Abstract
Long non-coding RNAs (lncRNAs) were reported to potentially play a regulatory role in the process of myocardial regeneration in the neonatal mouse. N6-methyladenosine (m6A) modification may play a key role in myocardial regeneration in mice and regulates a variety of biological processes through affecting the stability of lncRNAs. However, the map of m6A modification of lncRNAs in mouse cardiac development still remains unknown. We aimed to investigate the differences in the m6A status of lncRNAs during mouse cardiac development and reveal a potential role of m6A modification modulating lncRNAs in cardiac development and myocardial regeneration during cardiac development in mice. Methylated RNA immunoprecipitation sequencing (MeRIP-seq) and RNA sequencing (RNA-seq) of the heart tissue in C57BL/6 J mice at postnatal day 1 (P1), P7 and P28 were performed to produce stagewise cardiac lncRNA m6A-methylomes in a parallel timeframe with the established loss of an intrinsic cardiac regeneration capacity and early postnatal development. There were significant differences in the distribution and abundance of m6A modifications in lncRNAs in the P7 vs P1 mice. In addition, the functional role of m6A in regulating lncRNA levels was established for selected transcripts with METTL3 silencing in neonatal cardiomyocytes in vitro. Based on our MeRIP-qPCR experiment data, both lncGm15328 and lncRNA Zfp597, that were not previously associated with cardiac regeneration, were found to be the most differently methylated at P1-P7. These two lncRNAs sponged several miRNAs which further regulated multiple mRNAs, including some of which have previously been linked with cardiac regeneration ability. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes analysis revealed that differential m6A modifications were more enriched in functions and cellular signalling pathways related to cardiomyocyte proliferation. Our data suggested that the m6A modification on lncRNAs may play an important role in the regeneration of myocardium and cardiac development.
Similar content being viewed by others
Introduction
The heart is a terminally differentiated organ and it has been accepted that the heart lacks the ability to regenerate [1, 2]. Notably, excellent heart functions and long-term outcomes in human newborns and neonates/toddles were noted following treatment or surgery from massive cardiac infarction [3] or congenital heart disease [4], respectively, indicating that the young heart may have some regenerating capability. However, self-renewal of cardiomyocytes in humans, and that the renewal rate of cardiomyocytes decreases with age [5, 6]. The myocardium of the postnatal day 1 (P1) mouse can be completely regenerated after surgical resection of the apex of heart, this regenerative ability is lost at P7 [7, 8]. Although the underlying mechanisms of the lost ability remain unknown, these including cardiac polyploidy, multi-levelled early innate immune system, “cancer risk” suppression and cardiac thyroxin signaling activation may likely be a barrier for cardiomyocyte proliferation [9,10,11,12,13]. Thus, studying the changes of myocardial regeneration ability during the neonatal-to-adult mouse heart transition may help to elucidate the mechanism of myocardium regeneration.
N6-methyladenosine (m6A) is the most prevalent internal modification in eukaryotic cells [14]. m6A modification reflects a dynamic and reversible process, which involves methyltransferases (METTL3, METTL14 and WTAP), demethylases (FTO, ALKBH5 and ALKBH3) and RNA-binding proteins (YTHDF1-3, YTHDC1-2 and IGF2BPs) processing [15]. A previous study showed that METTL3 was downregulated in the postnatal day (P7) relative to P0 heart in rat newborns, and enhancing METTL3 expression level of P0 cardiomyocytes resulted in increased proliferation in vitro[16]; however, in opposition to this, a recent study showed identified global METTL3 knockout enhanced cardiac regeneration and repair after myocardial injury [17]. It was also reported that m6A methylation was significantly increased and the expression level of ALKBH5 was decreased after birth in the mouse heart and its overexpression promoted myocardial regeneration and repair after the myocardial infarction (MI) [18]. m6A methylation was reported to be the key regulator of cardiac development and myocardial regeneration [18, 19], but the underlying mechanisms remain elusive.
LncRNAs are a class of special RNA transcripts with more than 200 nucleotides and are widely known to participate in various biological and pathological processes [20, 21]. Myocardial regeneration is regulated by lncRNAs [22,23,24]. The lncRNA CAREL was upregulated in the P7 mouse heart in parallel with loss of cardiac regenerative capacity. Thus, lncRNA CAREL overexpression in cardiomyocytes inhibited its proliferation, whilst silencing it in the heart promoted myocardial regeneration and improved cardiac function in the infarcted area [25]. lncRNA ECRAR expression was the highest in the P1 rat heart and then was progressively decreased in rat hearts after birth; its overexpression not only promoted myocardial regeneration but also improved post-infarction cardiac function [26]. Taken together, current evidence strongly suggests that cardiomyocyte proliferation can be triggered by a variety of endogenous factors.
Here, we performed a m6A-specific analysis and related bioinformatics analysis of m6A on lncRNAs from the heart tissue of C57BL/6 J mice at P1, P7, and P28. These approaches produced multiple datasets describing cardiac lncRNA m6A methylomes from different developmental stages early after birth which appear in a parallel timeframe with the established loss of intrinsic cardiac regeneration capacity. Thus, our work reported here may open avenues for identifying novel regeneration-related lncRNAs and molecular pathways regulated by m6A.
Results
Motif analysis
HOMER software was applied to map the myocardial lncRNA m6A methylomes (P1, P7 and P28). RRACH (R = G or A; H = A, C or U) is the most conserved sequence motif in P1 (Fig. 1A, P < 0.05), which is in line with the previous studies [27, 28]. Meanwhile, P7 conserved motif is RRA (A/N)(A/N), and P28 conserved motif is RRAHC (Fig. 1B, C, P < 0.05).
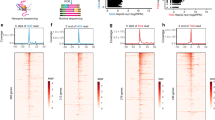
A–C top m6A motifs enriched from all identified m6A peaks in postnatal day 1 (P1), P7, P28 heart tissue; P < 0.05 (n = 3) (hypergeometric test). D–F Pie chart showing the distribution profiles of m6A peaks for host genes of lncRNA in P1, P7, P28 heart tissue. G–I The distribution of m6A peaks on exon length; Adjusted p values with False Discovery Rate; p < 0.05 (n = 3) (hypergeometric test). (control: lncRNAs without m6A peaks; peak: lncRNAs with m6A peaks).
