Abstract
Tissue macrophages in many adult organs originate from yolk sac (YS) progenitors, which invade the developing embryo and persist by means of local self-renewal. However, the route and characteristics of YS macrophage trafficking during embryogenesis are incompletely understood. Here we show the early migration dynamics of YS-derived macrophage progenitors in vivo using fate mapping and intravital microscopy. From embryonic day 8.5 (E8.5) CX3CR1+ pre-macrophages are present in the mouse YS where they rapidly proliferate and gain access to the bloodstream to migrate towards the embryo. Trafficking of pre-macrophages and their progenitors from the YS to tissues peaks around E10.5, dramatically decreases towards E12.5 and is no longer evident from E14.5 onwards. Thus, YS progenitors use the vascular system during a restricted time window of embryogenesis to invade the growing fetus. These findings close an important gap in our understanding of the development of the innate immune system.
Similar content being viewed by others
Introduction
Macrophages are important effectors of innate immunity. They have a critical function in organ development, maintenance of tissue homeostasis and host protection during infection1. Furthermore, they have distinct functions in chronic inflammatory diseases such as atherosclerosis, metabolic disorders and cancer2,3,4,5.
Historically, macrophages were believed to originate exclusively from bone marrow (BM)-derived monocytes after their recruitment to tissues. However, first macrophage progenitors develop in the YS, the first site of hematopoiesis in mammals6. Fate mapping analyses and lineage tracing experiments have indicated that tissue macrophages in many organs are of early embryonic origin7,8,9,10,11. Macrophages have been proposed to develop in the YS in two successive waves. First, primitive macrophage progenitors appear around E7.25 in the YS and predominantly infiltrate the brain after the onset of circulation7,12,13,14. Second, multipotent erythro-myeloid progenitors (EMP) develop from E8.25 onwards and are a major source of tissue resident macrophages6,15,16. Substantially later, starting from E10.5, hematopoietic stem cells (HSC) arise in the aorto-gonado-mesonephros (AGM) region and migrate to the liver to initiate fetal definitive hematopoiesis, which later shifts to the BM17,18,19. In adult life, monocytes and other short-lived myeloid cells are continuously replaced, whereas tissue macrophages can self-renew and persist independently of definitive BM hematopoiesis for prolonged periods of time, potentially throughout life8,16,20. However, even though macrophage trafficking during early embryonic development is a prerequisite for establishing this important component of the innate immune system, the kinetics of this process have not been defined.
EMPs are characterized by expression of the receptor tyrosine kinase KIT as well as the macrophage colony-stimulating factor 1 receptor (CSF1R). Subsequent differentiation into pre-macrophages is indicated by expression of the fractalkine receptor CX3CR1, whereas mature tissue resident macrophages additionally express F4/8012,21,22,23. Thus, these markers allow genetic labeling and visualization of macrophage progenitors early during embryonic development.
In this study, we present in vivo data of EMP and pre-macrophage trafficking and proliferation in mouse YS and embryo. We identify a restricted time frame in which these cells travel from the YS to infiltrate the embryo via the bloodstream before becoming phenotypically mature macrophages within their tissue of residence.
Results
Intravital microscopy of YS macrophages
To visualize the early development of macrophages in the mouse YS (Fig. 1a–f) and their trafficking to the embryo (Figs. 1g–k and 2a–f) we established an in vivo imaging strategy (Fig. 1b, c; Methods).
Intravascular trafficking of CX3CR1+ YS pre-macrophages. a, b Schematic graphs of the mouse model (a) and the intravital imaging setup in Cx3cr1GFP/+ mice (b). c E16.5 Cx3cr1GFP/+ embryo with surrounding YS and its dense vascular network. d YS tissue with CX3CR1+ pre-macrophages at indicated time points. Pictures show a representative microscopic field of 400 × 400 µm. e Corresponding quantification of CX3CR1+ cells per microscopic field of 400 × 400 µm; *** p < 0.001 (two-tailed Mann–Whitney test); graph shows median with interquartile range (±IQR, error bars). f In vivo YS staining for the endothelial marker CD31 (red) in Cx3cr1GFP/+ (green) embryos at E10.5. g, h, j Images extracted from E10.5 video sequences of CX3CR1+ YS tissue show a cell entering the vasculature (g, from Supplementary Movie 1) as well as intravascular non-adhering (h, from Supplementary Movie 3) and rolling (j) cells. CX3CR1+ cells in focus are indicated by arrowheads; direction of flow from bottom to top (g), left to right (h), left to right (j). i Quantification of intravascular CX3CR1+ cells in the YS in an average-sized vessel; median ± IQR. k Velocity of intravascular non-adhering (n = 50) and rolling (n = 5) CX3CR1+ cells in the YS at E10.5; median ± IQR. Scale bars are 1 mm (c), 100 µm (d, f, g, h, j)
Pre-macrophages infiltrate embryonic tissues. a Isolated E10.5 Cx3cr1GFP/+ embryo. b, c Visualization (b) and quantification (c) of CX3CR1+ macrophages in different embryonic regions at indicated time points; * p < 0.05 (one-way ANOVA with Tukey’s multiple comparisons test: head vs. trunk p = 0.0107, head vs. tail p = 0.0121, trunk vs. tail p = 0.9860); graph shows median ± IQR. d Quantification by flow cytometry of CX3CR1 GFP+ cells in the embryonic YS, brain and trunk at indicated time points; (**) p = 0.0017 (two-tailed Mann–Whitney test); median ± IQR. e CX3CR1 GFP+ cells (green) in the embryonic brain at E16.5 additionally stained for the endothelial marker CD31 (red). f Image series extracted from live E10.5 video sequences (Supplementary Movie 6) show a cell attaching to the endothelium in the embryonic head region; direction of flow from top to bottom. Scale bars are 1 mm (a), 100 µm (b, e, f)
We first took advantage of Cx3cr1GFP/+ knock-in mice to visualize pre-macrophages and macrophages21. Green fluorescent protein (GFP)-labeling in Cx3cr1GFP/+ embryos corresponds to the CX3CR1+ CD45+ c-KIT- (“A3-like”) population previously characterized in vitro, while other hematopoietic lineages are not labeled (Supplementary Fig. 1a–c)8,12. CX3CR1+ positive cells were not detectable in the YS by epifluorescence microscopy at E8.5; however, a rapidly expanding population of CX3CR1+ pre-macrophages appeared shortly thereafter (Fig. 1d). Macrophage numbers per area increased significantly in tissues during embryogenesis. Maximum cell density was approximately 100 cells per microscopic field (400× 400 µm) in the YS and reached a plateau by E12.5 with stable cell densities thereafter (Fig. 1d, e). In parallel to the increase in cell numbers, CX3CR1+ pre-macrophages underwent morphological changes from spherical shape to mature cells with multiple dendrites (Fig. 1d) potentially engaging in direct cell-to-cell contacts, as it is typically found in adult tissues24. Moreover, pre-macrophages appeared partially associated with vascular structures (Fig. 1f and Supplementary Fig. 1d) and the first detection of CX3CR1+ pre-macrophages was temporally correlated with the formation of a dense vascular network (Fig. 1d, f and Supplementary Fig. 1d, e).
