Abstract
The completion of the human genome project has enabled researchers to characterize the breakpoints for various chromosomal structural abnormalities including deletions, duplications or translocations. This in turn has shed new light on the molecular mechanisms underlying the onset of gross chromosomal rearrangements. On the other hand, advances in genetic manipulation technologies for various model organisms has increased our knowledge of meiotic chromosome segregation, errors which, contribute to chromosomal aneuploidy. This review focuses on the current understanding of germ line chromosomal abnormalities and provides an overview of the mechanisms involved. We refer to our own recent data and those of others to illustrate some of the new paradigms that have arisen in this field. We also discuss some perspectives on the sexual dimorphism of some of the pathways that leads to these chromosomal abnormalities.
Similar content being viewed by others
The random nature of gross chromosomal rearrangements
The development of chromosomal structural abnormalities, also known as gross chromosomal rearrangements (GCR), is essentially dependent on two distinct processes: double-strand breaks (DSB) and DSB repair. DSBs can result from exogenous agents such as ionizing radiation and chemotherapeutic drugs, and also from endogenously generated reactive oxygen species and mechanical stresses on the chromosomes.1 DSBs can impair cellular function and eventually cause cell death by triggering apoptosis.2 To counteract such deleterious effects, DSBs are often repaired by using error-free systems; that is, homologous recombination (HR). However, GCRs are occasionally the result of an illegitimate error-prone repair system, non-homologous end joining or single-strand annealing. Moreover, the experimental induction of only two DSBs is sufficient to produce GCRs in mammalian cells.3 DSBs that occur on the same chromosome produce deletions or inversions, whereas those on different chromosomes yield translocations.
In general, DSBs occur randomly, and thus GCR is fundamentally a random and non-recurrent event. However, some unusual situations can arise that modulate either of the two DSB-related events, induction and repair and occasionally trigger recurrent GCRs. Genomic sequence analyses of the break point region of such recurrent GCRs have revealed characteristic genomic architectures including direct or inverted repeats and the critical break points often reside within these regions. When a DSB occurs within one copy of a repeat segment, illegitimate DSB repair through the HR pathway using another copy of the repeat segments may be induced, and this is the most likely mechanism of recurrent GCR. To date, a substantial number of disease-related GCRs initiated by this mechanism have been identified, and are thus referred to as ‘genomic disorders’.4
Non-random GCR: mediation of deletion and duplication events by low-copy repeats
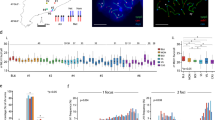
One of the best-studied examples of this group of disease is Charcot-Marie-Tooth disease type 1A (CMT1A), which is an inherited peripheral neuropathy that results from duplication of dosage-sensitive gene, PMP22. A common 1.4 Mb duplication of genomic segment including PMP22 gene at chromosome 17p11.2 is found in majority of patients with CMT1A. Both proximal and distal end points of the duplication are located within 24 kb low-copy repeats (LCR) that have almost an identical sequence (>98%).5 Thus, HR between the LCRs is the likely mechanism of the duplication. On the other hand, hereditary neuropathy with liability to pressure palsies (HNPP) is another type of hereditary neuropathy caused by a deletion of 17p11.2. Interestingly, the deleted segment in HNPP is identical to the duplicated region in CMT1A and the deletion end points are located within the same LCRs (Figure 1a).6
LCR-mediated deletion and duplication. (a) Example of two disorders with deletion or duplication resulting from reciprocal exchange. Black boxes indicate LCRs at 17p11.2. A 1.4 Mb genomic segment (white) is deleted in HNPP patients or duplicated in CMT1A cases. (b) Deletion mapping of 243 cases with DS22. LCRs are indicated by white boxes. Approximately 90% of the cases carry a 3 Mb standard deletion spanning LCR22-A and LCR22-D. Other minority cases also carry deletions between each LCR. TBX1, the gene responsible for the phenotype, is located between LCR22-A and LCR22-B. (c) Possible mechanism of LCR-mediated deletion and duplication. Two homologous chromosomes (paternal and maternal) are indicated by bold and dotted lines, respectively. In normal meiotic recombination, because crossover is placed on the equivalent position for each chromosome, the copy numbers of each chromosomal region do not change (left). However, the highly homologous nature of LCRs gives rise to mispairing of the homologous chromosomes, which induces NAHR (middle). An alternative hypothesis is that intrachromosomal recombination is followed by a looping out of the intervening region (right).
Similarly, 22q11 deletion syndrome (DS22) is another example of genomic disorder characterized by a combination of congenital cardiac defect, dysmorphic face, mental retardation, thymic defect and hypoparathyroidism. Initially considerable phenotypic variability of DS22 invokes the idea of contiguous gene deletion syndrome: the variation of deleted region contributes to variation of the phenotype. However, deletion mapping revealed that the deleted region of 90% of cases was almost identical regardless of their phenotypic difference.7 Genomic sequence of this region was complicated and challenging, because more than seven copies of LCRs clustered at 22q11.8 Finally, DS22 critical region were found to contain four copies of the LCRs, two of which contributed to generation of the 3 Mb standard deletion. Although the remaining cases with DS22 represent minor type of deletion, all deletion end points are located within the four LCRs (Figure 1b). Later, mental retardation patients with 22q11 duplication that is reciprocal to the 22q11 deletion were reported, suggesting a common mechanism that is shared with CMT and HNPP.9
Discoveries of these LCRs have opened up a consecutive identification of similar syndromes caused by recurrent deletion or duplication where LCRs are invariantly located at both the end points. A growing number of examples include Prader–Willi syndrome/Angelman syndrome (15q11–13), Williams–Beuren syndrome (7q11), spinal muscular atrophy (5q12–13), neurofibromatosis type 1 (NF1) (17q11), Smith–Magenis syndrome (17p11) and Sotos syndrome (5q35).10, 11, 12, 13, 14, 15, 16
DS22 arises at a frequency of 1 in 3000 live births including a high incidence of de novo cases. Indeed, PCR detects frequent de novo deletions in sperm from healthy donors.16 This high recurrent rate invokes an idea of involvement of a physiological event, meiotic recombination, in generation of deletion/duplication. In meiosis I, programmed induction of DSBs by SPO11 endonuclease is followed by homologous chromosome pairing and subsequent error-free DSB-repair by HR. A subset of HR results in a crossover of homologous chromosomes that contributes to exchange in genetic materials and diversity of gametes (Figure 1c, left). However, LCR flanking the DSB site might induce mispairing to another copy of LCR with high homology, resulting in non-allelic homologous recombination (NAHR) (Figure 1c, middle). The possible outcome is a deletion on one allele and a duplication on the other.4 Another pathway that can potentially produce LCR-mediated deletion is an intrachromosomal NAHR between the LCRs on the same chromosome (Figure 1c, right). In this case, segment between the two LCRs would be looped out, generating only a deletion.17, 18 The LCRs involved in CMT1A or DS22 manifest recombination frequency of average levels, suggestive of no DSB or recombination hotspot activity of the LCRs.19, 20 Thus, recurrent GCRs of this type are unlikely to result from non-random DSBs, but from non-random DSB repair through illegitimate NAHR.