Distribution of m6A peaks for host genes of lncRNAs
We analyzed the data and plotted pie charts to identify the distribution profiles of m6A peaks for host genes of lncRNAs (Fig. 1D–F). Most m6A peaks were enriched in the promoter region, followed by 3′ UTR region. During the development of the mouse heart, m6A peaks showed great changes in the promoter region between P1 (65.3%) and P7 (50.7%), but there was a marginal difference between P7 (50.7%) and P28 (51.7%) groups. Additionally, the exon length of lncRNA with m6A peaks had no significant difference among the three groups (Fig. 1G–I, Supplemental Fig. S1). These data suggested that from day 1 to 7 after birth, the distribution of m6A peaks underwent a major reorganization in a short period of 7 days, while the lncRNA m6A methylome remained relatively stable in terms of m6A peak distribution across lncRNAs after P7.
Abundance features of m6A peaks on lncRNAs and Chromosomes
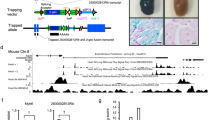
To gain insight into transcriptome-wide m6A methylation, m6A-sequencing of lncRNAs in C57BL/6 J mouse heart tissue of P1, P7, and P28 was performed. lncRNAs m6A peaks were to be 248 at P1, increased to 1,239 at P7 and decreased to 808 at P28. The differences and overlaps of m6A peaks among these three groups are displayed in a Venn diagram, and some redundancy data were merged in a statistical mapping (group P1 were compared with group P7, and one peak in P1 overlapped with two or more peaks in P7, namely redundancy). Among them, only 1 m6A peak (host gene name: Aopep, Aminopeptidase O) was common between the P1 and P7 mice, and this single conserved m6A lncRNA peak across all study groups (Fig. 2A, B, D). However, there are 605 overlapping m6A peaks between P7 and P28 (Fig. 2C). The majority of lncRNAs contained only one m6A peak (more than 50%), and this ratio is highest at P1 (P1:66.5%, P7:52.1%, P28:56.4%) (Fig. 2E). Towards adolescence, the m6A in lncRNAs within mouse myocardium was increased in stoichiometry, but the datasets showed that multiply methylated lncRNAs was not associated with different expression levels. Circos software was used to analyze the distribution of lncRNA methylation sites on the chromosomes (Fig. 2F). It can be seen that there was a disappearance of the highly prominent P1 lncRNA m6A peak at chromosome 14 around Myh6-Myh7 host genes by the P7 and P28.
A Venn diagram showing the overlap of m6A peaks within lncRNAs in postnatal day 7 (P7) and P1 heart tissue. B Venn diagram showing the overlap of m6A peaks within lncRNAs in P28 and P1 heart tissue. C Venn diagram showing the overlap of m6A peaks within lncRNAs in P28 and P7 heart tissue. D Venn diagram showing the overlap of m6A peaks within lncRNAs in P1, P7 and P28 heart tissue. E Percentage of lncRNAs harboring different m6A peaks in these three groups, it seems that most of the methylated lncRNAs harbring only one m6A peak. F Circos plot showing the distribution of the m6A sites within lncRNAs on each chromosome in P1, P7 and P28 heart tissue (red: P1; blue: P7; green: P28).
Correlation analysis between m6A modifications and lncRNAs expression
To uncover whether the differentially expressed lncRNAs were associated with m6A methylation changes, we performed the cross-analysis of the RNA-Seq and m6A-Seq data. The upregulated lncRNAs solely were either hypomethylated or not methylated at all, while the downregulated lncRNAs were either hypermethylated, hypomethylated or not methylated when comparing P7 to P1 (Fig. 3A). These indicated that the methylation-status associated poorly with downregulation of lncRNA expression whilst the lncRNA hypomethylation was associated more consistently with the respective transcript overexpression. However, in comparisons with P28 vs P1 and P28 vs P7 together, there was a non-existent correlation with the lncRNA expression (Fig. 3B, C). To further assess the m6A methylation levels difference of lncRNAs in these three developmental stages of mouse heart tissue, the clustering of methylation differences were clearly be distinguished in the P1, P7 and P28 mouse heart (Fig. 3D). The differentially methylated and expressed lncRNA transcripts were more numerous in P7 vs P1, indicating that m6A modification might be associated with and even regulate lncRNA expression especially in P1 to P7.
A–C. The relationship between lncRNAs expression level and methylation level; P < 0.05 (n = 3) (Student t-test), Red dot (High): m6A level increased (FC > 1.5, P < 0.05), Blue dot (Low): m6A level decreased (FC > 1.5, P < 0.05), Gray dot (No): m6A level variation: |FC | < 1.5. D Cluster analysis of m6A in postnatal day 1 (P1), P7 and P28 heart tissue, red color for hypermethylated and blue for hypomethylated. The gradient color legend unit indicates the fold change number of m6A, and use the pheatmap function of the pheatmap package in R to normalize the fold change number.
Among the 40 lncRNAs with the largest methylation modified fold changes (Supplemental Table S2-5), some of these lncRNAs had big differences in m6A methylation such as lncRNA Myh7. Interestingly, Myh7 gene was reported to be a classic biomarker for cardiac development and a potential target for attenuating cardiomyocyte hypertrophy [29, 30]. However, lncRNA Myh7 was not changed in our study which may indicate that the change of lncRNA Myh7 m6A methylation during P1-P28 was unlikely converted to the altered itself expression.
m6A RNA methylation-related lncRNAs (m6A-lncRNAs) may have potential effect on heart development
By cross-analysis of the m6A-Seq and RNA-seq data, a total of 38 lncRNAs was found be with expression changes closely related to m6A modifications in the P7 vs P1 group (Fig. 4A and Supplemental Table S6).
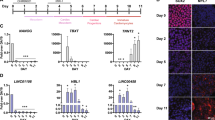
A Veen diagram of lncRNAs of different expression and methylation including up-regulated and down-regulated between postnatal day 1 (P1) and P7. B RT-qPCR assay was applied to determine the expression of 9 genes related to cardiac disease. C RT-qPCR analysis of METTL3 in cultured cardiomyocytes transfected with siRNA and negative control siRNA (si-NC). D Western blot analysis of METTL3 of cardiomyocytes transfected with siRNA and siRNA-ctrl, quantitated by Image J. E The expression of lncRNAs by RT-qPCR in siRNA-transfected cardiomyocytes, Data expressed as mean ± SEM (n = 5); *P < 0.05 (Student t-test). F MeRIP-qPCR results of lncRNA Snhg3 and lncRNA Nedd4 transfected with si-Mettl3 of cardiomyocytes, showing the m6A levels of lncRNA Snhg3 and lncRNA Nedd4 in the way of bounding with anti-m6A antibody, Data expressed as mean ± SEM (n = 3); *P < 0.05 (Student t-test). %input means the percentage of genes that has been methylated. %input=2(-ΔCt normalized RIP). n.s.: P > 0.05; *P < 0.05; **P < 0.01; ***P < 0.001.
