Abstract
Hepatitis C virus infection represents a major problem of public health with around 350 millions of chronically infected individuals worldwide. The frequent evolution towards severe liver disease and cancer are the main features of HCV chronic infection. Antiviral therapies, mainly based on the combination of IFN and ribavirin can only assure a long term eradication of the virus in less than half of treated patients. The mechanisms underlying HCV pathogenesis and persistence in the host are still largely unknown and the efforts made by researchers in the understanding the viral biology have been hampered by the absence of a reliable in vitro and in vivo system reproducing HCV infection. The present review will mainly focus on viral pathogenetic mechanisms based on the interaction of HCV proteins (especially core, NS3 and NS5A) with host cellular signaling transduction pathways regulating cell growth and viability and on the strategies developed by the virus to persist in the host and escape to antiviral therapy. Past and recent data obtained in this field with different experimental approaches will be discussed.
Similar content being viewed by others
Introduction
Hepatitis C Virus (HCV) is a major pathogen given its extremely high prevalence (around 350 millions of chronically infected individuals worldwide), the high rate of chronic infection and the significant risk of severe chronic active hepatitis and cirrhosis among chronically infected subjects. HCV infection induces chronic infection in up to 60–80% of infected adults. The pathogenic effects of chronic HCV infections is highly variable; some patients will only show minimal liver lesions while others (around 20%) will develop after 5–10 years follow-up severe fibrosis and cirrhosis. Finally, 30–50% of patients with cirrhosis, whatever the etiological factor, develop HCC after a 10 year follow-up.
Acute and chronic hepatitis induced by most hepatitis viruses clearly implicate the host immune response to viral proteins. The development of molecular biology has led to realize that some hepatitis viruses also directly modulate liver cell proliferation and viability. This issue has important implications for understanding the molecular bases of the liver lesions and liver carcinogenesis, as well as for developing new therapeutical strategies. The present review will focus on the viral pathogenetic mechanisms based on the interaction of HCV proteins with host cellular signaling transduction pathways regulating cell growth and viability and on the strategies developed by the virus to persist in the host and escape to antiviral therapy.
Hepatitis C virus biology
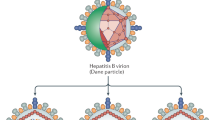
HCV belongs to the genus of Hepacivirus and is a member of the Flaviviridae, together with the Pestiviruses and Flaviviruses. Its genome is a positive, single strand RNA molecule which includes two untranslated regions at the 5′ and 3′ ends, and a large open reading frame encoding for a 3010–3030 amino acid polyprotein. This polyprotein is post-translationally processed into structural and non-structural proteins, the cleavage being dependent on host and viral encoded enzymes (Figure 1). The structural proteins, encoded in the N-terminal region, include the core protein followed by two envelope glycosylated proteins: E1 and E2. The non-structural domain encodes for six proteins: NS2, 3, 4A, 4B and 5A, 5B (see article from Dr Bartenschlager in this Special Issue).
HCV RNA shows significant genetic variability, and exhibits an estimated rate of nucleotide change of approximately 10−3 substitutions/site/year (see article from Dr Simmonds in this Special Issue). This genetic variability can be seen in all domains of the HCV RNA and the concept of ‘quasi-species’ has been introduced. In fact, HCV RNA circulates, in most cases, as a population of RNA molecules which also differs in serum, liver and PBMC. Some ‘hot spots’ of mutation have been identified in the E2 envelope protein encoding sequences where there are two hypervariable sequences (HVR 1 and 2), located in the 5′ part of this domain, with a high rate of non-synonymous mutations (i.e. leading to amino acid changes). Some of the non-structural (NS5A in particular) as well as the capsid encoding sequences show a lower, yet significant, rate of variability. In contrast, the 5′ untranslated region is highly conserved among different isolates although mutations can be identified at some sites. At present three major HCV types are distinguished: one, two and three as well as three to seven other types, according to the classification proposed. There is a general agreement that the response to interferon therapy depends on the infecting HCV type, while the association between certain types and liver lesions severity is highly debated.1 The existence of a population of HCV RNA quasispecies may favor the selection of RNA molecules that are ‘resistant’ to anti-viral factors and some evidence have been provided for a time-dependent selection of some of these HCV species (reviewed in 2).