Pelizaeus–Merzbacher disease is one of the hereditary neurodegenerative disorders resulting from deletion/duplication of the chromosomal region including dosage-sensitive PLP1 gene on Xq22. In contrast to other genomic disorders, the analysis of break points revealed no LCR-like genomic structures at either ends of deletions or duplications. Recently, sequence refractory fragments that preclude the progression of DNA polymerase have been consistently identified at these rearrangement junctions.21 This observation suggests that a unique mechanism related to DNA replication-stalling that might cause non-random DSB formation and might potentially be another source of recurrent GCRs.
Non-random GCR: palindrome-mediated translocation
Somatic translocations are often identified in association with certain tumors and leukemias. A translocation that occurs in a single cell should normally be harmless, even if it disrupts an important gene. However, when a translocation accidentally activates the oncogenic process through the generation of a chimeric oncogene, the cell can acquire a growth advantage that causes unrestricted clonal expansion. Even in recurrent tumor-related translocations that appear to be cytogenetically identical, such as the well-known t(9;22)(q34;q11) observed in chronic myeloblastic leukemia, the translocation break points have been shown to be non-identical when examined at a molecular level.22 In other words, although they often involve the same genes, their molecular location within those genes varies considerably. However, a subset of oncogenic translocations does arise in a non-random fashion. These distinct break points are the sites of translocation often associated with lymphoblastic leukemias/lymphomas that are often found at immunoglobulin or T-cell receptor gene loci. As these genes normally undergo somatic rearrangement to achieve diversity of the encoded protein, errors in this physiological DSB and DSB-repair process may be involved in the generation of translocation events.23
Similarly, most constitutional translocations are random events associated with non-recurrent break points. However, rare recurrent constitutional translocations have also been revealed, and are reminiscent of a specific genomic structure, such as the LCR. The Robertsonian translocation is one of the best known non-random constitutional translocations and is characterized by a fusion of the centromeres of two acrocentric chromosomes. Because the centromeric region is composed of long stretches of specific repeat sequences, HR events between these repeats may be involved in generation of this translocation.24 In support of this hypothesis is the high frequency of de novo Robertsonian translocations in the germ line, which is akin to the LCR-mediated deletion/duplication. The short arms of acrocentric chromosomes are rich in ribosomal genes associated with the nucleolar organizer region in interphase nuclei, and this might also affect the high incidence of the translocations.25
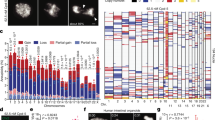
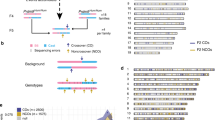
The t(11;22)(q23;q11) is another example of a recurrent constitutional translocation in humans. Although balanced carriers of the t(11;22) usually manifest no clinical symptoms, translocation carriers often have problems with reproduction, such as male infertility, recurrent spontaneous abortions and the birth of offspring with a chromosomal imbalance known as Emanuel syndrome (supernumerary der(22)t(11;22) syndrome). We have previously identified palindromic AT-rich repeats (PATRRs) at both break point regions on 11q23 and 22q11.26, 27, 28 The PATRRs on 11q23 and 22q11 (PATRR11, PATRR22) are 450 and 590 bp in length, respectively, with a high AT content of more than 90%, constituting a nearly perfect palindromic structure with 98% identity between its proximal and distal arms (Figure 2a). However, in spite of their similarity with regard to AT-richness, no substantial homology was observed between PATRR11 and PATRR22 (58% identity), suggesting that HR is not responsible for this recurrent translocation. A more detailed characterization of the junction involved further revealed remnants of DSBs at the center of the two PATRRs, followed by repair through the non-homologous end joining pathway.29, 30, 31, 32
Palindrome-mediated translocations. (a) Predicted secondary structure for palindromic sequences. Short palindromic sequences have the potential to form double-stranded cruciform structures by intrastrand base pairing in single-stranded DNA. DNA sequences indicated by dotted white arrows are complementary to those depicted by the white-dotted black arrows. (b) Cruciform extrusion of the plasmid harboring the PATRR11 region. The plasmid bearing the PATRR11 insert was fixed using psoralen treatment followed by ultraviolet exposure. The PATRR11 fragments were then released with restriction enzyme digestion and were visualized using atomic force microscopy. (c) Translocation-specific PCR. Arrows indicate each arm of the PATRR11 (solid arrows) and the PATRR22 (hatched arrows) regions. Translocations can be detected using one of the primers flanking PATRR11 and with one of the primers flanking PATRR22. PCR primers for the detection of der(11) are also indicated (triangles). Centromeres are represented as circles. (d) Detection of de novo translocation. Genomic DNA was extracted from sperm samples and subjected to translocation-specific PCR by using multiple batches of template DNA. The gel images show the PCR results for sperm DNA (upper panel) and lymphoblast DNA (lower panel). Lane N, negative control; lane P, genomic DNA from a t(11;22) balanced carrier serving as a positive control.