Trafficking of YS macrophage progenitors via the bloodstream
To date two alternate routes have been postulated of how macrophage progenitors enter the embryo, trans-tissue migration or trans-vascular trafficking via the bloodstream7,13,22,25,26,27. To address these two hypotheses, we directly imaged the trafficking routes of YS macrophage progenitors during embryogenesis using intravital microscopy. We observed CX3CR1+ pre-macrophages entering the YS vasculature (Fig. 1g, Supplementary Movies 1, 2) and traveling towards the embryo (Fig. 1h, i, Supplementary Fig. 1g and Supplementary Movies 3–5). The majority of pre-macrophages were trafficking in the YS bloodstream with an average velocity of about 210 µm/s, while some CX3CR1+ cells displayed slow surface translocation (rolling) at significantly lower mean velocity of 25 µm/s (Fig. 1j, k and Supplementary Fig. 1h). On occasion, pre-macrophages re-adhered transiently to YS endothelial structures after their vascular infiltration (Supplementary Movie 2). However, the biological meaning of these transient interactions remains unknown.
Restricted time window of vasculature-mediated infiltration
In vivo microscopy of the YS vasculature showed that CX3CR1+ cells appeared in the bloodstream shortly after the formation of a vascular network around E9 (Fig. 1i and Supplementary Fig. 1e)7,28. Subsequently, we observed a rapid increase in intravascular pre-macrophages trafficking to the embryo up to a maximum of about 30 cells/min at E10.5 in an average-sized vessel of about 60 µm in diameter at this developmental stage (Fig. 1i and Supplementary Fig. 1f, g). The highest number of pre-macrophages trafficking through the bloodstream per minute was observed between E9.5 and E10.5. At E12.5 the number dropped by more than 90% and no trafficking cells were observed at E14.5 (Fig. 1i, Supplementary Movies 3–5).
Yolk sac macrophages in humans
The process of hematopoietic development is conserved between mouse and humans29. In men, the YS serves as the initial site of hematopoiesis and gives rise to granulo-macrophage progenitors30,31. We collected fetal material from week 9 of gestation to determine the phenotype of human YS macrophages. YS macrophages were spherical as their mouse counterparts, while mature macrophages in the placenta presented the typical morphology of mature phagocytes with multiple dendrites (Supplementary Fig. 2a, b). Besides the human macrophage markers macrosialin (CD68) and epidermal growth factor module-containing mucin-like receptor 2 (EMR2, CD312)32,33, macrophages of the human YS expressed CX3CR1 as in mice (Supplementary Fig. 2a, b). Thus, we identified similarities between mouse and human YS macrophages, opening the possibility of analogies in their development. The surface expression of EMR2 is of potential interest, since differential expression of EMR2 has been linked to macrophage maturation33.
Trafficking concurs with vascular network formation
YS CX3CR1+ pre-macrophages are located in close proximity to vascular structures (Fig. 1f and Supplementary Fig. 1d), which is consistent with histological reports34,35. In the embryo, the primary target for YS-derived pre-macrophages was the developing brain, which is also characterized by the early establishment of a vascular network (Fig. 2a–f and Supplementary Movie 6)8,35. Although some CX3CR1+ cells were found in close proximity to the brain vasculature, they distributed throughout the head region forming a dense cellular network (Fig. 2b, e and Supplementary Movie 6). The infiltration of other tissues started thereafter reaching comparable cell densities of about 80 CX3CR1+ cells/ microscopic field at E12.5 (Fig. 2b–d). Macrophage numbers in YS and embryonic tissues remained stable from E12.5, supporting the notion that infiltration of the embryo by tissue macrophage progenitors was mostly completed at this time.
Trafficking is restricted to spherical-shaped macrophages
Next we addressed whether alterations in cell morphology accompanying macrophage maturation were associated with macrophage trafficking. Pre-macrophages in the bloodstream displayed a spherical shape (Supplementary Movies 1–6). In contrast, dendrite-shaped YS macrophages did not enter the circulation or traveled through blood. The findings established with epifluorescence imaging were confirmed by high resolution spinning disc confocal microscopy in Cx3cr1Cre:Rosa26mT/mG embryos (Fig. 3a, b and Supplementary Movie 7). Thus, alterations in cell morphology correlate with restricted trafficking potential of YS macrophages. In line with this, large numbers of matured macrophages with dendrites remained in YS tissue after E12.5 (Fig. 1d, e, i). Interestingly, respective macrophages with mature morphology extended their dendrites into the YS tissue and also into the vessel lumen (Fig. 3c–f and Supplementary Movie 8).
Trafficking is associated with cellular morphology. Intravital spinning disc microscopy of YS vasculature in E10.5 Cx3cr1Cre:Rosa26mT/mG embryos. a Schematic graph for the Cx3cr1Cre:Rosa26mT/mG mouse model. b Image series of CX3CR1 GFP+ (green) YS pre-macrophages; direction of flow from top to bottom (from Supplementary Movie 7). c–f Dendrite-shaped CX3CR1 GFP+ macrophages (green) with intravascular protrusions (c, d) further illustrated by multiplanar reconstructions in XY (e, from Supplementary Movie 8) and orthogonal YZ views including heat map (f); endothelium (mTomato, red). Scale bars are 20 µm (b, c, e), 10 µm (d)
YS macrophage trafficking is independent of MYB and CX3CR1
Trafficking of pre-macrophages from the YS to the embryo took place mostly between E9.5 and E10.5, significantly decreased towards E12.5 and was not observed after E14.5 (Fig. 1i). This raised the possibility that pre-macrophage trafficking was restricted by the onset of fetal definitive hematopoiesis. HSCs arise in the AGM region from E10.5 and are found in the fetal liver by E12, where they produce hematopoietic cells36. We therefore revisited macrophage development in mice lacking fetal definitive hematopoiesis due to genetic absence of the transcription factor MYB (Fig. 4a, b). From E9.5 through E16.5 CX3CR1+ cell numbers and densities in YS and embryonic tissues were not altered in the absence of HSCs and their progeny (Fig. 3c, d and Supplementary Fig. 3a–c)8,22. Using intravital microscopy, we found that numbers and timing of pre-macrophage trafficking from the YS to the embryo in Myb knockout mice (Myb−/−:Cx3cr1GFP/+) were comparable to wild type (Myb+/+:Cx3cr1GFP/+) littermates (Fig. 4e and Supplementary Movies 9, 10). Thus, onset of fetal definitive hematopoiesis did not influence, or even restrict, pre-macrophage trafficking from the YS. These data also showed that most CX3CR1+ cells identified in tissues after the onset of fetal hematopoiesis were derived from YS hematopoiesis but not from MYB-dependent hematopoietic progenitors. This is noteworthy, because CX3CR1 is potentially expressed on various mature blood cells including fetal liver and BM monocytes37, which could have biased our analysis at later stages of embryonic development (i.e. from E12.5). Further, our findings support the notion that YS-derived tissue macrophages are independent of the transcription factor MYB, which is consistent with previous reports8.
