Abstract
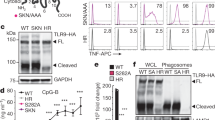
Sensors of the innate immune system that detect intracellular nucleic acids must be regulated to prevent inappropriate activation by endogenous DNA and RNA. The exonuclease Trex1 regulates the DNA-sensing pathway by metabolizing potential DNA ligands that trigger it. However, an analogous mechanism for regulating the RIG-I-like receptors (RLRs) that detect RNA remains unknown. We found here that the SKIV2L RNA exosome potently limited the activation of RLRs. The unfolded protein response (UPR), which generated endogenous RLR ligands through the cleavage of cellular RNA by the endonuclease IRE-1, triggered the production of type I interferons in cells depleted of SKIV2L. Humans with deficiency in SKIV2L had a type I interferon signature in their peripheral blood. Our findings reveal a mechanism for the intracellular metabolism of immunostimulatory RNA, with implications for specific autoimmune disorders.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Barbalat, R., Ewald, S.E., Mouchess, M.L. & Barton, G.M. Nucleic acid recognition by the innate immune system. Annu. Rev. Immunol. 29, 185–214 (2011).
Pichlmair, A. & Reis e Sousa, C. Innate recognition of viruses. Immunity 27, 370–383 (2007).
Kato, H. et al. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J. Exp. Med. 205, 1601–1610 (2008).
Stetson, D.B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Xiao, T.S. & Fitzgerald, K.A. The cGAS-STING pathway for DNA sensing. Mol. Cell 51, 135–139 (2013).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z.J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Jin, T. et al. Structures of the HIN domain:DNA complexes reveal ligand binding and activation mechanisms of the AIM2 inflammasome and IFI16 receptor. Immunity 36, 561–571 (2012).
Liao, J.C. et al. Interferon-inducible protein 16: insight into the interaction with tumor suppressor p53. Structure 19, 418–429 (2011).
Civril, F. et al. Structural mechanism of cytosolic DNA sensing by cGAS. Nature 498, 332–337 (2013).
Stetson, D.B., Ko, J.S., Heidmann, T. & Medzhitov, R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 134, 587–598 (2008).
Crow, Y.J. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 cause Aicardi-Goutieres syndrome at the AGS1 locus. Nat. Genet. 38, 917–920 (2006).
Gall, A. et al. Autoimmunity initiates in nonhematopoietic cells and progresses via lymphocytes in an interferon-dependent autoimmune disease. Immunity 36, 120–131 (2012).
Garneau, N.L., Wilusz, J. & Wilusz, C.J. The highways and byways of mRNA decay. Nat. Rev. Mol. Cell Biol. 8, 113–126 (2007).
Schneider, C. & Tollervey, D. Threading the barrel of the RNA exosome. Trends Biochem. Sci. 38, 485–493 (2013).
Brown, J.T., Bai, X. & Johnson, A.W. The yeast antiviral proteins Ski2p, Ski3p, and Ski8p exist as a complex in vivo. RNA 6, 449–457 (2000).
Fernando, M.M. et al. Identification of two independent risk factors for lupus within the MHC in United Kingdom families. PLoS Genet. 3, e192 (2007).
Saito, T., Owen, D.M., Jiang, F., Marcotrigiano, J. & Gale, M. Jr. Innate immunity induced by composition-dependent RIG-I recognition of hepatitis C virus RNA. Nature 454, 523–527 (2008).
Roberts, Z.J. et al. The chemotherapeutic agent DMXAA potently and specifically activates the TBK1-IRF-3 signaling axis. J. Exp. Med. 204, 1559–1569 (2007).
Conlon, J. et al. Mouse, but not human STING, binds and signals in response to the vascular disrupting agent 5,6-dimethylxanthenone-4-acetic acid. J. Immunol. 190, 5216–5225 (2013).
Yamamoto, M. et al. Role of adaptor TRIF in the MyD88-independent Toll-like receptor signaling pathway. Science 301, 640–643 (2003).
Bernales, S., Papa, F.R. & Walter, P. Intracellular signaling by the unfolded protein response. Annu. Rev. Cell Dev. Biol. 22, 487–508 (2006).
Travers, K.J. et al. Functional and genomic analyses reveal an essential coordination between the unfolded protein response and ER-associated degradation. Cell 101, 249–258 (2000).
Cox, J.S., Shamu, C.E. & Walter, P. Transcriptional induction of genes encoding endoplasmic reticulum resident proteins requires a transmembrane protein kinase. Cell 73, 1197–1206 (1993).
Mori, K., Ma, W., Gething, M.J. & Sambrook, J. A transmembrane protein with a cdc2+/CDC28-related kinase activity is required for signaling from the ER to the nucleus. Cell 74, 743–756 (1993).
Credle, J.J., Finer-Moore, J.S., Papa, F.R., Stroud, R.M. & Walter, P. On the mechanism of sensing unfolded protein in the endoplasmic reticulum. Proc. Natl. Acad. Sci. USA 102, 18773–18784 (2005).
Gardner, B.M. & Walter, P. Unfolded proteins are Ire1-activating ligands that directly induce the unfolded protein response. Science 333, 1891–1894 (2011).
Cox, J.S. & Walter, P. A novel mechanism for regulating activity of a transcription factor that controls the unfolded protein response. Cell 87, 391–404 (1996).
Calfon, M. et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 415, 92–96 (2002).
Sidrauski, C., Cox, J.S. & Walter, P. tRNA ligase is required for regulated mRNA splicing in the unfolded protein response. Cell 87, 405–413 (1996).
