Abstract
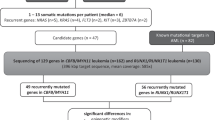
Core-binding factor acute myeloid leukemia (CBF-AML) is defined by the presence of either t(8;21)(q22;q22)/RUNX1-RUNX1T1 or inv(16)(p13.1q22)/t(16;16)(p13.1;q22)/CBFB-MYH11. The resulting fusion genes require a ‘second hit’ to initiate leukemogenesis. Mutation assessment of 177 adults with CBF-AML, including 68 with t(8;21) and 109 with inv(16)/t(16;16), identified not only mutations well known in CBF-AML but also mutations in the CCND1 and CCND2 genes, which represent novel frequent molecular alterations in AML with t(8;21). Altogether, CCND1 (n=2) and CCND2 (n=8) mutations were detected in 10 (15%) patients with t(8;21) in our cohort. A single CCND2 mutation was also found in 1 (0.9%) patient with inv(16). In contrast, CCND1 and CCND2 mutations were detected in only 11 (0.77%) of 1426 non-CBF-AML patients. All CCND2 mutations cluster around the highly conserved amino-acid residue threonine 280 (Thr280). We show that Thr280Ala-mutated CCND2 leads to increased phosphorylation of the retinoblastoma protein, thereby causing significant cell cycle changes and increased proliferation of AML cell lines. The identification of CCND1 and CCND2 mutations as frequent mutational events in t(8;21) AML may provide further justification for cell cycle-directed therapy in this disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Speck NA, Gilliland DG . Core binding factors in haematopoiesis and leukemia. Nat Rev Cancer 2002; 2: 502–513.
Mrózek K, Marcucci G, Paschka P, Bloomfield CD . Advances in molecular genetics and treatment of core-binding factor acute myeloid leukemia. Curr Opin Oncol 2008; 20: 711–718.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 2016; 127: 2391–2405.
Marcucci G, Mrózek K, Ruppert AS, Maharry K, Kolitz JE, Moore JO et al. Prognostic factors and outcome of core binding factor acute myeloid leukemia patients with t(8;21) differ from those with inv(16): a Cancer and Leukemia Group B study. J Clin Oncol 2005; 23: 5705–5717.
Appelbaum FR, Kopecky KJ, Tallman MS, Slovak ML, Gundacker HM, Kim HT et al. The clinical spectrum of adult acute myeloid leukaemia associated with core binding factor translocations. Br J Haematol 2006; 135: 165–173.
Sinha C, Cunningham LC, Liu PP . Core binding factor acute myeloid leukemia: new prognostic categories and therapeutic opportunities. Semin Hematol 2015; 52: 215–222.
Duployez N, Marceau-Renaut A, Boissel N, Petit A, Bucci M, Geffroy S et al. Comprehensive mutational profiling of core binding factor acute myeloid leukemia. Blood 2016; 127: 2451–2459.
Grimwade D, Ivey A, Huntly BJP . Molecular landscape of acute myeloid leukemia in younger adults and its clinical significance. Blood 2016; 127: 29–41.
Micol J-B, Duployez N, Boissel N, Petit A, Geffroy S, Nibourel O et al. Frequent ASXL2 mutations in acute myeloid leukemia patients with t(8;21)/RUNX1-RUNX1T1 chromosomal translocations. Blood 2014; 124: 1445–1449.
Hartmann L, Dutta S, Opatz S, Vosberg S, Reiter K, Leubolt G et al. ZBTB7A mutations in acute myeloid leukaemia with t(8;21) translocation. Nat Commun 2016; 7: 11733.
Lavallée V-P, Lemieux S, Boucher G, Gendron P, Boivin I, Armstrong RN et al. RNA-sequencing analysis of core binding factor AML identifies recurrent ZBTB7A mutations and defines RUNX1-CBFA2T3 fusion signature [letter]. Blood 2016; 127: 2498–2501.
Deshpande A, Sicinski P, Hinds PW . Cyclins and cdks in development and cancer: a perspective. Oncogene 2005; 24: 2909–2915.
Malumbres M, Barbacid M . To cycle or not to cycle: a critical decision in cancer. Nat Rev Cancer 2001; 1: 222–231.
Mrózek K, Carroll AJ, Maharry K, Rao KW, Patil SR, Pettenati MJ et al. Central review of cytogenetics is necessary for cooperative group correlative and clinical studies of adult acute leukemia: the Cancer and Leukemia Group B experience. Int J Oncol 2008; 33: 239–244.
Kroll KW, Eisfeld A-K, Lozanski A, Bloomfield CD, Byrd JC, Blachly JS . MuCor: mutation aggregation and correlation. Bioinformatics 2016; 32: 1557–1558.
Whitman SP, Archer KJ, Feng L, Baldus C, Becknell B, Carlson BD et al. Absence of the wild-type allele predicts poor prognosis in adult de novo acute myeloid leukemia with normal cytogenetics and the internal tandem duplication of FLT3: a Cancer and Leukemia Group B study. Cancer Res 2001; 61: 7233–7239.
Kida A, Kakihana K, Kotani S, Kurosu T, Miura O . Glycogen synthethase kinase-3β and p38 phosphorylate cyclin D2 on Thr280 to trigger its ubiquitin/proteasome-dependent degradation in hematopoietic cells. Oncogene 2007; 26: 6630–6640.