The progress in HCV research has been hampered by the lack of an efficient and reliable model supporting viral entry, replication and secretion of viral particles. However, some in vitro approaches has been recently developed such as the HCV replicons. This represents presently the most relevant model to investigate HCV RNA replication, HCV protein synthesis and maturation, and should allow new approaches for the discovery and characterization of potential antiviral molecules. The HCV replicons consist in genomic HCV–RNAs capable of autonomous replication exclusively in a particular hepatoma cell line (Huh7). First HCV replicons consisted in bicistronic constructs with a selection gene (neomycin) downstream the HCV internal ribosome entry segment (IRES) followed by encephalomyocarditis (EMCV) IRES driving the translation of a second cistron consisting in a subgenomic HCV–RNA including the non-structural genes of the virus (NS3 to NS5B). Such constructions only allowed the persistence of the replicon in cells where HCV IRES was active. The following generation of replicons has now included the complete HCV open reading frame, encoding for both structural and nonstructural proteins. Subgenomic and genome-length replicons were able to express HCV proteins and to persist for years in the cells, but no encapsidation or secretion of viral particles has been shown in this model. The amount of replicon RNA in the cells was dependent to the degree of confluence of cell culture, rapidly decreasing at high confluence and indicating that HCV RNA replication or translation is tightly linked to host cell metabolism.3,4,5 The replicon model confirmed in vitro the capacity of HCV replication machinery to adapt to the cellular context by point mutations with some hot-spots along the genome. In fact, some point mutations in key regions of HCV genome (namely NS3- and mostly NS5A-encoding sequences) strongly increased HCV replication, unmasking the role of some HCV proteins in HCV-RNA transcription and translation processes.3,4,6
The absence of small animal models for HCV infection has been also partially addressed by Mercer et al.7 The authors transplanted normal human hepatocytes in severe combined immunodeficient mice carrying the urokinase-like plasminogen activator transgene. Transplanted human cells, taking advantage to the toxicity of the transgene expressed in resident mice hepatocytes, extensively repopulate mouse liver and support productive HCV infection. Similar animal models hosting human hepatocytes have been previously proposed as an infection model for other hepatitis viruses.8,9 Finally, a different approach to study HCV infection in vivo has been recently developed; laser capture microdissection (LCM) technique allows the isolation and analysis of single infected cells chosen by histological and immunohistochemical criteria from liver sections.10 This strategy will open new avenues in the comprehension of HCV related pathogenesis, directly focusing on the analysis of infected cells.