Intravascular trafficking is independent of MYB and CX3CR1. a Schematic graph of the Cx3cr1GFP/+ Myb−/− mouse model. b Bright field images of isolated embryos at E16.5 indicating the absence of fetal definitive hematopoiesis (i.e. severe anemia). c, d Visualization (c) and quantification (d) of CX3CR1+ cells in the YS of Myb+/+ or Myb−/− mice at indicated time points; all comparisons of Myb+/+ vs. Myb−/− are n.s. (two-tailed t-test: E9.5 p = 0.9818, E10.5 p = 0.8894, E12.5 p = 0.6900, E16.5 p = 0.9543); mean ± standard deviation (SD, error bars). e Quantification of intravascular CX3CR1+ cells in a Myb+/+ and Myb−/− YS on indicated time points; E10.5: n.s. (two-tailed Mann–Whitney: E10.5 p = 0.4537); median ± IQR. f Quantification of CX3CR1+ cell densities per microscopic field of 400 × 400 µm in embryonic tissues at E10.5; *** p < 0.001 (two-tailed t-test); mean ± SD. g Quantification of intravascular CX3CR1+ cells in YS of Cx3cr1GFP/+ and Cx3cr1GFP/GFP E10.5 embryos; n.s. (two-tailed t-test: p = 0.6065); mean ± SD. Scale bar is 100 µm (c)
The chemokine receptor CX3CR1 mediates myeloid cell trafficking in inflammation38,39,40 and macrophage colonization of the mouse embryo occurs in a CX3CR1-dependent manner21. We therefore addressed whether pre-macrophage trafficking was modulated by CX3CR1. In E10.5 embryos we found an increased cell density in the YS of Cx3cr1GFP/GFP mice, while cell densities in embryonic tissues were decreased at this stage of development as previously reported (Fig. 4f)21. However, the number of cells per minute trafficking from the YS to the embryo was similar between Cx3cr1GFP/GFP (functional knockout) and Cx3cr1GFP/+ littermates (Fig. 4g). Further, the different hematopoietic lineages in the fetal liver were not altered by loss of one or both Cx3cr1 alleles (Supplementary Fig. 3d–f). Thus, the molecular cues restricting macrophage migration from the YS to the embryo remain to be determined.
Trafficking of CSF1R+ macrophage progenitors
The major source of YS CX3CR1+ pre-macrophages are EMPs21, and we have previously shown that they can be labeled using a CSF1R-driven fate mapping system16. To define the trafficking dynamics of CSF1R+ macrophage progenitors we first performed intravital microscopy in Csf1rCre:Rosa26eYFP embryos (Fig. 5a). We observed a strong increase in YFP+ cells in YS and embryonic tissues during the early development with the spatiotemporal distribution pattern being comparable to that of CX3CR1+ pre-macrophages (Fig. 5b–g). It should be noted that EMPs have multilineage (i.e. erythroid, megakaryocyte and myeloid) potential23,41, resulting in the labeling of other hematopoietic cells in addition to macrophages16. Importantly, CSF1R+ cells traveled with similar velocity to CX3CR1+ pre-macrophages in the YS vasculature and the maximum number of trafficking progenitors was reached between 9.5 and E10.5 in both models, thus showing similar trafficking dynamics (Fig. 5b–d and Supplementary Movies 11, 12).
Trafficking kinetics of CSF1R+ cells are similar to pre-macrophages. a Schematic graph for the Csf1rCre:Rosa26eYFP mouse model. b, e Fluorescence images of CSF1R YFP+ cells (green) in the YS (e) with corresponding quantifications (b) of cells per microscopic field at indicated time points; mean ± SD. c Quantification of intravascular CSF1R+ cells in an average-sized vessel; mean ± SD. d Velocity of intravascular non-adhering (n = 21) and rolling (n = 11) CSF1R+ cells in the YS at E9.5; median ± IQR. f, g Fluorescence images of CSF1R YFP+ cells (green) in different embryonic regions (f) with corresponding quantifications (g) of cells per microscopic field at indicated time points; ** p = 0.0072, n.s. p = 0.0569 (two-tailed Mann–Whitney test); median ± IQR. Scale bars are 100 µm (e, f)
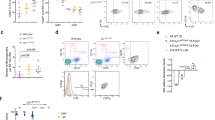
While labeling of CSF1R-expressing cells is more efficient in mice with constitutively active Cre recombinase as compared to a tamoxifen (TAM)-inducible system, onset of fetal definitive hematopoiesis may lead to a bias during microscopy of YS macrophage trafficking. In order to label YS-derived CSF1R+ cells and their progeny before the onset of fetal definitive hematopoiesis8, we crossed Csf1rMerCreMer mice with a Rosa26eYFP reporter and applied a single dose of 4-hydroxytamoxifen (OH-TAM) to induce expression of the yellow fluorescent protein (YFP) in a temporally-controlled manner (Fig. 6a–g). YFP labeling in E10.5 progenitors was low when OH-TAM was injected at E7.5, but increased about 6-fold when OH-TAM was injected at E8.5 (Fig. 6b, c), which is in line with previous reports that Csf1r becomes active during maturation12,16,34. E8.5 OH-TAM injection labeled about 10–20% erythrocytes, granulocytes and macrophages whereas cells of the lymphoid lineage were not labeled, which is consistent with EMP multilineage potential (Supplementary Fig. 4a–d)16,23.
Cellular expansion and morphology of CSF1R+ progenitors. a Schematic graph for the Csf1rMerCreMer:Rosa26eYFP mouse model with OH-TAM pulse labeling at E7.5 or E8.5 (red arrows) and subsequent analyses at indicated time points (black arrows). b, c Fluorescence pictures of E10.5 YS and embryo after OH-TAM pulse at E7.5 or E8.5 as indicated (b) with corresponding quantifications c; *** p < 0.001 (two-tailed Mann–Whitney test); median ± IQR. d Fluorescent pictures in the YS and embryo at E8.5, E9.0 and E9.5 with OH-TAM injection 24 h earlier. e, f Fluorescence pictures of a E12.5 YS and different embryonic regions after OH-TAM pulse labeling on E8.5 (e) with corresponding quantification of cluster size (f); *** p < 0.001 (Kruskal–Wallis test with Dunn’s multiple comparisons test); median ± IQR. g Quantification of intravascular cells at indicated time points in an average-sized vessel; median ± IQR. h Fluorescence pictures of a E10.25 YS in a Csf1rMerCreMer:Rosa26tdTomato (red):Cx3cr1GFP/+ (green) mouse model with OH-TAM pulse labeling at E8.5. Additional staining for F4/80 (purple) and CD31 (turquoise). i, j Adult liver sections of Csf1rMerCreMer:Rosa26eYFP mice, pulse-labeled at E8.5 (i) with additional stainings for CD31 (red) and DAPI (blue) (j). Scale bars are 100 µm (b, d, e, j), 1 mm (i), 50 µm (h)
YFP+ cells were first detected in the YS at E8.5 after OH-TAM pulse labeling at E7.5. Approximately 12 h later, coinciding with the onset of circulation, CSF1R+ cells could be also detected in the embryo (Fig. 6d). Trafficking kinetics of YFP+ cells in Csf1rMerCreMer:Rosa26eYFP embryos OH-TAM-pulsed at E8.5 were comparable with that of Csf1rCre:Rosa26eYFP and Cx3cr1GFP/+ embryos analyzed earlier (Fig. 6g and Supplementary Movies 13, 14). In general, infiltration of the embryo occurred mostly between E9.5 and E12.5 in these mice confirming that the trafficking of macrophages and their progenitors from the YS via the bloodstream was restricted to this well-defined time window of development (Fig. 6g). The difference in the number of trafficking cells at E12.5 between pulse-labeled Csf1rMerCreMer (Figs 5 and 6f) and Cx3cr1GFP/+ (Fig. 1i) mice could reflect CSF1R-labeling of EMPs as compared to matured “primitive” macrophages, the latter having completed their emigration from the YS at this time.