Two unrelated constitutional t(17;22)(q11;q11) translocations associated with NF1 disease have been reported.33, 34 These two translocations disrupt the NF1 gene on 17q11, which produces the NF1 phenotype in the affected patients. Further analysis of the break points revealed that the translocation occurred between PATRR22 and another ∼200bp PATRR within intron 31 of the NF1 gene.34, 35, 36 Subsequent molecular cloning of other translocation break points has demonstrated similar palindromic, often AT-rich sequences on partner chromosomes, such as 4q35.1, 1p21.2 and 8q24.1.37, 38, 39 Hence, this subset of translocations seems to occur in a non-random fashion possibly mediated by the DSB susceptibility of palindromic DNA.
Small palindromic sequences such as the PATRRs have the potential to form stem–loop structures by intrastrand base pairing within single-stranded DNA. As a consequence, they form a specific secondary structure, either a single-stranded hairpin or a double-stranded cruciform (Figures 2a and b).40 It is not yet certain whether such cruciform DNA actually exists in living cells. However, this phenomenon is indirectly evidenced by the fact that the polymorphisms within the PATRRs affect the susceptibility to translocation. Longer and more symmetric PATRR alleles with potent cruciform-forming propensity in vitro show a strong predisposition to translocation events.41, 42 It is proposed that the secondary structures adopted by palindromic DNA induce a greater susceptibility to DSBs thus leading to recurrent chromosomal translocations in humans.26 This hypothesis has been recently verified using a plasmid-based model system that recapitulates PATRR-mediated translocation.43 Such secondary structure-induced GCR has also been evidenced by the presence of DNA segments that could potentially adopt triplex DNA conformations at the break points of the recurrent t(14;18)(q32;q21) translocation in follicular lymphoma.44
In a manner that is analogous to the Robertsonian translocation, the intranuclear localization of the associated break point regions might also affect the recurrent nature of the t(11;22) translocation. Recent data have supported an emerging consensus that chromosomes are compartmentalized into discrete territories.45 In B-cell lymphoma, a prevailing hypothesis is that the spatial proximity of the two break point loci affects the translocation-susceptibility levels.46 With regards to t(11;22), the break point region at 11q23 is located closer to the other breakpoint at 22q11 than to other loci, which might facilitate illegitimate repair leading to translocation.47
The dimorphic features of GCR
We established a t(11;22)-specific PCR system that utilizes sequence data for the region around the PATRR. PATRR-flanking primers were designed both on 11q23 and 22q11 to amplify the junction fragments of the der(11) or the der(22) (Figure 2c). We thereby successfully amplified the der(11) and der(22) junction fragments of balanced t(11;22) carriers as well as the der(22) of patients with Emanuel syndrome.29 Translocation-specific PCR was then performed using conditions that would allow for the detection of a single molecule of the target DNA. When DNA derived from sperm samples obtained from healthy male volunteers was amplified using our assay, we obtained both positive and negative PCR results, indicating that we had successfully detected translocation products that were de novo in origin (Figure 2d).48
Surprisingly, translocation-specific PCR has never detected the de novo occurrence of translocation in any tissues other than sperm.48 Diverse human tissues such as peripheral leukocytes and skin fibroblasts have been found to be consistently negative using a PCR assay for translocation. We also tested various long-term cultured somatic cell lines derived from human cells, but all proved to be negative. The fact that only sperm samples produce de novo t(11;22)s leads us to speculate that PATRR-mediated translocations occur primarily during meiosis. These findings are thus quite unusual and seem to be inconsistent with established mechanisms regarding the instability of palindromic DNA sequences that have been described for many experimental organisms.49, 50 Such genomic instability seems to be primarily mediated by stalling of the DNA replication fork at a region that forms a hairpin structure.51 However, the apparently sperm-specific de novo occurrence of PATRR-mediated translocations may suggest a novel paradigm regarding palindrome instability that is independent of DNA replication. The frequency of de novo t(11;22) translocations in sperm has been found to be independent of the age of the donors.52 If the translocation events occur during DNA replication, the samples from older donors should include a greater number of translocations as they have undergone a greater number of cell divisions (Figure 3a). Thus, these findings lend support to the possibility of replication-independency of this phenomenon.
To further evaluate the possible existence of such a meiosis-specific translocation mechanism, it must be determined whether female germ cells can also produce de novo t(11;22) events. However, translocation-specific PCR is not feasible for this purpose because the number of human oocytes that can be examined is limited. As an alternative approach, we collected samples from individuals who had undergone a de novo t(11;22) and also their parents to determine the origin of the translocation by assessing the sequence variation within the PATRRs. By segregation analysis, we have so far found that the de novo events are exclusively of paternal origin in our patient population although the number of samples examined thus far is small (unpublished data). This finding implies that it is not necessarily meiosis, but a spermatogenesis-specific mechanism that permits the development of the t(11;22) translocation. DNA breakage might occur during late spermatogenesis when it is packaged into dense chromatin. A plausible explanation is also that during the transition of the histones to protamine, their release from nucleosomes may have a role in the release of free-negative supercoiling and thus facilitate cruciform extrusions at the PATRR.40, 53
Whereas 90% of numerical chromosomal abnormalities in the germ line are derived from maternal errors in meiosis, 80% of the structural chromosomal abnormalities are of paternal origin.54, 55 This is not surprising because it is quite likely that the relatively higher number of cell divisions that occurs during spermatogenesis compared with oogenesis (Figure 3a) increases the probability of DSB formation, whereas a lack of efficient DNA repair in late spermatogenesis may also contribute to this disparity. However, our current hypothesis points to the importance of post-meiotic chromatin dynamism and changes in the DNA topology during spermatogenesis for the generation of translocations. This has important implications for the novel hypothesis that de novo GCRs in humans are predominantly of paternal origin.