To explore the impact of m6A modifications on lncRNAs expression in mouse heart development, 9 of these 38 lncRNAs (Supplemental Table S1) were found either to be related to myocardial development or regeneration, or otherwise, may be the new molecules that potentially regulates myocardial regeneration. Eight of 9 lncRNAs expression in P1 and P7 mouse heart tissue were consistent with qPRCR sequence (Fig. 4B). It is known that METTL3 is the catalytic subunit responsible for the m6A writing by the MTC (methyltransferase complex) [15]. Additionally, METTL3 mediated m6A methylation within myocardium plays an indispensable role in cardiac homeostasis especially in aging [19]. We then used siRNA targeting METTL3 to reduce the endogenous expression of METTL3 in neonatal mouse cardiomyocytes (NMCMs) (Fig. 4C, D). As shown in Fig. 4E, knockdown of METTL3 in NMCMs, among the lncRNAs with changes in expression, lncRNA Snhg3 was the most significant increase (Fold Change = 3.02) and lncRNA Nedd4 was the most significant decrease. Meanwhile, m6A modification of ‘lncRNA Snhg3’ and ‘lncRNA Nedd4’ were decreased upon METTL3 knockdown as shown by MeRIP-qPCR assay (Fig. 4F), suggesting that these two lncRNAs may be the target of METTL3. Moreover, the Nedd4 has earlier been reported to be a potential key factor for myocardial regeneration [31].
lncRNAs can act as microRNA (miRNA) sponges in regulating protein-coding gene expression [32]. The lncRNA-miRNA-mRNA regulatory network plays critical role in cardiac regeneration [33]. Among the 38 lncRNAs, two lncRNAs (LncGm15328: the largest increase of m6A modification; lncRNA Zfp597: the largest decrease of m6A modification) with the biggest difference m6A modification were selected to establish a ceRNA network (Fig. 5). The lncRNA-miRNA-mRNA network LncGm15328 and lncRNA Zfp597 was related to the target of miR-19a/19b and miR-9, respectively. Existing literatures reported that miR-19a/19b was directly involved in regulating myocardial regeneration, and miR-9 was shown to be correlated with cardiogenin-treated regenerating heart [34, 35].
Gene ontology and Kyoto Encyclopedia of Genes and Genomes analyses of lncRNAs harboring differentially methylated N6-methyladenosine sites
To investigate the function of differentially methylated lncRNAs in P1, P7, and P28 heart tissue, we performed Gene Ontology (GO) enrichment analysis, which contains biological processes (BP), cellular components (CC), and molecular functions (MF). Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis was used to identify pathways in which differentially methylated lncRNAs may be involved. There was little difference in m6A modification of lncRNAs between P7 and P28 group, so we focused on P7 vs P1 and P28 vs P1 for analysis.
P7 vs P1: GO enrichment analysis showed that lncRNAs with up-methylated m6A sites were mainly enriched in the cellular response to angiotensin, myosin filament and actin-dependent ATPase activity, whereas lncRNAs with down-methylated sites were mostly enriched in connective tissue development, regulation of organ growth and growth factor binding (Fig. 6A, B). KEGG analysis showed that lncRNAs with up-methylated m6A sites were highly related to ‘Cardiac muscle contraction’ and ‘Adrenergic signaling in cardiomyocytes’. However, lncRNAs with down-methylated sites were significantly enriched in ‘Protein digestion and absorption’ (Fig. 7A, B).
A Top ten GO terms of biological processes(BP), cellular components (CC) and molecular functions (MF) were significantly enriched for the hypermethylated lncRNAs in postnatal day 7 (P7) vs P1. B Top ten GO terms of BP, CC and MF were significantly enriched for the hypomethylated lncRNAs in P7 vs P1. C Top ten GO terms of BP, CC and MF were significantly enriched for the hypermethylated lncRNAs in P28 vs P1. D Top ten GO terms of BP, CC and MF were significantly enriched for the hypomethylated lncRNAs in P28 vs P1; P < 0.05 (n = 3) (Hypergeometric test).
A Bar plot showing the top ten significant enrichment pathway for the hypermethylated lncRNAs in postnatal day 7 (P7) vs P1. B Bar plot showing the top ten significant enrichment pathway for the hypomethylated lncRNAs in P7 vs P1. C Bar plot showing the top ten significant enrichment pathway for the hypermethylated lncRNAs in P28 vs P1. D Bar plot showing the top ten significant enrichment pathway for the hypomethylated lncRNAs in P28 vs P1.
P28 vs P1: GO enrichment data showed that lncRNAs with up-methylated m6A sites were especially enriched in regulation of alternative mRNA splicing via spliceosome, secretory granule membrane and SH2 domain binding. Impressively, lncRNAs with down-methylated sites were especially enriched in insulin-like growth factor receptor signaling pathway and response to thyroid hormone (Fig. 6C, D). It has been demonstrated that the insulin-like growth factor pathway is intricately related to hypertrophy [36] and thyroid hormone was responsible for murine MYH7 (beta-MHC, myosin heavy chain)-to-MYH6 (alpha-MHC) cardiac myosin isoform switch in early postpartum development [37].
Moreover, data from KEGG analysis showed that ‘Nicotinate and nicotinamide metabolism’ and ‘Renin-angiotensin system’ were significantly enriched in lncRNAs with up-methylated m6A sites. The ‘Nicotinate and nicotinamide’ are intricately related, among others, to glycolysis. Previous studies have reported that nicotinamide can stimulate glycolysis in cardiomyocytes [38], and glycolysis was heavily associated with cardiac regenerative ability within “hypoxic cardiomyocyte niches” [39]. Notably, postpartum relative hyperoxia inhibited glycolysis leading to halt regenerative ability [40], while hypoxia-induced anaerobic metabolism elevated glycolysis to promote regenerative milieu within the myocardium [41,42,43]. Angiotensin was known to promote interstitial cardiac fibrosis and hypertrophy [44]. In contrast, lncRNAs with down-methylated sites were involved in ‘Cardiac muscle contraction’ and ‘Cell cycle-yeast’ (Fig. 7C, D).
Discussion
It has been established that the mouse myocardium poses an inherent capacity for major regeneration. However, this capacity is operative only during the first week after birth [7]. After which, its regenerative capability was lost and that may likely be due to the underlying mechanisms of ex utero relative hyperxia-induced DNA damage, cardiac polyploidy, multi-levelled early innate immune system, “cancer risk” suppression mechanism and cardiac thyroxin signaling activation [9,10,11,12,13, 40]. Interestingly, m6A modification, as the most common modification in mRNA and lncRNA, is involved in a variety of biological processes [14, 15]. However, m6A modification within lncRNA in the neonatal mouse heart still remains unexplored territory. In this study, we performed m6A-seq and RNA-seq to sequence the genome-wide profiling of methylation-modified lncRNAs in the heart of P1, P7 and P28 mice, investigated differences between these three developmental stages and found a potential role for m6A methylation of lncRNAs in cardiac development.
