, A.F. Positional cloning identifies zebrafish one-eyed pinhead as a permissive EGF-related ligand required during gastrulation. Cell 92, 241–251 (1998).
Lister, J.A., Robertson, C.P., Lepage, T., Johnson, S.L. & Raible, D.W. nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 126, 3757–3767 (1999).
Parichy, D.M., Rawls, J.F., Pratt, S.J., Whitfield, T.T. & Johnson, S.L. Zebrafish sparse corresponds to an orthologue of c-kit and is required for the morphogenesis of a subpopulation of melanocytes, but is not essential for hematopoiesis or primordial germ cell development. Development 126, 3425–3436 (1999).
Tavernarakis, N., Wang, S.L., Dorovkov, M., Ryazanov, A. & Driscoll, M. Heritable and inducible genetic interference by double-stranded RNA encoded by transgenes. Nature Genet. 24, 180–183 (2000).
Wang, H., Long, Q., Marty, S.D., Sassa, S. & Lin, S. A zebrafish model for hepatoerythropoietic porphyria. Nature Genet. 20, 239–243 (1998).
Krauss, S., Concordet, J.P. & Ingham, P.W. A functionally conserved homolog of the Drosophila segment polarity gene hh is expressed in tissues with polarizing activity in zebrafish embryos. Cell 75, 1431–1444 (1993).
Ekker, S.C. et al. Patterning activities of vertebrate hedgehog proteins in the developing eye and brain. Curr. Biol. 5, 944–955 (1995).
Summerton, J. & Weller, D. Morpholino antisense oligomers: design, preparation, and properties. Antisense Nucleic Acid Drug Dev. 7, 187–195 (1997).
Arora, V. et al. c-Myc antisense limits rat liver regeneration and indicates role for c- Myc in regulating cytochrome P-450 3A activity. J. Pharmacol. Exp. Ther. 292, 921–928 (2000).
Qin, G., Taylor, M., Ning, Y.Y., Iversen, P. & Kobzik, L. In vivo evaluation of a morpholino antisense oligomer directed against tumor necrosis factor-α. Antisense Nucleic Acid Drug Dev. 10, 11–16 (2000).
Heasman, J., Kofron, M. & Wylie, C. β-catenin signaling activity dissected in the early Xenopus embryo: a novel antisense approach. Dev. Biol. 222, 124–134 (2000).
Fisher, S., Amacher, S.L. & Halpern, M.E. Loss of cerebrum function ventralizes the zebrafish embryo. Development 124, 1301–1311 (1997).
Hammerschmidt, M. et al. dino and mercedes, two genes regulating dorsal development in the zebrafish embryo. Development 123, 95–102 (1996).
Gritsman, K. et al. The EGF-CFC protein one-eyed pinhead is essential for nodal signaling. Cell 97, 121–132 (1999).
Halpern, M.E. et al. Genetic interactions in zebrafish midline development. Dev. Biol. 187, 154–170 (1997).
Schier, A.F., Neuhauss, S.C., Helde, K.A., Talbot, W.S. & Driever, W. The one-eyed pinhead gene functions in mesoderm and endoderm formation in zebrafish and interacts with no tail. Development 124, 327–342 (1997).
Fekany, K. et al. The zebrafish bozozok locus encodes Dharma, a homeodomain protein essential for induction of gastrula organizer and dorsoanterior embryonic structures. Development 126, 1427–1438 (1999).
Koos, D.S. & Ho, R.K. The nieuwkoid/dharma homeobox gene is essential for bmp2b repression in the zebrafish pregastrula. Dev. Biol. 215, 190–207 (1999).
Haffter, P. et al. The identification of genes with unique and essential functions in the development of the zebrafish, Danio rerio. Development 123, 1–36 (1996).
Kappas, A., Sassa, S., Galbraith, R.A. & Nordmann, Y. The Metabolic Basis of Inherited Diseases (eds Scriver, C.R., Beaudet, A.L. & Sly, W.S.) 2103–2159 (McGraw-Hill, New York, 1995).
Wallis, D.E. & Muenke, M. Molecular mechanisms of holoprosencephaly. Mol. Genet. Metab. 68, 126–138 (1999).
Belloni, E. et al. Identification of Sonic hedgehog as a candidate gene responsible for holoprosencephaly. Nature Genet. 14, 353–356 (1996).
Roessler, E. et al. Mutations in the human Sonic Hedgehog gene cause holoprosencephaly. Nature Genet. 14, 357–360 (1996).
Schauerte, H.E. et al. Sonic hedgehog is not required for the induction of medial floor plate cells in the zebrafish. Development 125, 2983–2993 (1998).
Concordet, J.P. et al. Spatial regulation of a zebrafish patched homologue reflects the roles of sonic hedgehog and protein kinase A in neural tube and somite patterning. Development 122, 2835–2846 (1996).
Miller-Bertoglio, V., Fisher, S., Sanchez, A., Mullins, M. & Halpern, M.E. Differential regulation of chordin expression domains in zebrafish. Dev. Biol. 192, 537–550 (1998).
Sasai, Y. et al. Xenopus chordin: a novel dorsalizing factor activated by organizer-specific homeobox genes. Cell 79, 779–790 (1994).
Acknowledgements
We thank S. Bingham, A. Chandrasekhar and M. Mieda for communication of results before publication; D. Mohn and P. Hackett for the E-line strain; A. Davidson, K. Finley and K. Markley for homozygous E-line fish; M. Halpern for the oep template DNA; A. Davidson, A. Kattan, K. Finley and C. Saunders for independently corroborating this method; P. Ingham for the ptc probe; S. Schulte-Merker for anti-Ntl antibody; J. Larson for technical support; M. Kofron, J. Heasman, J. Summerton and P. Morcos for sharing ideas on the possible uses of morpholinos as antisense inhibitors; and the zebrafish community for their suggestions and encouragement.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nasevicius, A., Ekker, S. Effective targeted gene ‘knockdown’ in zebrafish. Nat Genet 26, 216–220 (2000). https://doi.org/10.1038/79951
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/79951
This article is cited by
-
Exploring hematopoiesis in zebrafish using forward genetic screening
Experimental & Molecular Medicine (2024)
-
Zebrafish cardiac repolarization does not functionally depend on the expression of the hERG1b-like transcript
Pflügers Archiv - European Journal of Physiology (2024)
-
Modelling human lower urinary tract malformations in zebrafish
Molecular and Cellular Pediatrics (2023)
-
Molecular Characterization of U6 Promoters from Orange-Spotted Grouper (Epinephelus coioides) and Its Application in DNA Vector-Based RNAi Technology
Marine Biotechnology (2023)
-
Unlocking the Potential of Zebrafish Research with Artificial Intelligence: Advancements in Tracking, Processing, and Visualization
Medical & Biological Engineering & Computing (2023)