In sharp contrast, Robertsonian translocations preferentially occur during female meiosis.56 This clearly reflects the difference in the development of the Robertsonian translocation, in which meiotic recombination might be involved. Consistent with this is the observation that LCR-mediated deletions are predominantly derived from errors in female meiosis.57 The higher recombination rate found in female meioses compared with their male counterparts may also be relevant to these observations.58
Genetic susceptibility to germ line chromosomal aneuploidy
Other types of chromosomal aberrations include numerical abnormalities such as trisomy or monosomy, also known as aneuploidy. In the conceptus or fetus this occurs in at least 5% of all pregnancies and is a common reproductive problem in humans.59 Some fetuses with aneuploidy survive to term but suffer from disorders associated with congenital anomalies and mental retardation such as Down syndrome with trisomy 21. However, most conceptuses of autosomal aneuploidy die in utero, resulting in early pregnancy loss.
Most germ line aneuploidy occurs as a consequence of the non-disjunction of homologous chromosomes during meiosis I. A considerable body of evidence obtained from the analysis of genetically manipulated model organisms has been accumulated in relation to how the meiotic machinery is involved in non-disjunction.60, 61 There is an emerging consensus that the events that occur during prophase in meiosis I are essential for the proper segregation of homologous chromosomes.62 Homologous chromosomes that behave independently during mitotic division have to segregate into two different daughter cells in meiosis I. To accomplish this process, homologous chromosomes interact with each other utilizing a specialized HR pathway (Figure 4a). Initially, chromosomal DNA undergoes programmed cleavage through SPO11 endonuclease at more than 100 sites dispersed throughout the entire genome. To repair these DSBs correctly, a subsequent HR pathway is activated and the broken DNA ends begin looking for homologous regions with the aid of RAD51 and RAD52. As a consequence, two homologous chromosomes are brought together in close association, known as homolog-pairing. A proteinous structure known as the synaptonemal complex is then formed between the paired homologous chromosomes and the DNA lesions are subsequently repaired through HR. At the final step in HR, a four-stranded DNA structure, the Holliday junction that physically links two chromosomes is resolved in one of two ways, crossover or non-crossover. Crossover maintains the physical linkage of the chromosomes (chiasmata) and produces the appropriate bi-orient tension at the opposite spindle poles during metaphase in meiosis I. This then transmits signals to the checkpoint machinery allowing cells to enter anaphase (Figure 4b). Thus, the number and location of the crossover events are strictly regulated (crossover assurance and interference). Meiotic recombination, which is well known as a mechanism that shuffles genetic material to produce variation among individuals, is also indispensable for the proper segregation of homologous chromosomes.
Mechanisms that drive the segregation of homologous chromosomes in meiosis I. (a) Schematic representation of meiotic recombination. Initially, programmed DSBs are induced by SPO11 endonuclease (pink circles). The 5′-ends of the DSBs are then resected and 3′-protruding single-stranded ends are generated. With the aid of RAD51 and RAD52, the DNA ends produce nucleofilament complexes (blue circles), which facilitate genome-wide homology scanning to find homologous chromosome (red lines). Next, the single-stranded DNA end invades the homologous duplex DNA and forms a D-loop structure. DNA synthesis seals the DSBs and double Holliday junctions emerge. These Holliday junctions are resolved one of two ways, crossover (right) or non-crossover (left), with Holliday junction resolvase (green wedges). (b) Three critical steps that affect chromosomal segregation in meiosis I. In pre-meiotic S-phase, both maternal (red) and paternal (green) chromosomes are replicated and are tightly connected with cohesin complexes (purple circles). After the induction of DSBs, two homologous chromosomes undergo pairing and a synaptonemal complex (pink ladder) is established between them. Successive DSB-repair by HR results in the production of at least one obligatory chiasmata by crossover. Afterwards, oogenesis enters a period of long meiotic arrest until meiosis I restarts at puberty. Just before ovulation, two homologous chromosomes are pulled in opposite directions to two spindle poles (gray). At this moment, two homologous chromosomes are connected only by a cohesin complex distal to the chiasmata. A full colour version of this figure is available at the Journal of Human Genetics journal online.
It is not unreasonable to hypothesize that genetic defects in the meiotic machinery in humans induce a greater susceptibility to non-disjunction and the generation of an aneuploid conceptus.63 There has been an anecdotal consensus that a woman with a previous aneuploid conception has an increased recurrence risk for the same or a different aneuploidy.64, 65 Recently, we identified mutations in the SYCP3 gene in two women with a recurrent pregnancy loss at between 6 and 10 weeks, possibly due to a repeated aneuploidy.66 The SYCP3 protein is an essential component of this synaptonemal complex and promotes the connection of homologous chromosomes during prophase in meiosis I (Figure 4b).67 Both identified mutations in this gene are heterozygous, possibly affecting the function of the wild-type SYCP3 protein through a dominant-negative effect. The phenotypes of the women were similar to that of female mice deficient in SYCP3, some offspring of which are affected by aneuploidy and succumb in utero during embryonic development.68 As defects of other meiotic genes are reported to induce aneuploidy in oocytes in female mice, it is possible that these genes also contribute to the susceptibility to aneuploidy in humans.69, 70
In relation to SYCP3, it is of interest that a mutation in this gene was also identified in two human patients with azoospermia.71 Indeed, male mice that are deficient in SYCP3 are sterile due to the onset of meiotic arrest.72 Three stages in prophase of meiosis I are believed to be critical for the proper segregation of meiotic chromosomes including (1) cohesion between sister chromosomes, (2) synapsis between homologous chromosomes and (3) the location and frequency of meiotic recombination. Notably, the phenotype of mice deficient in these genes often differs between male and female mice.73 Whereas male mice always manifest complete infertility as a result of meiotic arrest, female mice often show aneuploidy in oocytes during meiosis. This might be primarily due to differences in checkpoint robustness between female and male mice. Males may have a more stringent checkpoint during prophase of meiosis I that prevents aneuploidy in gametogenesis. Thus, the extra chromosomes in trisomic fetuses or conceptuses predominantly originate from non-disjunction events in female meiosis I.55, 59 Furthermore, chromosome imbalances in offspring, in which either of the parents carries chromosomal structural abnormalities such as translocations or inversions, often originate from maternal gametes, whereas such chromosomal abnormalities occasionally cause azoospermia due to meiotic arrest when they are present in male mice.74 As these structural abnormalities obviously impair complete synapsis during meiotic prophase, this phenotypic disparity may also be accounted by the checkpoint differences between female and male mice.