During myocardial injury, such as myocardial infarction (MI), cardiomyocytes in extremis may be highly susceptible to arrhythmias due to metabolic disturbances, and necrosis may lead to acute cardiac dysfunction. Then fibrosis of the damaged area occurs in the following days to weeks, which, depending on the extent of the resulting scar, often leads to heart failure as a late-stage complication [45]. For example, there are 2 to 4 billion (3.2 ± 0.75 billion) cardiomyocytes in the human left ventricle [6], and MI can destroy 25% of these cardiomyocytes depending on the occlusion site and revascularization time [46]. Therefore, identifying methods to promote the proliferation of cardiomyocytes in the area where cardiomyocytes are lost is an urgent medical need. Activating proliferation of cardiomyocytes may be an attractive approach to repair myocardial injury caused by MI or other heart diseases [47]. Although regenerative approaches are receiving significant attention, the mechanism underpinning the regulation of myocardial regeneration is complex and still remains incompletely understood.
m6A modification changes appear dynamically at different stages of organ development, especially in mammalian hearts, suggesting that m6A methylation plays an important role in heart growth [48, 49]. Additionally, accumulating evidence showed that the m6A level is closely related to the development of heart disease; for example, METTL3 overexpression can induce eccentric remodeling and cardiac dysfunction alone without an additional stressor [19]. m6A is also involved in the process of myocardial regeneration [16,17,18]. It has previously been documented that compared with normal heart tissue, the level of m6A modification is higher and FTO expression level is lower in myocardium tissue of mice with ischemic damage and heart failure [50]. In a MI mouse model, FTO overexpression can reduce myocardial fibrosis, enhance angiogenesis, and improve cardiac function [50]. Another interesting new finding was that m6A modification enhanced the recruitment of miR-133a-RISC (RNA-induced silencing complexes)-AGO2 (Argonaute 2-Insulin-like growth factor 2) to its targets in heart development and in response to cardiac hypertrophy and proliferation [51]. The above evidence presented thus far supports the idea that m6A may become a new target for the treatment of cardiovascular diseases in the future. There is already strong evidence of the importance of lncRNAs in the regulation of cardiomyocyte proliferation. Indeed, lnc CPR and Sirt1 AS lncRNA has previously been shown to regulate cardiomyocyte proliferation in vitro and vivo [52, 53]. The effects of two lncRNAs lnc CAREL and lnc DACH1 in cardiomyocytes proliferation were also investigated in induced pluripotent stem cells (iPSC)-derived cardiomyocytes [25, 54]. Nevertheless, whether m6A modification can regulate myocardial regeneration through controlling the expression of myocardial regeneration related lncRNAs still remains unknown. Therefore, exploring the interaction between m6A and lncRNAs in cardiac development will improve our understanding the mechanism of myocardial regeneration.
We identified methylation peaks on lncRNAs and found significant differences in the distribution and abundance of methylation peaks between P1 and P7, but such difference was not detectable at the later stage after P7 at P28. Indeed, here we found that the most prominent reorganization in the m6A peak distribution within lncRNAs occurred at across the 3′UTR and promoter regions. Namely, the relative fraction of m6A peaks from the promoter region at P1 was reduced and the fraction of m6A peaks at 3′UTR was increased at P7. Considering that the m6A in 3′UTR has been shown to regulate the stability of the respective RNA [55], the difference in the distribution of m6A modification in the 3′UTR region may affect the stability of lncRNAs during heart development to control myocardial regeneration ability. Recent studies have reported that lncRNAs harboring open reading frame (ORF) sequences can encode proteins/peptides [56]. For example, LINC00998 was originally described as a non-coding transcript, but a 180 nucleotide (nt) smORF (small ORF) was found in exon 3, which encoded a small integral membrane protein 30 (SMIM30) [57]. However, we discovered that the length of the exons contained in the m6A-modified lncRNA has no significant difference during heart development. Therefore, we speculated that the m6A modification may act principally by regulating the stability of lncRNAs, but has little effect on the protein-encoding process of lncRNAs. The exact m6A reader involved in the stabilization and degradation of lncRNA and the functional role of these several unveiled m6A modified lncRNA are unknown and all these need to be explored further in vivo cardiac regeneration models in the future. In addition to the distribution of m6A methylation, there are also great difference in the m6A abundance of the P1, P7 and P28 heart. Indeed, we noticed a near-total reorganization of m6A peaks within lncRNAs from P1 to P7 myocardium with only one conserved peak within lncRNA Aopep. In addition, with more than 600 conserved lncRNA m6A peaks, such major reorganization was not anymore detectable within the later stages when P7 and P28 were compared, suggesting that the methylome remains highly redundant but it was shrinking with an age increase (Fig. 2A–C). Notably, we found a major m6A peak at a rough locus of 50–55kB within chromosome 14 at P1, which disappeared at P7 and P28 (Fig. 2F). Through the analysis of the data, the decrease of the m6A level of lncRNA Myh6 and lncRNA Myh7 may be responsible for the disappearance of this single m6A peak (Supplemental Table S5). Myh6 was primarily expressed in the cardiomyocytes in adult mice but Myh7 was expressed in embryonic cardiomyocytes [37]. Hypertrophy of adult hearts is also associated with Myh6 downregulation and Myh7 induction, returning to a fetal state of MHC expression and thus controlling the expression of MHC may be an attractive approach for heart failure therapy [58]. It has been reported that inhibition of thyroid hormone activaty prolonged myocardium regenerative ability [13], but the role of myosin isoforms, especially from epitranscriptomic m6A, in this process has not been established yet. Whether m6A modification is involved in the shift from Myh6 to neonatal Myh7 expression needs to be studied further. Taking these above observations into account, during the period of rapid changes of myocardial regeneration ability (P1 to P7), m6A may exert potential influence within the loss of myocardial regenerative capacity after birth.
Subsequently, through combined analysis of the m6A-Seq and RNA-seq data, it revealed the correlation between m6A modification and lncRNA expression level in the developing mouse hearts(P1, P7 and P28). We explored that in P7 vs P1, m6A modification was associated with lncRNA expression levels to some extent (Fig. 3A). However, no association was seen between m6A modification and lncRNA expression level in P28 vs P1 and P7 vs P28 (Fig. 3B, C). We speculate that m6A modifications may affect the expression of certain lncRNAs in the heart from P1 to P7, and that this effect was diminished after P7. Interestingly, our previous study found that the expression of m6A reader, insulin-like growth factor-binding protein 3(IGF2BP3), was decreased gradually after birth [59], and its overexpression was also reported to enhance cardiomyocyte proliferation both in vitro and in vivo [49]. Furthermore, it is very likely that lncRNAs were differently expressed due to differential m6A methylation. Indeed, we found that lncRNA Snhg3 was downregulated while lncRNA Nedd4 was upregulated due to m6A.