Mirzaa GM, Parry DA, Fry AE, Giamanco KA, Schwartzentruber J, Vanstone M et al. De novo CCND2 mutations leading to stabilization of cyclin D2 cause megalencephaly-polymicrogyria-polydactyly-hydrocephalus syndrome. Nat Genet 2014; 46: 510–515.
Diehl JA, Cheng M, Roussel MF, Sherr CJ . Glycogen synthase kinase-3β regulates cyclin D1 proteolysis and subcellular localization. Genes Dev 1998; 12: 3499–3511.
Alt JR, Cleveland JL, Hannink M, Diehl JA . Phosphorylation-dependent regulation of cyclin D1 nuclear export and cyclin D1-dependent cellular transformation. Genes Dev 2000; 14: 3102–3114.
Shao J, Sheng H, DuBois RN, Beauchamp RD . Oncogenic Ras-mediated cell growth arrest and apoptosis are associated with increased ubiquitin-dependent cyclin D1 degradation. J Biol Chem 2000; 275: 22916–22924.
Ely S, Di Liberto M, Niesvizky R, Baughn LB, Cho HJ, Hatada EN et al. Mutually exclusive cyclin-dependent kinase 4/cyclin D1 and cyclin dependent kinase 6/cyclin D2 pairing inactivates retinoblastoma protein and promotes cell cycle dysregulation in multiple myeloma. Cancer Res 2005; 65: 11345–11353.
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 2013; 368: 2059–2074.
Tartaglia M, Kalidas K, Shaw A, Song X, Musat DL, van der Burgt I et al. PTPN11 mutations in Noonan syndrome: molecular spectrum, genotype-phenotype correlation, and phenotypic heterogeneity. Am J Hum Genet 2002; 70: 1555–1563.
Gibson WT, Hood RL, Zhan SH, Bulman DE, Fejes AP, Moore R et al. Mutations in EZH2 cause Weaver syndrome. Am J Hum Genet 2012; 90: 110–118.
Van Houdt KJK, Nowakowska BA, Sousa SB, van Schaik BDC, Seuntjens E, Avonce N et al. Heterozygous missense mutations in SMARCA2 cause Nicolaides-Baraitser syndrome. Nat Genet 2012; 44: 445–449.
Boyle MI, Jespersgaard C, Brøndum-Nielsen K, Bisgaard A-M, Tümer Z . Cornelia de Lange syndrome. Clin Genet 2015; 88: 1–12.
Musgrove EA, Caldon CE, Barraclough J, Stone A, Sutherland RL . Cyclin D as a therapeutic target in cancer. Nat Rev Cancer 2011; 11: 558–572.
Mao X, Cao B, Wood TE, Hurren R, Tong J, Wang X et al. A small-molecule inhibitor of D-cyclin transactivation displays preclinical efficacy in myeloma and leukemia via phosphoinositide 3-kinase pathway. Blood 2011; 117: 1986–1997.
Acknowledgements
We thank the patients who consented to participate in these clinical trials and the families who supported them; to Donna Bucci and the CALGB/Alliance Leukemia Tissue Bank at The Ohio State University Comprehensive Cancer Center, Columbus, OH, USA for sample processing and storage services and Lisa J Sterling and Chris Finks for data management. This work was supported in part by the National Cancer Institute (grants CA101140, CA140158, CA180861, CA196171, CA016058, CA180821, CA180882 and CA077658), the Leukemia Clinical Research Foundation, the Warren D. Brown Foundation, the Pelotonia Fellowship Program (to A-K Eisfeld) and by an allocation of computing resources from The Ohio Supercomputer Center.
Author contributions
A-KE, JCB, KM and CDB contributed to the study design; A-KE, AdlC, JCB, SS, CJW, KM and CDB contributed to data interpretation, A-KE, KM, AdlC and CDB wrote the manuscript; A-KE and SO generated the libraries for the targeted sequencing approach; A-KE analyzed the sequencing data; CS, MB and CJW performed laboratory-based research; JSB and KWK performed the data processing; JK and DN performed statistical analysis; AJC, JCB, JEK, MRB, RMS, BLP, KM and CDB were involved directly or indirectly in the care of patients and/or sample procurement. All authors read and agreed on the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Eisfeld, AK., Kohlschmidt, J., Schwind, S. et al. Mutations in the CCND1 and CCND2 genes are frequent events in adult patients with t(8;21)(q22;q22) acute myeloid leukemia. Leukemia 31, 1278–1285 (2017). https://doi.org/10.1038/leu.2016.332
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2016.332
This article is cited by
-
Functional characterization of cooperating MGA mutations in RUNX1::RUNX1T1 acute myeloid leukemia
Leukemia (2024)
-
CCND2 and miR-206 as potential biomarkers in the clinical diagnosis of thyroid carcinoma by fine-needle aspiration cytology
World Journal of Surgical Oncology (2023)
-
The Prognostic Significance of the BIN1 and CCND2 Gene in Adult Patients with Acute Myeloid Leukemia
Indian Journal of Hematology and Blood Transfusion (2022)
-
Cytogenetic and mutational analysis and outcome assessment of a cohort of 284 children with de novo acute myeloid leukemia reveal complex karyotype as an adverse risk factor for inferior survival
Molecular Cytogenetics (2021)
-
Inhibition of CDK4/6 and autophagy synergistically induces apoptosis in t(8;21) acute myeloid leukemia cells
International Journal of Hematology (2021)