Age-related chromosomal non-disjunction in meiosis I
In human trisomy 21, three general characteristics are generally acknowledged: (1) in most cases the extra chromosome 21 is of maternal origin; (2) most cases are derived from a non-disjunction event in meiosis I; and (3) the frequency of these errors increases with maternal age.63 The basis for the age-dependent increase in non-disjunction in meiosis I has been a long-standing enigma in this field. The first specific evidence related to this issue was the observation that oocytes from older mice display a decrease in number of chiasmata, which might predispose the germ line cells to non-disjunction in meiosis I.75 Studies using polymorphic markers have provided supportive findings showing that the number of recombination events in chromosome 21 is reduced in human trisomy 21 cases.76, 77 The ‘production line hypothesis’ has also been entertained, that is, there is a gradient in the fetal ovary such that the first-formed oocytes have a higher frequency of chiasmata than those formed at a later stage, and that the oocytes ovulate in the same order in which they enter meiosis.75
Extensive studies using polymorphic markers have additionally revealed that the sites of recombination have a distal location bias in cases of trisomy 21.78 More detailed analyses have further demonstrated that distal recombination is a risk only in younger mothers, but that recombination pattern is not atypical in trisomy 21 offspring of an older mother.79 In females, the oocytes enter meiosis during the fetal period and their development arrests in the middle of prophase of meiosis I (dictyate stage) until they restart maturation just prior to ovulation at the reproductive age (Figure 3b). Thus, meiosis I is a decade-long process that likely puts some unusual stress on the segregation apparatus during this vulnerable period. The fact that even chromosomes with typical patterns of recombination undergo non-disjunction in older women implies that oocytes that were originally normal can acquire compromised chiasmatic functions that are common to distally located recombinations and could lead to the same outcome.79
Recent findings in mice that are deficient in SMC1B, one of the meiosis-specific components of the cohesin complex, have shed light on this proposition. In the female mutant mice, although the pattern of recombination locations detected at the MLH1 foci was found to be similar to wild-type mice, the chiasmata were predominantly located at the distal region and the number of bivalents in meiosis I was decreased in older mutant mice. This suggests that defects in cohesin instigates a slippage of the chiasmata towards the ends of chromosome during long meiosis I in older female mice.80 These findings may support the hypothesis that the degradation of the cohesin complex contributes to reduced number of chiasmata that will lead to an age-dependent increase of non-disjunction events in meiosis I. Although the mode of turnover for the cohesin complex has remained elusive to date, it is very possible that this will be revealed in the near future.
Conclusions
Elucidation of the mechanisms that govern the generation of chromosomal abnormalities will allow evidence-based risk assessments for the next generation. Combined with the recent advances in pre-implantation genetic diagnoses that can be an option for individuals at risk, this will provide a valuable source of information for clients receiving genetic counseling.
References
Khanna, K. K. & Jackson, S. P. DNA double-strand breaks: signaling, repair and the cancer connection. Nat. Genet. 27, 247–254 (2001).
van Gent, D. C., Hoeijmakers, J. H. & Kanaar, R. Chromosomal stability and the DNA double-stranded break connection. Nat. Rev. Genet. 2, 196–206 (2001).
Richardson, C. & Jasin, M. Frequent chromosomal translocations induced by DNA double-strand breaks. Nature 405, 697–700 (2000).
Lupski, J. R. Genomic disorders: structural features of the genome can lead to DNA rearrangements and human disease traits. Trends Genet. 14, 417–422 (1998).
Pentao, L., Wise, C. A., Chinault, A. C., Patel, P. I. & Lupski, J. R. Charcot-Marie-Tooth type 1A duplication appears to arise from recombination at repeat sequences flanking the 1.5 Mb monomer unit. Nat. Genet. 2, 292–300 (1992).
Chance, P. F., Alderson, M. K., Leppig, K. A., Lensch, M. W., Matsunami, N., Smith, B. et al. DNA deletion associated with hereditary neuropathy with liability to pressure palsies. Cell 72, 143–151 (1993).
Shaikh, T. H., Kurahashi, H., Saitta, S. C., O'Hare, A. M., Hu, P., Roe, B. A. et al. Chromosome 22-specific low copy repeats and the 22q11.2 deletion syndrome: genomic organization and deletion endpoint analysis. Hum. Mol. Genet. 9, 489–501 (2000).
Dunham, I., Shimizu, N., Roe, B. A., Chissoe, S., Hunt, A. R., Collins, J. E. et al. The DNA sequence of human chromosome 22. Nature 402, 489–495 (1999).
Edelmann, L., Pandita, R. K., Spiteri, E., Funke, B., Goldberg, R., Palanisamy, N. et al. A common molecular basis for rearrangement disorders on chromosome 22q11. Hum. Mol. Genet. 8, 1157–1167 (1999).