To dissect the different stages of macrophage development in the YS, we harnessed Csf1rMerCreMer:Rosa26tdTomato:Cx3cr1GFP/+ mice, in which we combined the E8.5 CSF1R-mediated OH-TAM pulse labeling approach with the constitutive Cx3cr1GFP/+ model to identify maturing macrophages. By E10.25/E10.5 a large proportion of CSF1R+ EMPs had acquired CX3CR1 (Fig. 6h, Supplementary Fig. 4c), which is in line with the recent reports on the sequence of macrophage maturation12,21. CSF1R+ CX3CR1+ pre-macrophages displayed a spherical shape and were frequently found to be traveling to the embryo (Fig. 6h, Supplementary Movies 3–5 and 7). F4/80 was expressed on mature macrophages with dendrites. These cells were located throughout the YS and often adjacent to YS vessels. They displayed reduced migration capacity and did not travel intraluminally (Figs. 3, 6h and Supplementary Movies 7, 8). Thus, trafficking of macrophage progenitors to the embryo correlated with defined stages of their development.
Expansion of CSF1R+ macrophage progenitors
The proliferative potential of YS EMPs has previously been demonstrated in vitro using clonogenic assays16. In Csf1rMerCreMer:Rosa26eYFP embryos in vivo, E8.5 OH-TAM pulse-labeled CSF1R+ progenitors displayed cell clusters in tissues (Fig. 6d, e and Supplementary Movies 13, 14). In the YS, cell clusters were relatively large (about 20 cells) while in the embryo cell conglomerates mostly consisted of few YFP+ cells (2–7 cells) at E12.5 (Fig. 6e, f). This suggests that macrophage progenitors expand in the YS before their emergence and further expansion takes place inside the embryo. This is in line with the proliferation dynamics of YS-derived EMPs in the fetal liver16 and of tissue macrophages in the epidermis of adult mice42. Notably, in liver sections of 1 year-old Csf1rMerCreMer:Rosa26eYFP mice pulsed at E8.5, large clusters of YFP+ Kupffer cells were present (Fig. 6i, j). Thus, the patterns of expansion initiated during early embryonic development can persist in adult mice. Future fate mapping experiments harnessing multicolor reporter mice are necessary to proof clonality of these cell clusters.
Trafficking dynamics of KIT+ EMPs
Application of a OH-TAM pulse at E8.5 in Csf1rMerCreMer:Rosa26eYFP embryos effectively labeled YS-derived EMPs (Supplementary Fig. 4b)16,43. However, maturing macrophages that arise from the first (primitive) wave of YS hematopoiesis may also be labeled once Csf1r has become active6,44. Flow cytometry analyses of the YS revealed a CD16/32+ KIT- population of primitive macrophages progenitors before the first appearance of EMPs suggesting their independent origin in line with previous reports (Supplementary Fig. 5a)12,23. Only a few hours later KIT+ EMPs develop in the YS and seed the fetal liver (Supplementary Fig. 5a–d) before they differentiate into KIT- CX3CR1+ pre-macrophages as described previously (Supplementary Fig. 5b, e)12,23.
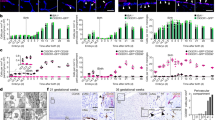
To more specifically determine the trafficking of YS-derived EMPs we carried out intravital microscopy in mice, in which Cre-dependent recombination was under control of the Kit promoter. KIT has previously been identified as a marker of EMPs12,45, and distinguishes them from maturing YS macrophages and primitive progenitors22,23. In order to label KIT+ progenitors, we crossed KitMerCreMer mice with Rosa26mT/mG reporter mice and applied a single dose of OH-TAM to induce GFP expression (Fig. 7a). In KitMerCreMer:Rosa26mT/mG embryos, OH-TAM application from E7.5 to E8.5 resulted in increased labeling of EMPs (Fig. 7b–d). Intravital microscopy at E10.5 showed that numerous EMPs, or their progeny, migrated to the embryo via the YS vasculature (Fig. 7e, f and Supplementary Movie 15). Cells derived from Kit-expressing progenitors thereby infiltrated all embryonic regions including the brain (Fig. 7b, c, g). However, the number of trafficking cells in E8.5 pulse-labeled KitMerCreMer embryos was lower than in E8.5 pulse-labeled Csf1rMerCreMer mice at E10.5, when trafficking reached its maximum (Figs. 6g and 7e). The difference might be explained by labeling of both EMPs and differentiating “primitive” macrophages in the Csf1r-dependent model. However, other factors such as differences in Cre recombination efficiency might also play a role. In summary, the experiments indicate that of the different cell types labeled by applying a OH-TAM pulse, EMPs and their progeny represent an integral part trafficking from the YS to the embryo, which is in line with their role in seeding the fetal liver13,16,46.
Trafficking of KIT+ EMPs. a Schematic graph for the KitMerCreMer:Rosa26mT/mG mouse model with OH-TAM pulse labeling at E7.5 or E8.5 (red arrows) and subsequent analyses at E10.5 (black arrow). b–d Fluorescent pictures of KIT+ cells in the YS and embryo on E10.5 after pulse labeling at indicated time points (b, c) with corresponding quantifications (d); *** p < 0.001 (Mann–Whitney test); median ± IQR. e Quantification of labeled KIT+ cells/min in an average-sized YS vessel at E10.5; median ± IQR. f Velocity of intravascular non-adhering KIT+ cells in the YS at E10.5; mean ± SD. g Flow cytometry plots of CD45+ gated KIT+ vs. GFP+ cells from pulse-labeled KitMerCreMer:Rosa26mT/mG mice in YS, head and trunk region at E10.5. Plots show an individual representative experiment. At least 3 embryos of minimum 2 independent litters were analyzed per time point. Scale bars are 100 µm (b), 1 mm (c)
Discussion
Our study provides a detailed characterization of early YS macrophage development and trafficking to embryonic tissues in high spatiotemporal resolution. Shortly after their appearance in the YS, macrophage progenitors—in both EMP and pre-macrophage stages—travel within the bloodstream to infiltrate the embryo. The trafficking process reaches a maximum around E10.5 and is mostly completed by E12.5. YS macrophages thereby traffic in spherical mobile shape. A dendrite-rich form is acquired by YS-derived macrophage progenitors once target tissues have been reached and further maturation proceeds. Thus, alterations in cell morphology accompanying macrophage maturation correlate with their distribution through the bloodstream (Fig. 8). However, the transition from round cells to dendritic macrophages could be happening independently of the trafficking process within tissues.