Hollien, J. & Weissman, J.S. Decay of endoplasmic reticulum-localized mRNAs during the unfolded protein response. Science 313, 104–107 (2006).
Hollien, J. et al. Regulated Ire1-dependent decay of messenger RNAs in mammalian cells. J. Cell Biol. 186, 323–331 (2009).
Malathi, K., Dong, B., Gale, M. Jr. & Silverman, R.H. Small self-RNA generated by RNase L amplifies antiviral innate immunity. Nature 448, 816–819 (2007).
Takahasi, K. et al. Nonself RNA-sensing mechanism of RIG-I helicase and activation of antiviral immune responses. Mol. Cell 29, 428–440 (2008).
Cho, J.A. et al. The unfolded protein response element IRE1α senses bacterial proteins invading the ER to activate RIG-I and innate immune signaling. Cell Host Microbe 13, 558–569 (2013).
Han, D. et al. IRE1alpha kinase activation modes control alternate endoribonuclease outputs to determine divergent cell fates. Cell 138, 562–575 (2009).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mali, P., Esvelt, K.M. & Church, G.M. Cas9 as a versatile tool for engineering biology. Nat. Methods 10, 957–963 (2013).
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Sun, Q. et al. The specific and essential role of MAVS in antiviral innate immune responses. Immunity 24, 633–642 (2006).
Fabre, A. et al. SKIV2L mutations cause syndromic diarrhea, or trichohepatoenteric syndrome. Am. J. Hum. Genet. 90, 689–692 (2012).
Fabre, A., Martinez-Vinson, C., Goulet, O. & Badens, C. Syndromic diarrhea/Tricho-hepato-enteric syndrome. Orphanet J. Rare Dis. 8, 5 (2013).
Hartley, J.L. et al. Mutations in TTC37 cause trichohepatoenteric syndrome (phenotypic diarrhea of infancy). Gastroenterology 138, 2388–2398 (2010).
Fabre, A. et al. Novel mutations in TTC37 associated with tricho-hepato-enteric syndrome. Hum. Mutat. 32, 277–281 (2011).
Rice, G.I. et al. Assessment of interferon-related biomarkers in Aicardi-Goutieres syndrome associated with mutations in TREX1, RNASEH2A, RNASEH2B, RNASEH2C, SAMHD1, and ADAR: a case-control study. Lancet Neurol. 12, 1159–1169 (2013).
Acknowledgements
We thank M. Gale, Jr. (University of Washington) for Mavs−/− mice, and members of the Stetson laboratory for discussions. Supported by the National Institute of Allergy and Infectious Disease (AI084914 to D.B.S.), the Nuclease Immune Mediated Brain and Lupus-like conditions (NIMBL) project of the European Union Seventh Framework Programme 2007-2013 (241779 to Y.J.C. and D.B.S.), the Lupus Research Institute (D.B.S.), the Cancer Research Institute (E.E.G.) and the Rita Allen Foundation (D.B.S.).
Author information
Authors and Affiliations
Contributions
S.C.E. did all experiments with mouse cells; G.I.R. and Y.J.C. did the ISG analysis of human cells and helped write the manuscript; A.F., C.B. and J.L.H. obtained peripheral blood samples from patients with THES and control subjects, provided intellectual input and insights into human THES and helped write the manuscript; E.E.G. developed the Lenti-CRISPR system; and S.C.E. and D.B.S. designed the study and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Restriction fragment length polymorphism (RFLP) analysis of mouse fibroblasts in which Ern1 was targeted.
(a) MEFs immortalized with SV40 Large T antigen were transduced with lentiviruses encoding CAS9 alone, or CAS9 plus one of two guide RNAs targeting the murine Ern1 gene. Genomic DNA was isolated at the indicated times post selection of transduced cells, amplified with PCR primers flanking the targeted site, and digested with the indicated restriction enzymes. For guide RNA 1 (top), the digested wild-type DNA results in products of 191 bp and 103 bp. Targeted alleles are resistant to BsajI digestion and are ~294 bp, with a concomitant loss of the wild-type digestion products. For guide RNA 2 (bottom), the digested wild-type allele results in products of 157 bp and 96 bp. The targeted alleles are resistant to MlucI digestion and are ~253 bp. The asterisk in the bottom panel denotes a MlucI digestion product of the PCR that is outside the targeted region.
(b) A schematic view of the restriction enzyme sites within amplified genomic DNA for each guide RNA. The NHEJ product denotes insertions/deletions at the CRISPR/CAS9 target site that disrupt the restriction enzyme site.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Supplementary Table 1 (PDF 576 kb)
Rights and permissions
About this article
Cite this article
Eckard, S., Rice, G., Fabre, A. et al. The SKIV2L RNA exosome limits activation of the RIG-I-like receptors. Nat Immunol 15, 839–845 (2014). https://doi.org/10.1038/ni.2948
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2948
This article is cited by
-
Emerging role of the RNA-editing enzyme ADAR1 in stem cell fate and function
Biomarker Research (2023)
-
Cellular functions of eukaryotic RNA helicases and their links to human diseases
Nature Reviews Molecular Cell Biology (2023)
-
RBP–RNA interactions in the control of autoimmunity and autoinflammation
Cell Research (2023)
-
The genetics of monogenic intestinal epithelial disorders
Human Genetics (2023)
-
Cellular origins of dsRNA, their recognition and consequences
Nature Reviews Molecular Cell Biology (2022)