Infection by HCV of non-hepatocytic cells
There is now evidence that HCV also infects extrahepatic cells. In vivo, HCV RNA sequences have been detected in fresh peripheral blood mononuclear cell (PBMC) preparations from HCV infected patients.11,12 However, such an approach does not allow distinction between absorption of serum particles and true infection of PBMC in viremic patients. Still, several observations support this hypothesis including: (a) short-term cultures of PBMC yield a significant increase in the amount of viral RNA on stimulation by PHA and PMA;11 (b) in situ hybridization showed viral RNA in a limited percentage (around 1%) of circulating PBMC and lymph nodes;13,14 (c) PBMC from normal individuals can be infected by HCV;15 (d) there is evidence that Epstein–Barr virus-immortalized B-cells and T-cell clones from HCV carriers show long-term persistence of HCV genomes (Bronowicki, personal communication and 5); and (e) we have presented strong evidence for the persistence of HCVRNA in PBMC obtained from HCV-positive subjects and injected into SCID mice. Artifacts due to contamination with serum particles were excluded by this approach which showed infection of these mononuclear cells.16
A second issue concerns the detection of replicative forms of HCV RNA in PBMC. This point has been discussed in regards to the specificity of negative HCV RNA detection. Thus, although such negative strands are clearly detected in some patients and on stimulation with mitogens, the real prevalence of this phenomenon remains unsettled.17 Interestingly, it has been suggested that HCV type 1 might show a distinct lymphotropism, in comparison to other frequent HCV types.17 Also, using a highly strand-specific method of RT–PCR, Shimizu et al.18 were able to demonstrate HCV RNA of negative polarity in PBMC samples from infected chimpanzees. In addition, they found that the capacity of infected PBMC and/or liver cells in chimpanzees, as well as in human lymphocyte cell lines infected in vitro, varied among different HCV strains suggesting the existence of lymphotropic HCV strains.18 The possibility of specific cell subsets harboring HCV sequences has also been discussed. HCV negative strand RNA, possibly reflecting viral replication, was found in all cell subsets in some studies11 and in specific cell subpopulations (B lymphocytes and monocytes) in other ones.17,19
Finally, the possibility of PBMC infection by HCV has recently been confirmed in studies indicating that HCV infection of lymphoid cells may favor selection of distinctive viral variants. In vitro and in vivo studies, based on HVR1 sequence analysis, showed different quasispecies patterns between serum, liver and PBMC.20,21 Thus, these observations support the concept of a ‘compartmentalization’ of some HCV genomes. The differences in quasispecies composition between these tissues may arise from cellular factors in the liver and PBMC, which would favor the growth of certain variants over others.20 It is not established, however, that distinct cellular tropism might be shown by some HCV variants.
Other cell types are permissive to HCV infection; we have recently demonstrated the possibility to infect, at a low level, biliary cells in primary culture.22,23 Biliary lesions are frequently observed during chronic HCV infection24 and HCV infection of the biliary cells might be involved in this phenomenon. Overall, infection of non-hepatocytic cells might constitute a ‘reservoir’ which would favor selection of HCV variants and viral persistence. The almost universal recurrence of HCV infection after orthotopic liver transplantation corroborates the existence of extra-hepatic sites where HCV can persist and replicate.25,26 PBMC and bone marrow cell infection by HCV might also induce specific cell alterations that can be interpreted as the potential pathogenetic strategies of the wide range of lymphoproliferative disorders frequently appearing during the course of chronic hepatitis C virus infection (for review see15,27). Some mechanisms underlying the lymphoproliferative disorders associated to HCV chronic infection have been recently proposed and involve the anti-apoptotic protein BCL2. In fact, a chromosomal rearrangement, t(14;18), resulting in the translocation of the bcl-2 open-reading frame under the transcriptional control of the immunoglobulin heavy-chain gene (IgH) with consequent over-expression of bcl-2, has been found with high prevalence in chronic HCV infected subjects. The presence of t(14;18) translocation in B-cells was highly prevalent in HCV positive subject developing cryoglobulinemia and non-Hodgkin's lymphoma suggesting that bcl-2 overexpression may play a relevant role in extending B-cell clone life and increasing the probability of additional mutational events driving B-cells to malignancy.27,28,29 The relative contribution of the strong polyclonal B-cell stimulation during HCV infection and of putative direct signals triggered by HCV proteins on cellular membrane molecules (CD81, the presumed HCV receptor) remain to be investigated. Finally, the actual impact of PBMC infection on the quality of the immune response is unclear, even if some recent reports described a contribution of viral proteins in disturbing host immune response.
The interactions between HCV and major cellular metabolic networks
HCV interactions with interferon signaling
Chronically HCV-infected patients show an inappropriate interferon production and this has provided rationale for the use of interferon α in treating, now in combination with ribavirin, patients with HCV-related chronic active hepatitis. There is also emerging evidences that HCV might, as for many other viruses, evolve mechanisms to directly inhibit the interferon signaling in the infected cells. HCV polyprotein expression in an osteosarcoma cell line inhibits the antiviral effect of interferon α on vesicular stomatitis (VSV) and encephalomyocarditis (EMCV) viruses and decreases STAT3 DNA binding;30 this study did not identify, however, the viral protein(s) involved.