YS-derived macrophages traffic during defined stages of development. Graphical summary of morphology-dependent trafficking dynamics: macrophage progenitors expand in the YS from E8.5 onwards. They traffic to embryonic organs via vascular routes during a restricted time frame until E14.5. This trafficking period is characterized by a spherical cell shape. Along with progressive macrophage maturation an increasing amount of dendrites is formed and macrophages lose intravascular trafficking capacity
By applying intravital microscopy we provided direct evidence that YS macrophage progenitors colonize the embryo via the vasculature. There has been an ongoing debate on how YS macrophages enter the embryo, that is, by trans-tissue migration or trans-vascular trafficking via the bloodstream. The early appearance of myeloid colony-forming units in embryonic blood before the onset of fetal definitive hematopoiesis indicates trans-vascular trafficking13. Further, infiltration of embryonic tissues by EMPs and maturing macrophages is impaired in mice lacking a heartbeat supporting the role of a functional circulation for macrophage trafficking7,22,47,48,]. However, this model is limited by its dramatic impairment of overall mouse physiology resulting in early embryonic lethality49. In contrast to the data supporting trans-vascular migration, it has been suggested for various species that macrophages can colonize embryonic tissues independently of blood vessels via extravascular routes25,26,27,35. In line with this, functional circulation is not required for EMP emergence in the YS22. We interpret our data to indicate that macrophage progenitors infiltrate the embryo predominantly by intravascular trafficking. However, the microscopy tools applied in this paper are suited for imaging fast intravital processes and, therefore, we cannot quantify nor exclude trans-tissue migration.
Macrophage trafficking reached its maximum around E10.5, and we detected hardly any intravascular CX3CR1+ pre-macrophages after E12.5. Blood flow velocity in the YS vasculature still increases at this gestational age in mice. It is half maximal at E10.5 and does not reach a plateau before E13.550. Therefore, flow velocity does not seem to relevantly contribute to the trafficking of macrophages to the embryo. Further, trafficking is not influenced by the onset of fetal definitive hematopoiesis as demonstrated in Myb-deficient mouse embryos. The absence of CX3CR1 chemokine receptor signaling resulted in higher macrophage densities in the YS and impaired colonization of the embryo21; however, the quantity of intravascular cells was not altered. Thus, CX3CR1 does not affect intravascular trafficking of macrophage progenitors from the YS, it might however alter macrophage proliferation in the YS51 or modulate survival in peripheral embryonic tissues52,53, which remains to be determined. The molecular cues regulating macrophage trafficking from the YS to the embryo will need to be addressed in future studies. In addition to potential soluble molecules in the YS environment or cell surface receptors expressed on YS progenitors, endothelium-derived signals have recently been reported to modulate the seeding of macrophages in their destined tissue within the embryo54.
YS hematopoiesis gives rise to multipotent cells that can develop into fetal lymphoid progenitors55,56,57,58. The EMPs and pre-macrophages progenitors pulse-labeled in Csf1rMerCreMer:Rosa26eYFP embryos did not express markers of lymphoid cells. Although we did not carry out clonogenic assays and we therefore cannot formally exclude the lymphoid potential of trafficking CSF1R+ cells22,59, our findings are in line with a recent study by McGrath and colleagues who phenotyped purified EMPs and demonstrated their lack of B-lymphoid potential23. Further, lineage tracing in Rag1Cre:Rosa26eYFP mice indicated that tissue macrophages develop independently of lymphoid-primed YS progenitors59.
Using intravital microscopy, we show that YS macrophage progenitors migrate to the embryo in both EMP and pre-macrophage stages. This is in line with the concept that EMPs travel to the embryo to colonize the nascent fetal liver before E10.5, where they give rise to tissue macrophages and other myeloid cells (Supplementary Fig. 4b)13,16,46. The observation that more fluorescent cells were trafficking in E10.5 Csf1rMerCreMer embryos than in KitMerCreMer mice might be explained by the labeling of both maturing macrophages derived from primitive hematopoiesis and EMPs in the CSF1R-dependent model, whereas pulse labeling in KitMerCreMer embryos is more specific for fate mapping of EMPs23.
Most macrophage progenitors travel between E9.5 and E10.5; however first pre-macrophages enter the embryo around E9.0 with the onset of functional circulation. This population of macrophages predominantly infiltrates the embryonic brain (Figs. 2a–d and 5f, g and Supplementary Fig. 3a–c). It is possible that these cells arise from the first wave of KIT- CD16/32+ primitive progenitors6. However, this population might be heterogeneous and additional markers to unequivocally identify primitive macrophages remain to be determined. In close temporal proximity, emerging EMPs give rise to an increasing pool of macrophage progenitors that seed all embryonic tissues. Depending on the organ environment, these YS-derived resident macrophages then persist (e.g. brain, liver) or are replaced by circulating monocytes, either rapidly (e.g. intestine) or over time (e.g. lung, serous cavities)7,16,60,61,62,]. In summary, our study provides novel insights into the emergence of macrophage progenitors from the YS, a key process in the establishment of the innate immune system.
Methods
Mice
Cx3cr1GFP/+ 37, Cx3cr1Cre 9, Myb−/− 63, Csf1rCre expressing an improved Cre sequence under control of the Csf1r promoter64, TAM-inducible Csf1rMerCreMer 65 and KitMerCreMer 66, as well as Rosa26eYFP 67, Rosa26mT/mG 68 and Rosa26tdTomato 69 reporter mice have been previously described. Csf1rCre and Csf1rMerCreMer were on FVB/N background, all other mice were on C57BL/6J background (backcrossed for at least 5 generations). Timed matings were carried out using 10–20 weeks old mice. Embryos were analyzed at indicated time points without determining their sex. Mice were maintained in a specific pathogen-free environment and fed standard mouse diet ad libitum. All animal procedures were performed in adherence to our project licence (55.2-1-54-2532-181-13) issued by the German regional council at the Government of Bavaria (Regierungspräsidium Oberbayern), Munich, Germany, and by the Institutional Review Board (IACUC 15-04-006) of the Memorial Sloan Kettering Cancer Center, New York, USA.
Timed matings and pulse labeling
Mice of desired genotypes were mated overnight. Embryonic development was estimated considering midday of the vaginal plug as embryonic day 0.5 (E0.5). Pregnant females were used for subsequent experiments on indicated time points. For genetic cell labeling, we crossed TAM-inducible Csf1rMerCreMer or KitMerCreMer with Rosa26eYFP or Rosa26mT/mG. Recombination was induced in embryos by single injection of 75 µg per g (body weight) of 4-hydroxytamoxifen (Sigma) into pregnant females at indicated time points.
Intravital epifluorescence microscopy
Experiments were performed in deep anesthesia. Access to the abdominal cavity of pregnant mice was provided by a cesarean section. Depending on the developmental stage, intravital imaging was predominantly carried out in separation from the mother animal (E8.5–E12.5) for reasons of image quality or in continuous connection to the maternal circulation (E12.5–E16.5). Quantity and velocity of trafficking CX3CR1+ cells in separated E10.5 embryos was similar to embryos being in continuous connection to the mother (Supplementary Fig. 1g, h). Imaging of all embryos was performed in a heated environment in 37 °C PBS with continuous temperature monitoring. Average sized vessels of the YS (Supplementary Fig. 1f) were imaged using an Olympus BX51WI epifluorescence microscope with a Plan N 10x/0.25 or PlanApo N 1.25x/0.04 objective for 3–15 min in different areas of the YS or the embryo using the Olympus cell R imaging software. Maximum total imaging periods per YS were about 60 min. Vascular borders were identified by spontaneous image contrast in the GFP channel and confirmed by an intravital endothelial staining with a rat anti-CD31 (PECAM-1) antibody (clone MEC 13.3, Biolegend, 1:100) (Figs. 1f and 2e). Cell clusters were defined as a cell-to-cell distance <2 cell diameters (Fig. 6f). After imaging, mice were sacrificed and embryos were used for further analysis.