We further identified 38 lncRNAs with different m6A modification and lncRNA expression levels in the P7 vs P1 (Fig. 4A, Supplemental Table S6) and 9 of these 38 (Supplemental Table S1) may be potentially related to cardiac development or regeneration or may be the new molecules that potentially regulates myocardial regeneration. After removing lncRNA Ece1, which was not match with the expression change in P1 to P7 of RNA-sequence (Fig. 4B), we performed knock-down METTL3 in neonatal mouse cardiomyocytes to verify the expression of the remaining 8 lncRNAs, there were 6 lncRNAs having notable changes in the expression level (Fig. 4C, D). We chose the lncRNA with the largest increase in expression (lncRNA Snhg3) and the lncRNA with the largest decrease in expression (lncRNA Nedd4) after knockdown of METTL3 to conduct MeRIP-qpcr, and the results of MeRIP-qpcr data showed that the m6A abundances of ‘lncRNA Snhg3’ and ‘ lncRNA Nedd4’ were also decreased (Fig. 4E, F). This interesting finding suggests that m6A modification may indeed affect the expression of the key lncRNAs involved in the cardiac development but the potential underlying role of m6A readers executing such divergent responses still need further study. Recently, ubiquitination as an important protein post-translational modification has been demonstrated to be closely related to cardiovascular disease [60], and Nedd4 as the key enzyme in ubiquitination has been found to be involved in the regulation of myocardial regeneration and repair [31]. According to a recent study, USP12 (ubiquitin-specific protease 12) via enhancing p300/MTLL3 axis promote myocardial hypertrophy [61], and our finding may indicate the possibility of m6A methylation and ubiquitination cooperatively regulate heart growth. We also found that lncGm15328 and lncRNA Zfp597 may partake the loss of myocardial regeneration ability during mouse heart development from P1 to P7 (Fig. 5), and that m6A modification may also be involved in this process by regulating the expression levels of these two lncRNAs. The function of these two lncRNAs in heart growth has not been reported and may have great potential as the new targets for regulating myocardial regeneration, but warrants further study with MI models.
Additionally, GO and KEGG analysis revealed that both the hypermethylated and the hypomethylated lncRNAs (P7 vs P1 and P28 vs P1 group) were involved in many important biological functions and pathways. For instance, the GO results showed that hyper/hypomethylated lncRNAs may enrich in ‘regulation of organ growth’, ‘muscle organ development’ and ‘response to thyroid hormone’ functions. It was reported that the loss of heart regenerative capacity was triggered by increasing thyroid hormones [13]. In addition, the analysis indicated that some m6A differential lncRNAs were also enriched in ‘insulin-like growth factor receptor’ and ‘response to angiotensin’, which are participating in cardiac hypertrophy [36, 44]. m6A differential lncRNAs enriched in myocardial regeneration-related pathways, such as ‘TGF-beta signaling pathway’ is well documented [62]. All the above indicates that m6A differential lncRNAs may play an important role in the heart normal or dysfunction development.
In conclusion, we have provided an overall framework for m6A-lncRNA-heart development, and our data provides stagewise views into early postnatal myocardial N6-methyladenosine (m6A) methylomes specifically in long non-coding RNAs (lncRNAs), which, when correlated with the known simultaneously occurring loss of inherent cardiac regenerative capacity, were appropriately pointed out to be of interest regarding possible functional roles with the cardiac regeneration capacity. These findings identify potential m6A methylated lncRNA targets for future experimental heart regeneration-targeted studies.
Materials and methods
Animals studies
The animal study was reviewed and approved by the institutional committee of animal care and use of the Affiliated First Hospital of Guangdong Medical University (Guangdong, China). C57BL/6 mice, Male, aged 1-day-old, 7-day-old and 28-day-old, were purchased from the Experimental Animal Center of Southern Medical University (Guangdong, China). Under the premise of meeting the inclusion criteria, we randomly selected mice from different groups for heart extraction and sequencing. We assigned three groups in line with different ages (P1, P7, and P28) and collected three biological replicates (n = 3–5 mice per group) among which none was excluded. There was no blind selection involved. All mice in both groups were appropriately anesthetized with Ketamine (80 mg/kg, IP) + Xylazine (10 mg/kg, IP) and sacrificed by cervical dislocation. Their cardiac tissue was immediately collected, frozen in liquid nitrogen and stored at −80 °C for later preparation of total RNA.
Total RNA preparation
Total RNA was harvested and extracted from tissue samples using Trizol Reagent (Thermo Fisher Scientific, MA, USA). The accurate concentration and sample purity were detected through NanoDropND-2000 (Thermo Fisher Scientific, MA, USA). Finally, the degradation of total RNA was determined by agarose gel electrophoresis and Agilent 2100 Bioanalyzer (Agilent Technologies Inc, CA, USA). Only RIN (RNA integrity number)>7 of extracted RNA was used to ensure downstream high-quality total RNA-seq library construction (Supplemental Table S7). Qualified RNA acquisition carried out the above series of quality control (QC) processes.
lncRNAs Library Construction and sequencing
In brief, according to the manufacturer’s instructions, the total RNA was used for removing the rRNAs with Ribo-off rRNA Depletion Kit (H/M/R) (Vazyme Biotech, China) and purified by AMPure XP magnetic beads (Beckman Coulter, CA, USA). ABI 2720 Thermal Cycler (Thermo Fisher Scientific, USA) was used to construct RNA libraries using fragmented rRNA-depleted RNAs. The library concentration was accurately quantified by Qubit and the size distribution of library fragment was determined by Agilent 2100 Bioanalyzer (Agilent Technologies Inc, CA, USA). Then, the libraries were captured on Illumina cbot Cluster Station (Illumina, CA, USA) and finally sequenced and visualized for corresponding cycles on Illumina Hiseq 2500 (Illumina, CA, USA).
lncRNAs methylation-RNA immunoprecipitation (MeRIP) library construction and sequencing
MeRIP-Seq was based on previously published procedures [28]. Briefly, fragmented RNA was incubated with immunomagnetic beads premixed anti-m6A antibody. The mixture was then immunoprecipitated by incubation with protein-A beads. Next, purified RNA was used for the RNA-seq library by Illumina Hiseq 2500 (Illumina, CA, USA). The input sample without immunoprecipitation and the m6A IP samples were subjected to PE150 paired-end sequencing InIllumina Novaseq™ 6000. After removal of ribosomal sequences, the percentage of data quality values greater than Q30 was more than 90%. Finally, methylated sites on RNAs (peaks) were identified by the ChiPseeker package [63].
