Abstract
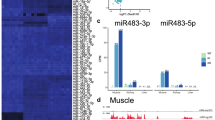
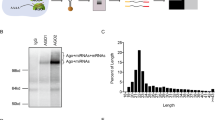
The H19 large intergenic non-coding RNA (lincRNA) is one of the most highly abundant and conserved transcripts in mammalian development, being expressed in both embryonic and extra-embryonic cell lineages, yet its physiological function is unknown. Here we show that miR-675, a microRNA (miRNA) embedded in H19’s first exon, is expressed exclusively in the placenta from the gestational time point when placental growth normally ceases, and placentas that lack H19 continue to grow. Overexpression of miR-675 in a range of embryonic and extra-embryonic cell lines results in their reduced proliferation; targets of the miRNA are upregulated in the H19 null placenta, including the growth-promoting insulin-like growth factor 1 receptor (Igf1r) gene. Moreover, the excision of miR-675 from H19 is dynamically regulated by the stress-response RNA-binding protein HuR. These results suggest that H19’s main physiological role is in limiting growth of the placenta before birth, by regulated processing of miR-675. The controlled release of miR-675 from H19 may also allow rapid inhibition of cell proliferation in response to cellular stress or oncogenic signals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Brannan, C. I., Dees, E. C., Ingram, R. S. & Tilghman, S. M. The product of the H19 gene may function as an RNA. Mol. Cell. Biol. 10, 28–36 (1990).
Seidl, C. I., Stricker, S. H. & Barlow, D. P. The imprinted Air ncRNA is an atypical RNAPII transcript that evades splicing and escapes nuclear export. EMBO. J. 25, 3565–3575 (2006).
Bartolomei, M. S. Genomic imprinting: employing and avoiding epigenetic processes. Genes Dev. 23, 2124–2133 (2009).
Gabory, A., Jammes, H. & Dandolo, L. The H19 locus: role of an imprinted non-coding RNA in growth and development. Bioessays 32, 473–480 (2010).
Hao, Y., Crenshaw, T., Moulton, T., Newcomb, E. & Tycko, B. Tumour-suppressor activity of H19 RNA. Nature 365, 764–767 (1993).
Yoshimizu, T. et al. The H19 locus acts in vivo as a tumor suppressor. Proc. Natl Acad. Sci. USA 105, 12417–12422 (2008).
Smits, G. et al. Conservation of the H19 noncoding RNA and H19-IGF2 imprinting mechanism in therians. Nat. Genet. 40, 971–976 (2008).
Gabory, A. et al. H19 acts as a trans regulator of the imprinted gene network controlling growth in mice. Development 136, 3413–3421 (2009).
Wilusz, J. E., Sunwoo, H. & Spector, D. L. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 23, 1494–1504 (2009).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Huntzinger, E. & Izaurralde, E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat. Rev. Genet. 12, 99–110 (2011).
Cai, X. & Cullen, B. R. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 13, 313–316 (2007).
Dudek, K. A., Lafont, J. E., Martinez-Sanchez, A. & Murphy, C. L. Type II collagen expression is regulated by tissue-specific miR-675 in human articular chondrocytes. J. Biol. Chem. 285, 24381–24387 (2010).
Chiang, H. R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Mineno, J. et al. The expression profile of microRNAs in mouse embryos. Nucleic Acids Res. 34, 1765–1771 (2006).
Yang, J. H., Shao, P., Zhou, H., Chen, Y. Q. & Qu, L. H. deepBase: a database for deeply annotating and mining deep sequencing data. Nucleic Acids Res. 38, D123-D130 (2010).
Lee, Y. S. & Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 21, 1025–1030 (2007).
Lim, L. P. et al. The microRNAs of Caenorhabditis elegans. Genes Dev. 17, 991–1008 (2003).
Knox, K. & Baker, J. C. Genomic evolution of the placenta using co-option and duplication and divergence. Genome Res. 18, 695–705 (2008).
Zhao, Z., Chang, F. C. & Furneaux, H. M. The identification of an endonuclease that cleaves within an HuR binding site in mRNA. Nucleic Acids Res. 28, 2695–2701 (2000).
Katsanou, V. et al. The RNA-binding protein Elavl1/HuR is essential for placental branching morphogenesis and embryonic development. Mol. Cell. Biol. 29, 2762–2776 (2009).
Kim, V. N., Han, J. & Siomi, M. C. Biogenesis of small RNAs in animals. Nat. Rev. Mol. Cell Biol. 10, 126–139 (2009).
Runge, S. et al. H19 RNA binds four molecules of insulin-like growth factor II mRNA-binding protein. J. Biol. Chem. 275, 29562–29569 (2000).
Mallanna, S. K. et al. Proteomic analysis of Sox2-associated proteins during early stages of mouse embryonic stem cell differentiation identifies Sox21 as a novel regulator of stem cell fate. Stem Cells 28, 1715–1727 (2010).
Coan, P. M., Ferguson-Smith, A. C. & Burton, G. J. Developmental dynamics of the definitive mouse placenta assessed by stereology. Biol. Reprod. 70, 1806–1813 (2004).
Angiolini, E. et al. Developmental adaptations to increased fetal nutrient demand in mouse genetic models of Igf2-mediated overgrowth. FASEB J. 25, 1737–1745 (2011).
Esquiliano, D. R., Guo, W., Liang, L., Dikkes, P. & Lopez, M. F. Placental glycogen stores are increased in mice with H19 null mutations but not in those with insulin or IGF type 1 receptor mutations. Placenta 30, 693–699 (2009).