Intravital spinning disc confocal microscopy
Experiments were performed in deep anesthesia. Embryos were prepared as described above for intravital epifluorescence microscopy. Analysis of YS-derived macrophage trafficking in real-time in vivo was performed in Cx3cr1Cre:Rosa26mT/mG embryos (E9.5, E10.5) using an upright spinning disk confocal microscope (Examiner, Zeiss, Germany) equipped with a confocal scanner unit CSU-X1 (Yokogawa Electric Corporation, Japan) and a CCD camera (Evolve, Photometrics, USA). Images were acquired with a 20x/ 1.0 NA or a 63x/ 1.0 NA water immersion objective (Plan Apochromat, Zeiss, Germany) using a laser with an excitation wavelength of 488 nm for the detection of macrophages and a laser with an excitation wavelength of 561 nm for the detection of the tomato fluorescence signal of all other tissues. Imaging and image processing was performed using Slidebook 6.0.11 Software (3i, USA) and Fiji (NIH, USA)70.
Antibodies for immunofluorescence
For cryosections of mouse tissues, we used a rat anti-CD31 (PECAM-1) antibody (clone MEC 13.3, Biolegend, 1:50) and a rabbit anti-GFP antibody (polyclonal, Invitrogen Life Technologies, 1:200). Cy3- and Alexa 488-conjugated secondary antibodies (Dianova Immuno Research) were added at a dilution of 1:500. For whole mount stainings we used a rat anti-CD31-PE antibody (clone MEC13.3, Biolegend, 1:200), an armenian hamster anti-CD31 antibody (clone 2H8, Thermo Scientific, 1:500) and a rat anti-F4/80 antibody (clone: Cl:A3-1, Biorad, 1:500). Secondary antibodies used were anti-armenian hamster DyLight 405 (Jackson ImmunoResearch, 1:500) and anti-rat Alexa Fluor 647 (Invitrogen, 1:500). Nuclei were stained using Hoechst (Hoechst 33342, ThermoFisher, 1:2000) or DAPI (ThermoFisher, 1 µg/ml).
Human cryosections were stained with a primary mouse anti-CD68 antibody (clone CL1346, Sigma, 1:1000), rabbit anti-CD68 antibody (polyclonal, Sigma, 1:1000), rabbit anti-CX3CR1 (polyclonal, Abcam, 1:100), or rat anti-EMR2 (clone W15101A, Biolegend, 1:100), as well as a secondary goat-anti-mouse IgG Cy2 antibody (Jackson ImmunoResearch, 1:100), goat anti-rat IgG Cy3 (Dianova, 1:100), goat anti-rabbit IgG Cy2 (Dianova, 1:100), or goat anti-rabbit IgG Cy3 antibody (Dako, 1:100). Nuclei were stained using Vectashield mounting medium for fluorescence with DAPI (Vector Laboratories, 1.5 µg/ml).
Human samples and laser scanning confocal microscopy
Preparation and analysis of human material was approved by the Institution’s ethics committee (reference #337-06 amended 26/08/2013). The study protocol conformed to the ethical guidelines of the 1975 Declaration of Helsinki and written informed consent was obtained from each patient included in the study. Human samples were fixed in 4% neutral buffered formalin, dehydrated and consecutively embedded in paraffin. Blocks were sliced in 5 µm sections. Afterwards sample sections were washed and stained with indicated primary antibody for 60 min at room temperature and secondary antibodies for 30 min at room temperature. Confocal fluorescence images were obtained with a LSM880 Zeiss microscope with a 20x/0,8 dry objective and an Airyscan module. Images were acquired using ZEN software (Zeiss, Germany).
Analysis of mouse liver sections
Timed matings were performed as described above and Csf1rMerCreMer:Rosa26eYFP embryos were pulse-labeled at E8.5. 1 year after birth, mice were euthanized under terminal anesthesia and livers were harvested. Organs were washed in PBS, fixed in 4% PFA and incubated in 30% sucrose overnight at 4 °C. Samples were embedded in Tissue-Tek OCT (Sakura Finetek) and cryoblocks were sliced into 15 μm sections. Sections were incubated in PBS containing BSA (5%) and normal goat IgG (1:60). Primary antibodies were added overnight at 4 °C. After washing, secondary antibodies were incubated for at least 2 h at room temperature. Nuclei were counterstained with DAPI. Sections were post-fixed with PFA 1% for 1 min and mounted with mounting medium (Vector Laboratories). Acquisitions were performed using a Leica TCS SP5 confocal microscope (Leica Microsystems) with tile scan software and 10x/0.4 (dry) as well as 20x/0.7 (water immersion) objective.
Microscopy of mouse whole mount stainings
YSs were mounted and visualized using a Axio Imager.M2 microscope (Zeiss, Germany) with a 20x/0.75 Plan Apochromat or 40x/0.8 Plan Neofluar objective and AxioVision Software (Zeiss, Germany) (refers to data presented in Supplementary Fig. 1d, e). Alternatively, a LSM880 Zeiss microscope with 40x/1.4 (oil) objective with ZEN black software (Zeiss, Germany) was used and data was analyzed with Imaris software (refers to data presented in Fig. 6g).
Flow cytometry of mouse embryonic tissues
Pregnant females were sacrificed by cervical dislocation. Embryos ranging from embryonic day (E) 8.5 to E16.5 were dissected out from the uterus and washed in 4 °C phosphate buffered saline (PBS, Invitrogen). The YS, fetal liver or embryonic regions (brain, trunk, tail) were harvested and digested in PBS containing 1 mg/ml Collagenase D (Roche), 100 U/ml Desoxyribonuclease I (Sigma) and 1% fetal bovine serum (PAA Laboratories) at 37 °C. Tissues were mechanically dissociated and passed through a 100 μm cell strainer (BD). Cells were centrifuged at 400 g for 5 min, resuspended in 4 °C PBS, plated in multi-well round-bottom plates and immunolabeled for flow cytometry analysis. Antibody mixes were added in 1% BSA/PBS and incubated for 20 min on 4 °C.
APC-Cy7-labeled anti-CD45.2 (clone 104) antibodies were from BD Pharmingen. Brilliant violet 421-labeled anti-F4/80 (clone BM8), APC-labeled anti-B220 (clone RA3-6B2), brilliant violet 421-labeled anti-CSF1R (CD115; clone AFS98), Alexa 647-labeled anti- CX3CR1 (clone SA011F11), PE-labeled anti-CD4 (clone H129.19), PE-labeled anti-CD19 (clone 6D5), APC-labeled anti-IL7Ra (CD127; clone A7R34), PE-labeled anti-Ter119 (clone Ter-119) and PE-labeled anti-CD16/32 (FcgRIII/II; clone 93) antibodies were from Biolegend. APC-labeled anti-KIT (CD117; clone 2B8) and PE-labeled anti-CD11b (clone M1/70) antibodies were from BD Biosciences. APC-labeled anti-Gr1 (Ly6G; clone RB6-8C5) antibodies were from eBiosciences. Flow cytometry was performed using a BD Biosciences LSR Fortessa flow cytometer and data were analyzed using FlowJo 10.