Amos-Landgraf, J. M., Ji, Y., Gottlieb, W., Depinet, T., Wandstrat, A. E., Cassidy, S. B. et al. Chromosome breakage in the Prader-Willi and Angelman syndromes involves recombination between large, transcribed repeats at proximal and distal breakpoints. Am. J. Hum. Genet. 65, 370–386 (1999).
Valero, M. C., de Luis, O., Cruces, J. & Pérez-Jurado, L. A. Fine-scale comparative mapping of the human 7q11.23 region and the orthologous region on mouse chromosome 5G: the low-copy repeats that flank the Williams-Beuren syndrome deletion arose at breakpoint sites of an evolutionary inversion(s). Genomics 69, 1–13 (2000).
Thompson, T. G., DiDonato, C. J., Simard, L. R., Ingraham, S. E., Burghes, A. H., Crawford, T. O. et al. A novel cDNA detects homozygous microdeletions in greater than 50% of type I spinal muscular atrophy patients. Nat. Genet. 9, 56–62 (1995).
Dorschner, M. O., Sybert, V. P., Weaver, M., Pletcher, B. A. & Stephens, K. NF1 microdeletion breakpoints are clustered at flanking repetitive sequences. Hum. Mol. Genet. 9, 35–46 (2000).
Potocki, L., Chen, K. S., Park, S. S., Osterholm, D. E., Withers, M. A., Kimonis, V. et al. Molecular mechanism for duplication 17p11.2- the homologous recombination reciprocal of the Smith-Magenis microdeletion. Nat. Genet. 24, 84–87 (2000).
Visser, R., Shimokawa, O., Harada, N., Kinoshita, A., Ohta, T., Niikawa, N. et al. Identification of a 3.0-kb major recombination hotspot in patients with Sotos syndrome who carry a common 1.9-Mb microdeletion. Am. J. Hum. Genet. 76, 52–67 (2005).
Turner, D. J., Miretti, M., Rajan, D., Fiegler, H., Carter, N. P., Blayney, M. L. et al. Germline rates of de novo meiotic deletions and duplications causing several genomic disorders. Nat. Genet. 40, 90–95 (2008).
Emanuel, B. S. & Shaikh, T. H. Segmental duplications: an ‘expanding’ role in genomic instability and disease. Nat. Rev. Genet. 2, 791–800 (2001).
Bailey, J. A. & Eichler, E. E. Primate segmental duplications: crucibles of evolution, diversity and disease. Nat. Rev. Genet. 7, 552–564 (2006).
Saitta, S. C., Harris, S. E., Gaeth, A. P., Driscoll, D. A., McDonald-McGinn, D. M., Maisenbacher, M. K. et al. Aberrant interchromosomal exchanges are the predominant cause of the 22q11.2 deletion. Hum. Mol. Genet. 13, 417–428 (2004).
Han, L. L., Keller, M. P., Navidi, W., Chance, P. F. & Arnheim, N. Unequal exchange at the Charcot-Marie-Tooth disease type 1A recombination hot-spot is not elevated above the genome average rate. Hum. Mol. Genet. 9, 1881–1889 (2000).
Lee, J. A., Carvalho, C. M. & Lupski, J. R. A DNA replication mechanism for generating nonrecurrent rearrangements associated with genomic disorders. Cell 131, 1235–1247 (2007).
Heisterkamp, N., Knoppel, E. & Groffen, J. The first BCR gene intron contains breakpoints in Philadelphia chromosome positive leukemia. Nucleic Acids Res. 16, 10069–10081 (1988).
Tsai, A. G., Lu, H., Raghavan, S. C., Muschen, M., Hsieh, C. L. & Lieber, M. R. Human chromosomal translocations at CpG sites and a theoretical basis for their lineage and stage specificity. Cell 135, 1130–1142 (2008).
Takahashi, Y., Fujita, H. & Nakamura, Y. & Kurahashi. H. Dual-color fish analysis of breakpoints on Robertsonian translocations. Jpn. J. Hum. Genet. 42, 517–523 (1997).
Kim, S. R. & Shaffer, L. G. Robertsonian translocations: mechanisms of formation, aneuploidy, and uniparental disomy and diagnostic considerations. Genet. Test 6, 163–168 (2002).
Kurahashi, H., Shaikh, T. H., Hu, P., Roe, B. A., Emanuel, B. S. & Budarf, M. L. Regions of genomic instability on 22q11 and 11q23 as the etiology for the recurrent constitutional t(11;22). Hum. Mol. Genet. 9, 1665–1670 (2000).
Kurahashi, H. & Emanuel, B. S. Long AT-rich palindromes and the constitutional t(11;22) breakpoint. Hum. Mol. Genet. 10, 2605–2617 (2001).
Kurahashi, H., Inagaki, H., Hosoba, E., Kato, T., Ohye, T., Kogo, H. et al. Molecular cloning of a translocation breakpoint hotspot in 22q11. Genome Res. 17, 461–469 (2007).
Kurahashi, H., Shaikh, T. H., Zackai, E. H., Celle, L., Driscoll, D. A., Budarf, M. L. et al. Tightly clustered 11q23 and 22q11 breakpoints permit PCR-based detection of the recurrent constitutional t(11;22). Am. J. Hum. Genet. 67, 763–768 (2000).
Edelmann, L., Spiteri, E., Koren, K., Pulijaal, V., Bialer, M. G., Shanske, A. et al. AT-rich palindromes mediate the constitutional t(11;22) translocation. Am. J. Hum. Genet. 68, 1–13 (2001).
Tapia-Paez, I., Kost-Alimova, M., Hu, P., Roe, B. A., Blennow, E., Fedorova, L. et al. The position of t(11;22)(q23;q11) constitutional translocation breakpoint is conserved among its carriers. Hum. Genet. 109, 167–177 (2001).
Kato, T., Inagaki, H., Kogo, H., Ohye, T., Yamada, K., Emanuel, B. S. et al. Two different forms of palindrome resolution in the human genome: deletion or translocation. Hum. Mol. Genet. 17, 1184–1191 (2008).