Leighton, P. A., Ingram, R. S., Eggenschwiler, J., Efstratiadis, A. & Tilghman, S. M. Disruption of imprinting caused by deletion of the H19 gene region in mice. Nature 375, 34–39 (1995).
Thorvaldsen, J. L., Duran, K. L. & Bartolomei, M. S. Deletion of the H19 differentially methylated domain results in loss of imprinted expression of H19 and Igf2. Genes Dev. 12, 3693–3702 (1998).
Ripoche, M. A., Kress, C., Poirier, F. & Dandolo, L. Deletion of the H19 transcription unit reveals the existence of a putative imprinting control element. Genes Dev. 11, 1596–1604 (1997).
Liu, J. P., Baker, J., Perkins, A. S., Robertson, E. J. & Efstratiadis, A. Mice carrying null mutations of the genes encoding insulin-like growth factor I (Igf-1) and type 1 IGF receptor (Igf1r). Cell 75, 59–72 (1993).
Baker, J., Liu, J. P., Robertson, E. J. & Efstratiadis, A. role of insulin-like growth factors in embryonic and postnatal growth. Cell 75, 73–82 (1993).
Jeyaraj, S. C., Dakhlallah, D., Hill, S. R. & Lee, B.S. Expression and distribution of HuR during ATP depletion and recovery in proximal tubule cells. Am. J. Physiol. Renal. Physiol. 291, F1255–F1263 (2006).
Kim, H. H., Abdelmohsen, K. & Gorospe, M. Regulation of HuR by DNA damage response kinases. J. Nucleic Acids 2010 (2010).
Pan, Y. X., Chen, H. & Kilberg, M. S. Interaction of RNA-binding proteins HuR and AUF1 with the human ATF3 mRNA 3’-untranslated region regulates its amino acid limitation-induced stabilization. J. Biol. Chem. 280, 34609–34616 (2005).
Wang, W. et al. HuR regulates p21 mRNA stabilization by UV light. Mol. Cell. Biol. 20, 760–769 (2000).
Blaxall, B. C. et al. Differential expression and localization of the mRNA binding proteins, AU-rich element mRNA binding protein (AUF1) and Hu antigen R (HuR), in neoplastic lung tissue. Mol. Carcinog. 28, 76–83 (2000).
Lebedeva, S. et al. Transcriptome-wide analysis of regulatory interactions of the RNA-binding protein HuR. Mol. Cell 43, 340–352 (2011).
Reddy, S. D., Ohshiro, K., Rayala, S. K. & Kumar, R. MicroRNA-7, a homeobox D10 target, inhibits p21-activated kinase 1 and regulates its functions. Cancer Res. 68, 8195–8200 (2008).
Saydam, O. et al. miRNA-7 attenuation in Schwannoma tumors stimulates growth by upregulating three oncogenic signaling pathways. Cancer Res. 71, 852–861 (2011).
Jiang, L. et al. MicroRNA-7 targets IGF1R (insulin-like growth factor 1 receptor) in tongue squamous cell carcinoma cells. Biochem. J. 432, 199–205 (2010).
Lim, D. H. & Maher, E. R. Genomic imprinting syndromes and cancer. Adv. Genet. 70, 145–175 (2010).
Tanaka, S., Kunath, T., Hadjantonakis, A. K., Nagy, A. & Rossant, J. Promotion of trophoblast stem cell proliferation by FGF4. Science 282, 2072–2075 (1998).
Ficz, G. et al. Dynamic regulation of 5-hydroxymethylcytosine in mouse ES cells and during differentiation. Nature 473, 398–402 (2011).
Caputi, M., Mayeda, A., Krainer, A. R. & Zahler, A. M. hnRNP A/B proteins are required for inhibition of HIV-1 pre-mRNA splicing. EMBO J. 18, 4060–4067 (1999).
Baroni, T. E., Chittur, S. V., George, A. D. & Tenenbaum, S. A. Advances in RIP-chip analysis: RNA-binding protein immunoprecipitation-microarray profiling. Methods Mol. Biol. 419, 93–108 (2008).
Acknowledgements
We thank D. L. Kontoyiannis (Alexander Fleming Biomedical Sciences Research Center, Greece) for supplying H u R−/− MEF cells. We also thank H. Jammes and A. Gabory for assisting with collection of the H19Δ3 phenotypic data, J. Webster for preparing samples for mass spectrometry and W. Dean for tissue collections. The authors would also like to thank K. Tabbada, A. Segonds-Pichon and S. Andrews for RNA sequencing, statistical and bioinformatic assistance, respectively. We thank M. Iacovino for creating the A2lox-cre ES cell line. We would also like to thank T. Hore, C. Krueger and J. Houseley for critical reading of the manuscript and all members of the laboratories of W. Reik, M. Hemberger, E. Vigorito and J. Houseley for helpful discussions. This work was supported by BBSRC, the Wellcome Trust, MRC, EU NoE EpiGeneSys, EU BLUEPRINT, NIH/NHLBI (U01HL100407) and the Cambridge Commonwealth Trust.
Author information
Authors and Affiliations
Contributions
A.K. designed and carried out experiments and interpreted results. D.O. performed mass spectrometry. P.M. collected and dissected placentas for RNA sequencing. M.K. made the A2lox.cre ES cell line. L.D., G.S. and W.R. designed and supervised this work and interpreted results. A.K. and W.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 792 kb)
Supplementary Tables 1–4
Supplementary Information (XLSX 914 kb)
Rights and permissions
About this article
Cite this article
Keniry, A., Oxley, D., Monnier, P. et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat Cell Biol 14, 659–665 (2012). https://doi.org/10.1038/ncb2521
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2521