Sequencing Data Analysis
To identify lncRNAs, the reported databases and software (EggNOG, CNCI, Pfam, CPC2) were applied based on the noncoding potential property of lncRNAs. The up- or downregulated expression of lncRNAs was set at absolute fold change (FC > 1.5, and P < 0.05). For m6A sequencing, methylated sites on lncRNAs (m6A peaks) were identified with the diffReps differential analysis package [64] with differential fold change evaluated of > 1.5 and a P value < 0.05. The correlation analysis between the lncRNAs (lncRNAs with changes in m6A modification) and co-expression mRNAs was evaluated using Pearson correlation by SPSS software (v22.0)
GO and KEGG Pathway Databases Analysis
LncRNAs expression with differentially methylated profiles were compared to functional differences between-group variance using enrichment analysis. Gene ontology (GO) was performed to annotate these genes. The functions were distinguished into three parts: cellular component (CC), molecular function (MF), and biological process (BP). The p value denotes the significance of GO term enrichment of the genes. In addition, Pathway enrichment analysis is a functional analysis that maps genes to the Kyoto Encyclopedia of Genes and Genomes (KEGG). The Fisher p-value denotes the significance of the pathway correlated to the conditions [65, 66].
Cardiomyocyte Isolation and Culture
Neonatal cardiomyocytes were isolated from 1 day-old (P1) and 7-day-old (P7) C57BL/6 mice by enzymatic dissociation. Shortly, P1 or P7 hearts were rapidly obtained and removed mostly blood in ice-cold PBS (C10010500BT, Gibco, USA) before being digested with trypsin enzyme (25200056, Gibco, USA). The separated cells were cultured in Gelatin-dealt wells (G8061, Solarbio, China) with 10% FBS DMEM medium (11995-065, Gibco, USA) supplemented with L-glutamine, 1% Antibiotics and 5% CO2 at 37 °C [67].
Transfection of Mettl3 siRNA (5′-GCUACCGUAUGGGCACUUATT-3′)/scrambled controls (RiboBio, Guangzhou, China) in cardiomyocytes was using Lipofectamine RNAiMAX reagent (Invitrogen, CA, USA) following the manufacturer’s instructions. Cardiomyocytes from neonatal heart were transfected with si-NC or si-METTL3 for 72 h, then the expression of METTLE3 and lncRNAs were determined.
Quantitative Real-time PCR
Total RNA isolation from tissue and the cultured primary cardiomyocytes were extracted with Trizol Reagent (Thermo Fisher Scientific, MA, USA). The cardiomyocyte lncRNA were measured by RT-qPCR. The ratio of OD260/280 to OD260/230 of the extracted RNA were detected by NanoDrop (Thermo Fisher Scientific, Waltham, MA, USA) as the samples purity index. If the OD260/280 value was between 1.8 and 2.2, OD260/230 ≥ 2.0, the RNA purity and integrity were suitable for downstream experiments. Quantitative reverse transcription-polymerase chain reaction (qRT-PCR) was performed on LightCycler 480 System (Roche, Germany) or Applied Biosystem 7500 qPCR system (Applied Biosystems,USA) by using SYBR Green PCR Master Mix (Thermo Fisher Scientific, MA, USA) and the prepared cDNAs.
Western blot analysis
Total protein was extracted with RIPA lysis buffer, and the protein concentrations were determined by enhanced bicinchoninic acid (BCA) protein assay kit (Beyotime, Shanghai, China). Furthermore, the equivalent amounts of protein were separated by SDS-PAGE on 10% acrylamide gels and transferred onto polyvinylidene fluoride (PVDF) membranes, blocked with 5% milk, and then incubated with primary target antibodies. The primary antibodies were anti-METTL3 (ab195352, Abcam, UK) and anti-GAPDH (AP0063, Bioword, USA). As the secondary antibody was goat polycolonal anti-Rabbit-IgG (14708 S, CST, USA). Quantitative analysis were performed by Image J software (NIH, Bethesda, USA).
m6A-RIP qPCR
To examine m6A modifications on individual genes, the MeRIP m6A Kit (GS-ET-001, CouldSeq, China) was used according to the manufacturer’s instructions. RNA in a certain ratio was severed as input control, and further analyzed by qPCR with the primers (Supplemental Table S1). The related enrichment of m6A in goal genes was calculated by normalizing the value of amplification cycle to the corresponding input portion.
Constructing the ceRNA network
The lncRNA-miRNA-mRNA co-expression network was visualized with the Cytoscape software (http://www.cytoscape.org/). Using the miRDB (Version5.0; http://mirdb.org) to predict the miRNAs targeted by lncRNAs. mRNAs targeted by the miRNAs were retrieved from the miRDB (Version 5.0; http://mirdb.org), miRTarBase (Version7.0; http://mirtarbase.mbc.nctu.edu.tw/), and TargetScan (Version 7.2; http://www.targetscan.org/vert_72/) databases [68,69,70].
Statistical analysis
All data were expressed as the mean ± SEM (standard error mean) and analyzed with Student t-test or hypergeometric test (Prism 9.0, GraphPad Software Inc, San Diego, CA). A P value less than 0.05 was considered to be of statistical significance.
Code availability
All computer codes were used to generate results during this study are available from the corresponding authors on reasonable request.
Data availability
The datasets generated and analyzed during the current study are available in the NCBI (https://www.ncbi.nlm.nih.gov/home/download/) repository, and BioProject ID: PRJNA828395.
References
Yuan X, Braun T. Multimodal Regulation of Cardiac Myocyte Proliferation. Circulation Res. 2017;121:293309.
Takeuchi T. Regulation of cardiomyocyte proliferation during development and regeneration. Nat Rev Cardiol 2014;56:402409.
Haubner BJ, Schneider J, Schweigmann U, Schuetz T, Dichtl W, Velik-Salchner C, et al. Functional Recovery of a Human Neonatal Heart After Severe Myocardial Infarction. Circ Res. 2016;118:216–21.
Azakie A, Russell JL, McCrindle BW, Van Arsdell GS, Benson LN, Coles JG, et al. Anatomic repair of anomalous left coronary artery from the pulmonary artery by aortic reimplantation: early survival, patterns of ventricular recovery and late outcome. Ann Thorac Surg. 2003;75:1535–41.
Bergmann O, Bhardwaj RD, Bernard S, Zdunek S, Barnabe-Heider F, Walsh S, et al. Evidence for cardiomyocyte renewal in humans. Science. 2009;324:98–102.