Kehrer-Sawatzki, H., Haussler, J., Krone, W., Bode, H., Jenne, D. E., Mehnert, K. U. et al. The second case of a t(17;22) in a family with neurofibromatosis type 1: sequence analysis of the breakpoint regions. Hum. Genet. 99, 237–247 (1997).
Kurahashi, H., Shaikh, T., Takata, M., Toda, T. & Emanuel, B. S. The constitutional t(17;22): another translocation mediated by palindromic AT-rich repeats. Am. J. Hum. Genet. 72, 733–738 (2003).
Inagaki, H., Ohye, T., Kogo, H., Yamada, K., Kowa, H., Shaikh, T. H. et al. A palindromic AT-rich repeat in the NF1 gene is hypervariable in humans and evolutionarily conserved among primates. Hum. Mutat. 26, 332–342 (2005).
Lewis, S. M., Chen, S., Strathern, J. N. & Rattray, A. J. New approaches to the analysis of palindromic sequences from the human genome: evolution and polymorphism of an intronic site at the NF1 locus. Nucleic Acids Res. 33, e186 (2005).
Nimmakayalu, M. A., Gotter, A. L., Shaikh, T. H. & Emanuel, B. S. A novel sequence-based approach to localize translocation breakpoints identifies the molecular basis of a t(4;22). Hum. Mol. Genet. 12, 2817–2825 (2003).
Gotter, A. L., Shaikh, T. H., Budarf, M. L., Rhodes, C. H. & Emanuel, B. S. A palindrome-mediated mechanism distinguishes translocations involving LCR-B of chromosome 22q11.2. Hum. Mol. Genet. 13, 103–115 (2004).
Gotter, A. L., Nimmakayalu, M. A., Jalali, G. R., Hacker, A. M., Vorstman, J. & Conforto-Duffy, D. et al. A palindrome-driven complex rearrangement of 22q11.2 and 8q24.1 elucidated using novel technologies. Genome Res. 17, 470–481 (2007).
Kurahashi, H., Inagaki, H., Yamada, K., Ohye, T., Taniguchi, M., Emanuel, B. S. et al. Cruciform DNA structure underlies the etiology for palindrome-mediated human chromosomal translocations. J. Biol. Chem. 279, 35377–35383 (2004).
Kato, T., Inagaki, H., Yamada, K., Kogo, H., Ohye, T., Kowa, H. et al. Genetic variation affects de novo translocation frequency. Science 311, 971 (2006).
Kogo, H., Inagaki, H., Ohye, T., Kato, T., Emanuel, B. S. & Kurahashi, H. Cruciform extrusion propensity of human translocation-mediating palindromic AT-rich repeats. Nucleic Acids Res. 35, 1198–1208 (2007).
Inagaki, H., Ohye, T., Kogo, H., Kato, T., Bolor, H., Taniguchi, M. et al. Chromosomal instability mediated by non-B DNA: Cruciform conformation and not DNA sequence is responsible for recurrent translocation in humans. Genome Res. 19, 191–198 (2009).
Raghavan, S. C., Swanson, P. C., Wu, X., Hsieh, C. L. & Lieber, M. R. A non-B-DNA structure at the Bcl-2 major breakpoint region is cleaved by the RAG complex. Nature 428, 88–93 (2004).
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2, 292–301 (2001).
Roix, J. J., McQueen, P. G., Munson, P. J., Parada, L. A. & Misteli, T. Spatial proximity of translocation-prone gene loci in human lymphomas. Nat. Genet. 34, 287–291 (2003).
Ashley, T., Gaeth, A. P., Inagaki, H., Seftel, A., Cohen, M. M., Anderson, L. K. et al. Meiotic recombination and spatial proximity in the etiology of the recurrent t(11;22). Am. J. Hum. Genet. 79, 524–538 (2006).
Kurahashi, H. & Emanuel, B. S. Unexpectedly high rate of de novo constitutional t(11;22) translocations in sperm from normal males. Nat. Genet. 29, 139–140 (2001).
Kurahashi, H., Inagaki, H., Ohye, T., Kogo, H., Kato, T. & Emanuel, B. S. Palindrome-mediated chromosomal translocations in humans. DNA Repair (Amst) 5, 1136–1145 (2006).
Kurahashi, H., Inagaki, H., Ohye, T., Kogo, H., Kato, T. & Emanuel, B. S. Chromosomal translocations mediated by palindromic DNA. Cell Cycle 5, 1297–1303 (2006).
Leach, D. R. Long DNA palindromes, cruciform structures, genetic instability and secondary structure repair. Bioessays 16, 893–900 (1994).
Kato, T., Yamada, K., Inagaki, H., Kogo, H., Ohye, T., Emanuel, B. S. et al. Age has no effect on de novo constitutional t(11;22) translocation frequency in sperm. Fertil. Steril. 88, 1446–1448 (2007).
Kovtun, I. V. & McMurray, C. T. Trinucleotide expansion in haploid germ cells by gap repair. Nat. Genet. 27, 407–411 (2001).
Buwe, A., Guttenbach, M. & Schmid, M. Effect of paternal age on the frequency of cytogenetic abnormalities in human spermatozoa. Cytogenet. Genome Res. 111, 213–228 (2005).
Cohen, J. Sorting out chromosome errors. Science 296, 2164–2166 (2002).
Page, S. L. & Shaffer, L. G. Nonhomologous Robertsonian translocations form predominantly during female meiosis. Nat. Genet. 15, 231–232 (1997).
Crow, J. F. The origins, patterns and implications of human spontaneous mutation. Nat. Rev. Genet. 1, 40–47 (2000).
Robinson, W. P. The extent, mechanism, and consequences of genetic variation, for recombination rate. Am. J. Hum. Genet. 59, 1175–1183 (1996).