Statistics
Data were tested for normal distribution using the D’Agostino & Pearson omnibus test. In case of normal distribution, comparisons between groups were calculated using unpaired, two-tailed Student’s t-test or one-way analysis of variance ANOVA (with Tukey’s multiple comparisons test). If normal distribution was not given, Mann-Whitney test or Kruskal-Wallis test (with Dunn’s multiple comparisons test) was used. A p-value of <0.05 was regarded as significant. Data is indicated as (***) p < 0.001, (**) p < 0.01, (*) p < 0.05, or not significant (n.s.) if p ≥ 0.05. Data are are expressed as mean ± standard deviation (SD) if normal, or as median with interquartile range (IQR) if not distributed normally. Dots represent individual values. All graphs and calculations were generated with GraphPad Prism 7 software.
Animal experiments were analyzed in a blinded manner where possible, i.e., when the genotype of the embryos was unknown at the time of experiments (analysis of Cx3cr1GFP/+ vs. Cx3cr1GFP/GFP, and Myb+/+ vs. Myb−/− mice).
Data availability
The authors declare that the data supporting the findings of this study are available within the article and its supplementary information files.
Change history
07 September 2018
This article contains errors in Figs. 5 and 6, for which we apologize. In Fig. 5f, the image ‘E12.5 tail’ was inadvertently replaced with a duplicate of the image ‘E12.5 trunk’ from the same panel. In Figure 6d, the image ‘E9.5/OH-TAM E8.5, embryo’ was inadvertently replaced with a duplicate of the image ‘E10.5/ OH-TAM E8.5, embryo’ from Fig. 6b. The corrected versions of these figures appear in the Author Correction associated with this Article.
References
Tauber, A. I. Metchnikoff and the phagocytosis theory. Nat. Rev. Mol. Cell. Biol. 4, 897–901 (2003).
Moore, K. J., Sheedy, F. J. & Fisher, E. A. Macrophages in atherosclerosis: a dynamic balance. Nat. Rev. Immunol. 13, 709–721 (2013).
McNelis, J. C. & Olefsky, J. M. Macrophages, immunity, and metabolic disease. Immunity 41, 36–48 (2014).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61 (2014).
Prinz, M. & Priller, J. Microglia and brain macrophages in the molecular age: from origin to neuropsychiatric disease. Nat. Rev. Neurosci. 15, 300–312 (2014).
McGrath, K. E., Frame, J. M. & Palis, J. Early hematopoiesis and macrophage development. Sem. Immunol. 27, 379–387 (2016).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Hashimoto, D. et al. Tissue-resident macrophages self-maintain locally throughout adult life with minimal contribution from circulating monocytes. Immunity 38, 792–804 (2013).
Epelman, S., Lavine, K. J. & Randolph, G. J. Origin and functions of tissue macrophages. Immunity 41, 21–35 (2014).
Bertrand, J. Y. et al. Three pathways to mature macrophages in the early mouse yolk sac. Blood 106, 3004–3011 (2005).
Palis, J., Robertson, S., Kennedy, M., Wall, C. & Keller, G. Development of erythroid and myeloid progenitors in the yolk sac and embryo proper of the mouse. Development 126, 5073–5084 (1999).
Sheng, J., Ruedl, C. & Karjalainen, K. Most tissue-resident macrophages except microglia are derived from fetal hematopoietic stem cells. Immunity 43, 382–393 (2015).
Gomez Perdiguero, E., Schulz, C. & Geissmann, F. Development and homeostasis of “resident” myeloid cells: the case of the microglia. Glia 61, 112–120 (2013).
Gomez Perdiguero, E. et al. Tissue-resident macrophages originate from yolk-sac-derived erythro-myeloid progenitors. Nature 518, 547–551 (2015).
Bertrand, J. Y. et al. Haematopoietic stem cells derive directly from aortic endothelium during development. Nature 464, 108–111 (2010).
Boisset, J.-C. et al. In vivo imaging of haematopoietic cells emerging from the mouse aortic endothelium. Nature 464, 116–120 (2010).
Kissa, K. & Herbomel, P. Blood stem cells emerge from aortic endothelium by a novel type of cell transition. Nature 464, 112–115 (2010).
Kanitakis, J., Petruzzo, P. & Dubernard, J. M. Turnover of epidermal Langerhans’ cells. N. Engl. J. Med. 351, 2661–2662 (2004).
Mass, E. et al. Specification of tissue-resident macrophages during organogenesis. Science 353, aaf4238 (2016).
Frame, J. M., Fegan, K. H., Conway, S. J., McGrath, K. E. & Palis, J. Definitive hematopoiesis in the yolk sac emerges from Wnt-responsive hemogenic endothelium independently of circulation and arterial identity. Stem Cells 34, 431–444 (2016).
McGrath, K. E. et al. Distinct sources of hematopoietic progenitors emerge before HSCs and provide functional blood cells in the mammalian embryo. Cell Rep. 11, 1892–1904 (2015).
Gordon, S., Pluddemann, A. & Martinez Estrada, F. Macrophage heterogeneity in tissues: phenotypic diversity and functions. Immunol. Rev. 262, 36–55 (2014).
Herbomel, P., Thisse, B. & Thisse, C. Zebrafish early macrophages colonize cephalic mesenchyme and developing brain, retina, and epidermis through a M-CSF receptor-dependent invasive process. Dev. Biol. 238, 274–288 (2001).
Chan, W. Y., Kohsaka, S. & Rezaie, P. The origin and cell lineage of microglia: new concepts. Brain Res. Rev. 53, 344–354 (2007).
Cuadros, M. A., Martin, C., Coltey, P., Almendros, A. & Navascues, J. First appearance, distribution, and origin of macrophages in the early development of the avian central nervous system. J. Comp. Neurol. 330, 113–129 (1993).
Ovchinnikov, D. A. et al. Expression of Gal4-dependent transgenes in cells of the mononuclear phagocyte system labeled with enhanced cyan fluorescent protein using Csf1r-Gal4VP16/UAS-ECFP double-transgenic mice. J. Leukoc. Biol. 83, 430–433 (2008).
Godin, I. & Cumano, A. The hare and the tortoise: an embryonic haematopoietic race. Nat. Rev. Immunol. 2, 593–604 (2002).
Takashina, T. Haemopoiesis in the human yolk sac. J. Anat. 151, 125–135 (1987).
Migliaccio, G. et al. Human embryonic hemopoiesis. Kinetics of progenitors and precursors underlying the yolk sac–liver transition. J. Clin. Invest. 78, 51–60 (1986).
Kwakkenbos, M. J. et al. The human EGF-TM7 family member EMR2 is a heterodimeric receptor expressed on myeloid cells. J. Leukoc. Biol. 71, 854–862 (2002).
Chang, G. W. et al. CD312, the human adhesion-GPCR EMR2, is differentially expressed during differentiation, maturation, and activation of myeloid cells. Biochem. Biophys. Res. Commun. 353, 133–138 (2007).
Lichanska, A. M. et al. Differentiation of the mononuclear phagocyte system during mouse embryogenesis: the role of transcription factor PU.1. Blood 94, 127–138 (1999).
Fantin, A. et al. Tissue macrophages act as cellular chaperones for vascular anastomosis downstream of VEGF-mediated endothelial tip cell induction. Blood 116, 829–840 (2010).
Ema, H. & Nakauchi, H. Expansion of hematopoietic stem cells in the developing liver of a mouse embryo. Blood 95, 2284–2288 (2000).
Jung, S. et al. Analysis of fractalkine receptor CX(3)CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Prinz, M. & Priller, J. Tickets to the brain: role of CCR2 and CX3CR1 in myeloid cell entry in the CNS. J. Neuroimmunol. 224, 80–84 (2010).