Bergmann O, Zdunek S, Felker A, Salehpour M, Alkass K, Bernard S, et al. Dynamics of Cell Generation and Turnover in the Human Heart. Cell. 2015;161:1566–75.
Porrello ER, Mahmoud AI, Simpson E, Hill JA, Richardson JA, Olson EN, et al. Transient regenerative potential of the neonatal mouse heart. Science. 2011;331:1078–80.
Mahmoud AI, Porrello ER, Kimura W, Olson EN, Sadek HA. (2014). Surgical models for cardiac regeneration in neonatal mice. Nat Protoc. 2014;9:305–11.
Derks W, Bergmann O. Polyploidy in Cardiomyocytes: Roadblock to Heart Regeneration? Circ Res. 2020;126:552–65.
Han C, Nie Y, Lian H, Liu R, He F, Huang H, et al. Acute inflammation stimulates a regenerative response in the neonatal mouse heart. Cell Res. 2015;25:1137–51.
D’Uva G, Aharonov A, Lauriola M, Kain D, Yahalom-Ronen Y, Carvalho S, et al. ERBB2 triggers mammalian heart regeneration by promoting cardiomyocyte dedifferentiation and proliferation. Nat Cell Biol. 2015;17:627–38.
Lindgren IM, Drake RR, Chattergoon NN, Thornburg KL. Down-regulation of MEIS1 promotes the maturation of oxidative phosphorylation in perinatal cardiomyocytes. FASEB J. 2019;33:7417–26.
Hirose K, Payumo AY, Cutie S, Hoang A, Zhang H, Guyot R, et al. Evidence for hormonal control of heart regenerative capacity during endothermy acquisition. Science 2019;364:184–8.
Wang X, He C. Dynamic RNA modifications in posttranscriptional regulation. Mol Cell. 2014;56:5–12.
Chelmicki T, Roger E, Teissandier A, Dura M, Bonneville L, Rucli S, et al. m(6)A RNA methylation regulates the fate of endogenous retroviruses. Nature. 2021;591:312–6.
Yang C, Zhao K, Zhang J, Wu X, Sun W, Kong X, et al. Comprehensive Analysis of the Transcriptome-Wide m6A Methylome of Heart via MeRIP After Birth: Day 0 vs. Day 7. Front Cardiovascular Med. 2021;8:633631.
Gong R, Wang X, Li H, Liu S, Jiang Z, Zhao Y, et al. Loss of m6A methyltransferase METTL3 promotes heart regeneration and repair after myocardial injury. Pharm Res. 2021;174:105845.
Han Z, Wang X, Xu Z, Cao Y, Gong R, Yu Y, et al. ALKBH5 regulates cardiomyocyte proliferation and heart regeneration by demethylating the mRNA of YTHDF1. Theranostics. 2021;11:3000–16.
Dorn LE, Lasman L, Chen J, Xu X, Hund TJ, Medvedovic M, et al. The N6-Methyladenosine mRNA Methylase METTL3 Controls Cardiac Homeostasis and Hypertrophy. Circulation. 2019;139:533–45.
Engreitz JM, Ollikainen N, Guttman M. Long non-coding RNAs: spatial amplifiers that control nuclear structure and gene expression. Nat Rev Mol Cell Biol. 2016;17:756–70.
Statello L, Guo CJ, Chen LL, Huarte M. Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol. 2021;22:96–118.
Meloni M, Riley PR, Baker AH. A new “lnc” between non-coding RNA and cardiac regeneration. Cardiovasc Res. 2018;114:1569–70.
Ounzain S, Crippa S, Pedrazzini T. Small and long non-coding RNAs in cardiac homeostasis and regeneration. Biochim Biophys Acta. 2013;1833:923–33.
Philippen LE, Dirkx E, Da CP, De Windt LJ. Non-coding RNA in control of gene regulatory programs in cardiac development and disease. J Mol Cell Cardiol. 2015;89:51–58.
Cai B, Ma W, Ding F, Zhang L, Huang Q, Wang X, et al. The Long Noncoding RNA CAREL Controls Cardiac Regeneration. J Am Coll Cardiol. 2018;72:534–50.
Chen Y, Li X, Li B, Wang H, Li M, Huang S, et al. Long Non-coding RNA ECRAR Triggers Post-natal Myocardial Regeneration by Activating ERK1/2 Signaling. Mol Ther. 2019;27:29–45.
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–6.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149:1635–46.
Han P, Li W, Lin CH, Yang J, Shang C, Nuernberg ST, et al. A long noncoding RNA protects the heart from pathological hypertrophy. Nature. 2014;514:102–6.
Kelly MA, Caleshu C, Morales A, Buchan J, Wolf Z, Harrison SM, et al. Adaptation and validation of the ACMG/AMP variant classification framework for MYH7-associated inherited cardiomyopathies: recommendations by ClinGen’s Inherited Cardiomyopathy Expert Panel. Genet Med. 2018;20:351–9.
Huang S, Li X, Zheng H, Si X, Li B, Wei G, et al. Loss of Super-Enhancer-Regulated circRNA Nfix Induces Cardiac Regeneration After Myocardial Infarction in Adult Mice. Circulation. 2019;139:2857–76.
Thomson DW, Dinger ME. Endogenous microRNA sponges: evidence and controversy. Nat Rev Genet. 2016;17:272–83.
Verjans R, van Bilsen M, Schroen B. Reviewing the Limitations of Adult Mammalian Cardiac Regeneration: Noncoding RNAs as Regulators of Cardiomyogenesis. Biomolecules. 2020;10:262.
Gao F, Kataoka M, Liu N, Liang T, Huang ZP, Gu F, et al. Therapeutic role of miR-19a/19b in cardiac regeneration and protection from myocardial infarction. Nat Commun. 2019;10:1802 https://doi.org/10.1038/s41467-019-09530-1
Lin X, Peng P, Cheng L, Chen S, Li K, Li ZY, et al. A natural compound induced cardiogenic differentiation of endogenous MSCs for repair of infarcted heart. Differentiation. 2012;83:1–9.
Friberg P, Isgaard J, Wåhlander H, Wickman A, Guron G, Adams MA. Cardiac hypertrophy and related growth processes: the role of insulin-like growth factor-I. Blood Press Suppl. 1995;2:22–9.
Morkin E. Control of cardiac myosin heavy chain gene expression. Microsc Res Tech. 2000;50:522–31.
Diguet N, Trammell SAJ, Tannous C, Deloux R, Piquereau J, Mougenot N, et al. Nicotinamide Riboside Preserves Cardiac Function in a Mouse Model of Dilated Cardiomyopathy. Circulation. 2018;137:2256–73.
Kimura W, Sadek HA. The cardiac hypoxic niche: emerging role of hypoxic microenvironment in cardiac progenitors. Cardiovasc Diagn Ther. 2012;2:278–89.