Hassold, T. & Hunt, P. To err (meiotically) is human: the genesis of human aneuploidy. Nat. Rev. Genet. 2, 280–291 (2001).
Eichenlaub-Ritter, U. Mouse genetic models for aneuploidy induction in germ cells. Cytogenet. Genome. Res. 111, 392–400 (2005).
Champion, M. D. & Hawley, R. S. Playing for half the deck: the molecular biology of meiosis. Nat. Cell Biol. 4, s50–56 (2002).
Neale, M. J. & Keeney, S. Clarifying the mechanics of DNA strand exchange in meiotic recombination. Nature 442, 153–158 (2006).
Hassold, T., Hall, H. & Hunt, P. The origin of human aneuploidy: where we have been, where we are going. Hum. Mol. Genet. 16, R203–208 (2007).
Rubio, C., Simón, C., Vidal, F., Rodrigo, L., Pehlivan, T., Remohí, J. et al. Chromosomal abnormalities and embryo development in recurrent miscarriage couples. Hum. Reprod. 18, 182–188 (2003).
Warburton, D., Dallaire, L., Thangavelu, M., Ross, L., Levin, B. & Kline, J. Trisomy recurrence: a reconsideration based on North American data. Am. J. Hum. Genet. 75, 376–385 (2004).
Bolor, H., Mori, T., Nishiyama, S., Ito, Y., Hosoba, E., Inagaki, H. et al. Mutations of the SYCP3 gene in women with recurrent pregnancy loss. Am. J. Hum. Genet. 84, 14–20 (2009).
Costa, Y. & Cooke, H. J. Dissecting the mammalian synaptonemal complex using targeted mutations. Chromosome Res. 15, 579–589 (2007).
Yuan, L., Liu, J. G., Hoja, M. R., Wilbertz, J., Nordqvist, K. & Hoog, C. Female germ cell aneuploidy and embryo death in mice lacking the meiosis-specific protein SCP3. Science 296, 1115–1118 (2002).
Woods, L. M., Hodges, C. A., Baart, E., Baker, S. M., Liskay, M. & Hunt, P. A. Chromosomal influence on meiotic spindle assembly: abnormal meiosis I in female Mlh1 mutant mice. J. Cell Biol. 145, 1395–1406 (1999).
Kuznetsov, S., Pellegrini, M., Shuda, K., Fernandez-Capetillo, O., Liu, Y., Martin, B. K. et al. RAD51C deficiency in mice results in early prophase I arrest in males and sister chromatid separation at metaphase II in females. J. Cell Biol. 176, 581–592 (2007).
Miyamoto, T., Hasuike, S., Yogev, L., Maduro, M. R., Ishikawa, M., Westphal, H. et al. Azoospermia in patients heterozygous for a mutation in SYCP3. Lancet 362, 1714–1719 (2003).
Yuan, L., Liu, J. G., Zhao, J., Brundell, E., Daneholt, B. & Hoog, C. The murine SCP3 gene is required for synaptonemal complex assembly, chromosome synapsis, and male fertility. Mol. Cell 5, 73–83 (2000).
Hunt, P. A. & Hassold, T. J. Sex matters in meiosis. Science 296, 2181–2183 (2002).
Oliver-Bonet, M., Ko, E. & Martin, R. H. Male infertility in reciprocal translocation carriers: the sex body affair. Cytogenet. Genome Res. 111, 343–346 (2005).
Henderson, S. A. & Edwards, R. G. Chiasma frequency and maternal age in mammals. Nature 218, 22–28 (1968).
Warren, A. C., Chakravarti, A., Wong, C., Slaugenhaupt, S. A., Halloran, S. L., Watkins, P. C. et al. Evidence for reduced recombination on the nondisjoined chromosomes 21 in Down syndrome. Science 237, 652–654 (1987).
Sherman, S. L., Takaesu, N., Freeman, S. B., Grantham, M., Phillips, C., Blackston, R. D. et al. Trisomy 21: association between reduced recombination and nondisjunction. Am. J. Hum Genet. 49, 608–620 (1991).
Sherman, S. L., Petersen, M. B., Freeman, S. B., Hersey, J., Pettay, D., Taft, L. et al. Non-disjunction of chromosome 21 in maternal meiosis I: evidence for a maternal age-dependent mechanism involving reduced recombination. Hum. Mol. Genet. 3, 1529–1535 (1994).
Lamb, N. E., Yu, K., Shaffer, J., Feingold, E. & Sherman, S. L. Association between maternal age and meiotic recombination for trisomy 21. Am. J. Hum. Genet. 76, 91–99 (2005).
Hodges, C. A., Revenkova, E., Jessberger, R., Hassold, T. J. & Hunt, P. A. SMC1beta-deficient female mice provide evidence that cohesins are a missing link in age-related nondisjunction. Nat. Genet. 37, 1351–1355 (2005).
Acknowledgements
We thank Dr Beverly S Emanuel (University of Pennsylvania) for helpful discussion. These studies were supported by a grant-in-aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan and Daiko Foundation to HK.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kurahashi, H., Bolor, H., Kato, T. et al. Recent advance in our understanding of the molecular nature of chromosomal abnormalities. J Hum Genet 54, 253–260 (2009). https://doi.org/10.1038/jhg.2009.35
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2009.35
Keywords
This article is cited by
-
Chromoanagenesis: cataclysms behind complex chromosomal rearrangements
Molecular Cytogenetics (2019)
-
Exonic duplication of the OTC gene by a complex rearrangement that likely occurred via a replication-based mechanism: a case report
BMC Medical Genetics (2018)
-
Molecular profile of atypical hyperplasia of the breast
Breast Cancer Research and Treatment (2018)
-
Preimplantation genetic diagnosis/screening by comprehensive molecular testing
Reproductive Medicine and Biology (2016)
-
Dual roles for the telomeric repeats in chromosomally integrated human herpesvirus-6
Scientific Reports (2014)