Carlin, L. M. et al. Nr4a1-dependent Ly6C(low) monocytes monitor endothelial cells and orchestrate their disposal. Cell 153, 362–375 (2013).
Stolla, M. et al. Fractalkine is expressed in early and advanced atherosclerotic lesions and supports monocyte recruitment via CX3CR1. PLoS ONE 7, e43572 (2012).
McGrath, K. E. et al. A transient definitive erythroid lineage with unique regulation of the beta-globin locus in the mammalian embryo. Blood 117, 4600–4608 (2011).
Ghigo, C. et al. Multicolor fate mapping of Langerhans cell homeostasis. J. Exp. Med. 210, 1657–1664 (2013).
Hoeffel, G. et al. C-Myb(+) erythro-myeloid progenitor-derived fetal monocytes give rise to adult tissue-resident macrophages. Immunity 42, 665–678 (2015).
Ginhoux, F. & Guilliams, M. Tissue-resident macrophage ontogeny and homeostasis. Immunity 44, 439–449 (2016).
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013).
Kieusseian, A. et al. Immature hematopoietic stem cells undergo maturation in the fetal liver. Development 139, 3521–3530 (2012).
Hoeffel, G. et al. Adult Langerhans cells derive predominantly from embryonic fetal liver monocytes with a minor contribution of yolk sac-derived macrophages. J. Exp. Med. 209, 1167–1181 (2012).
Lux, C. T. et al. All primitive and definitive hematopoietic progenitor cells emerging before E10 in the mouse embryo are products of the yolk sac. Blood 111, 3435–3438 (2008).
Cho, C. H., Kim, S. S., Jeong, M. J., Lee, C. O. & Shin, H. S. The Na+ -Ca2+ exchanger is essential for embryonic heart development in mice. Mol. Cells 10, 712–722 (2000).
Mu, J. & Adamson, S. L. Developmental changes in hemodynamics of uterine artery, utero- and umbilicoplacental, and vitelline circulations in mouse throughout gestation. Am. J. Physiol. Heart Circ. Physiol. 291, H1421–H1428 (2006).
Engel, D. R. et al. CX3CR1 reduces kidney fibrosis by inhibiting local proliferation of profibrotic macrophages. J. Immunol. 194, 1628–1638 (2015).
Lionakis, M. S. et al. CX3CR1-dependent renal macrophage survival promotes Candida control and host survival. J. Clin. Invest. 123, 5035–5051 (2013).
Ensan, S. et al. Self-renewing resident arterial macrophages arise from embryonic CX3CR1(+) precursors and circulating monocytes immediately after birth. Nat. Immunol. 17, 159–168 (2016).
Rantakari, P. et al. Fetal liver endothelium regulates the seeding of tissue-resident macrophages. Nature 538, 392–396 (2016).
Ueno, H. & Weissman, I. L. Clonal analysis of mouse development reveals a polyclonal origin for yolk sac blood islands. Dev. Cell. 11, 519–533 (2006).
Moore, M. A. & Metcalf, D. Ontogeny of the haemopoietic system: yolk sac origin of in vivo and in vitro colony forming cells in the developing mouse embryo. Br. J. Haematol. 18, 279–296 (1970).
Samokhvalov, I. M., Samokhvalova, N. I. & Nishikawa, S. Cell tracing shows the contribution of the yolk sac to adult haematopoiesis. Nature 446, 1056–1061 (2007).
Inlay, M. A. et al. Identification of multipotent progenitors that emerge prior to hematopoietic stem cells in embryonic development. Stem Cell Rep. 2, 457–472 (2014).
Boiers, C. et al. Lymphomyeloid contribution of an immune-restricted progenitor emerging prior to definitive hematopoietic stem cells. Cell. Stem Cell 13, 535–548 (2013).
Bain, C. C. et al. Long-lived self-renewing bone marrow-derived macrophages displace embryo-derived cells to inhabit adult serous cavities. Nat. Commun. 7, 11852 (2016).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Bain, C. C. et al. Constant replenishment from circulating monocytes maintains the macrophage pool in the intestine of adult mice. Nat. Immunol. 15, 929–937 (2014).
Mucenski, M. L. et al. A functional c-myb gene is required for normal murine fetal hepatic hematopoiesis. Cell 65, 677–689 (1991).
Deng, L. et al. A novel mouse model of inflammatory bowel disease links mammalian target of rapamycin-dependent hyperproliferation of colonic epithelium to inflammation-associated tumorigenesis. Am. J. Pathol. 176, 952–967 (2010).
Qian, B. Z. et al. CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature 475, 222–225 (2011).
van Berlo, J. H. et al. c-kit+ cells minimally contribute cardiomyocytes to the heart. Nature 509, 337–341 (2014).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank Nicole Blount and Beate Jantz for animal husbandry. We thank Sebastian Helmer, Marina Hofmann and Michael Lorenz for mouse genotyping. We thank Christina Kuhn for immunofluorescence stainings. We thank Steffen Jung, The Weizmann Institute of Science, for providing Cx3cr1Cre mice. This study was supported by the SFB914, projects A01 (M.M.), A10 (C.Schu.), B01 (M.S.), B02 (S.M.), Z01 (S.M., H.I.-A.) and Z03 (R.P., C.Schei.), as well as the DZHK (German Centre for Cardiovascular Research) and the BMBF (German Ministry of Education and Research) to C.S. (High Risk High Volume Late Translational project). C.Str. is supported by a Gerok position of the SFB914. F.G. was supported by NIH/NCI P30CA008748 MSKCC core grant and NIH/NIAID 1R01AI130345-01.
Author information
Authors and Affiliations
Contributions
C.Str. and C.Schu. designed the study, performed experiments and wrote the paper; R.S. and F.W. and performed experiments; A.M and M.S. helped to optimize intravital epifluoresence imaging of the embryo and performed associated experiments; R.P. and C.Schei. provided the spinning disc confocal microscopy platform and performed associated experiments; H.C.I.-A. provided the confocal microscopy platform and performed associated experiments, E.M. performed experiments with the Csf1rMerCreMer:Rosa26tdTomato:Cx3cr1GFP/+ mouse line; S.H. and U.J. provided human material and performed associated experiments; R.T., T.W., M.S., S.M. and F.G. discussed data and revised the manuscript; J.F., S.Y., R.V., J.D.M., S.K. and M.M. generated and provided mice and discussed data.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stremmel, C., Schuchert, R., Wagner, F. et al. Yolk sac macrophage progenitors traffic to the embryo during defined stages of development. Nat Commun 9, 75 (2018). https://doi.org/10.1038/s41467-017-02492-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-017-02492-2
This article is cited by
-
The niche matters: origin, function and fate of CNS-associated macrophages during health and disease
Acta Neuropathologica (2024)
-
Pathophysiology and probable etiology of cerebral small vessel disease in vascular dementia and Alzheimer’s disease
Molecular Neurodegeneration (2023)
-
Macrophage-driven cardiac inflammation and healing: insights from homeostasis and myocardial infarction
Cellular & Molecular Biology Letters (2023)
-
Microglia in neurodegenerative diseases: mechanism and potential therapeutic targets
Signal Transduction and Targeted Therapy (2023)
-
Mechanisms of myeloid cell entry to the healthy and diseased central nervous system
Nature Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.