Puente BN, Kimura W, Muralidhar SA, Moon J, Amatruda JF, Phelps KL, et al. The oxygen-rich postnatal environment induces cardiomyocyte cell-cycle arrest through DNA damage response. Cell 2014;157:565–79.
Nakada Y, Canseco DC, Thet S, Abdisalaam S, Asaithamby A, Santos CX, et al. Hypoxia induces heart regeneration in adult mice. Nature 2017;541:222–7.
Kimura W, Xiao F, Canseco DC, Muralidhar S, Thet S, Zhang HM, et al. Hypoxia fate mapping identifies cycling cardiomyocytes in the adult heart. Nature 2015;523:226–30.
Xie Y, Lampinen M, Takala J, Sikorski V, Soliymani R, Tarkia M, et al. Epicardial transplantation of atrial appendage micrograft patch salvages myocardium after infarction. J Heart Lung Transpl. 2020;39:707–18.
Adamcova M, Kawano I, Simko F. The Impact of microRNAs in Renin-Angiotensin-System-Induced Cardiac Remodelling. Int J Mol Sci. 2021;22:4762.
Reed GW, Rossi JE, Cannon CP. Acute myocardial infarction. Lancet. 2017;389:197–210.
Caulfield JB, Leinbach R, Gold H. The relationship of myocardial infarct size and prognosis. Circulation 1976;53:I141–4.
Karra R, Poss KD. Redirecting cardiac growth mechanisms for therapeutic regeneration. J Clin Invest. 2017;127:427–36.
Zhang H, Shi X, Huang T, Zhao X, Chen W, Gu N, et al. Dynamic landscape and evolution of m6A methylation in human. Nucleic Acids Res. 2020;48:6251–64.
Wang Z, Cui M, Shah AM, Ye W, Tan W, Min YL, et al. Mechanistic basis of neonatal heart regeneration revealed by transcriptome and histone modification profiling. Proc Natl Acad Sci USA. 2019;116:18455–65.
Mathiyalagan P, Adamiak M, Mayourian J, Sassi Y, Liang Y, Agarwal N, et al. FTO-Dependent N(6)-Methyladenosine Regulates Cardiac Function During Remodeling and Repair. Circulation. 2019;139:518–32.
Qian B, Wang P, Zhang D, Wu L. m6A modification promotes miR-133a repression during cardiac development and hypertrophy via IGF2BP2. Cell Death Disco. 2021;7:157.
Ponnusamy M, Liu F, Zhang Y, Li R, Zhai M, Liu F, et al. Long Noncoding RNA CPR (Cardiomyocyte Proliferation Regulator) Regulates Cardiomyocyte Proliferation and Cardiac Repair. Circulation. 2019;139:2668–84.
Li B, Hu Y, Li X, Jin G, Chen X, Chen G, et al. Sirt1 Antisense Long Noncoding RNA Promotes Cardiomyocyte Proliferation by Enhancing the Stability of Sirt1. J AM Heart Assoc. 2018;7:e9700.
Cai B, Ma W, Wang X, Sukhareva N, Hua B, Zhang L, et al. Targeting LncDACH1 promotes cardiac repair and regeneration after myocardium infarction. Cell Death Differ. 2020;27:2158–75.
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505:117–20.
Kong S, Tao M, Shen X, Ju S. Translatable circRNAs and lncRNAs: Driving mechanisms and functions of their translation products. Cancer Lett. 2020;483:59–65.
Pang Y, Liu Z, Han H, Wang B, Li W, Mao C, et al. Peptide SMIM30 promotes HCC development by inducing SRC/YES1 membrane anchoring and MAPK pathway activation. J Hepatol. 2020;73:1155–69.
McKinsey TA, Olson EN. Toward transcriptional therapies for the failing heart: chemical screens to modulate genes. J Clin Invest. 2005;115:538–46.
Yang Y, Shen S, Cai Y, Zeng K, Liu K, Li S, et al. Dynamic Patterns of N6-Methyladenosine Profiles of Messenger RNA Correlated with the Cardiomyocyte Regenerability during the Early Heart Development in Mice. Oxid Med Cell Longev. 2021;2021:5537804.
Martins-Marques T, Ribeiro-Rodrigues T, Pereira P, Codogno P, Girao H. Autophagy and ubiquitination in cardiovascular diseases. DNA Cell Biol. 2015;34:243–51.
Lu P, Xu Y, Sheng ZY, Peng XG, Zhang JJ, Wu QH, et al. De-ubiquitination of p300 by USP12 Critically Enhances METTL3 Expression and Ang II-induced cardiac hypertrophy. Exp Cell Res. 2021;406:112761.
Sorensen DW, van Berlo JH. The Role of TGF-β Signaling in Cardiomyocyte Proliferation. Curr Heart Fail Rep. 2020;17:225–33.
Yu G, Wang LG, He QY. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 2015;31:2382–3.
Shen L, Shao NY, Liu X, Maze I, Feng J, Nestler EJ. diffReps: detecting differential chromatin modification sites from ChIP-seq data with biological replicates. PLoS One. 2013;8:e65598.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–29.
Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13.
Etiology and Morphogenesis of Congenital Heart Disease. From Gene Function and Cellular Interaction to Morphology. Tokyo: Springer; 2016.
Wong N, Wang X. miRDB: an online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 2015;43:D146–D152.
Chou CH, Shrestha S, Yang CD, Chang NW, Lin YL, Liao KW, et al. miRTarBase update 2018: a resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018;46:D296–D302.
Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. Elife. 2015;4:e05005.
Acknowledgements
This work was supported by the Science and Technology Planning Project of Guangdong Province of China (2014A020212301) and the National Natural Science Fund of China (81870222).
Author information
Authors and Affiliations
Contributions
LZ and DM conceived the study. SS, SL and KL performed most of the bioinformatics analysis and wrote the manuscript. SR, ZX and JT helped project design and manuscript editing. YY and ZH performed the validation of this study. YH collected the mice heart samples. All authors participated manuscript writing and approved submission for a publication.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
The animal study was reviewed and approved by the institutional committee of animal care and use of the Affiliated First Hospital of Guangdong Medical University (Guangdong, China)
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shen, S., Liu, K., Li, S. et al. N6-methyladenosine modulates long non-coding RNA in the developing mouse heart. Cell Death Discov. 8, 329 (2022). https://doi.org/10.1038/s41420-022-01118-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-01118-x
This article is cited by
-
Effect of tumor exosome-derived Lnc RNA HOTAIR on the growth and metastasis of gastric cancer
Clinical and Translational Oncology (2023)
-
Circular RNAs in organ injury: recent development
Journal of Translational Medicine (2022)