NS5A is a candidate HCV protein, which is possibly involved in both the inhibition of the antiviral effects of interferon and the control of cell growth and viability (Figure 2). The pioneering studies conducted by Drs Enomoto31,32 and Katze et al.33,34 raised the hypothesis that NS5A might directly modulate the interferon transduction pathway by associating to and inhibiting the kinase activity of the double strand, RNA-stimulated protein kinase (PKR). In Japan, Enomoto et al. first demonstrated that mutations in a region of NS5A, which they referred to as the ‘Interferon Sensitive Determining Region: ISDR’ correlated with the response to interferon treatment. Thus, patients whose major circulating HCV genomes exhibited mutations in this region achieved the best response to treatment. Others,35,36 but not all,37 studies from Japan supported these findings. However, certain caveats should be borne in mind in this respect, which markedly complicate analysis of these findings. Several reports have shown that the strict correlation between ISDR mutations and treatment efficacy reported in Japan could not be demonstrated in Europe or the United States.38,39 The reasons for differences between clinical studies performed in Japan and Europe are not clear. It has been suggested that different interferon doses and regimens could affect the outcome of such comparative studies. However, this point has not been proved. It is also important to consider several virological parameters. The impact of NS5A mutations is dependent on the HCV type. Studies in Japan have mostly been performed in patients infected with HCV 1b or, to a lesser extent, HCV type 2, but NS5A mutations are not or only rarely identified in HCV type 3 genomes. NS5A variability in domains located outside the ISDR, and in particular in the C terminal part of the protein, may also be implicated in the control of therapeutic efficacy.40 The conclusions of these different studies may also differ because of methodological problems: other results have indeed been obtained when analysing only the major form of the HCV genome or complete quasispecies complexity.41,42 Moreover, a recent study suggested that ‘European’ and ‘Japanese’ HCV 1b genomes could exhibit different biological properties, a correlation between ISDR mutations and therapeutic efficacy only being demonstrated for the ‘Japanese’ isolates.43 These findings clearly need to be substantiated by further analyses. Finally, other major virological factors, such as the viral load, should also be taken into account.35
Several reports have now reinforced these clinical observations. In vitro, NS5A expression is sufficient to inhibit the antiviral effect of interferon α on VSV and EMCV viruses.44,45 We have in particular established this point by expressing several ‘natural’ NS5A mutants, isolated from patients with or without response to interferon, in the well differentiated hepatocytic cell line HuH7.46 The present data do not show, however, a correlation with the in vivo response to therapy.
Several important and fundamental problems remain to be solved regarding the molecular basis for the biological effects of NS5A. NS5A is located in the perinuclear area, and is a 56–58 Kd phosphoprotein. The kinase(s) involved are not identified, although casein-kinase II may be involved.47 Both a basal and ‘hyperphosphorylated’ status of NS5A have been suggested, and expression of the complete, non-structural HCV region is necessary in cis for the full phosphorylation of NS5A.48
Some reports suggest that NS5A might interact with at least two major cellular signaling pathways. NS5A might bind the catalytic domain of PKR, inhibiting its dimerisation and thus its activation.34,49 Furthermore, the domain involved in this interaction in NS5A might include the ISDR region and a domain adjacent to its 3′ extremity. Taken together, these data suggest an elegant model whereby expression of the NS5A protein may abolish the antiviral effect of PKR on interferon stimulation, this effect being abrogated by mutations which would then favor treatment efficacy. The implication of NS5A in the resistance to antiviral therapy has been recently confirmed, by the same group, expressing the non-structural HCV proteins in a subgenomic replicon model.50 It is important to note, however, that recent studies conducted by independent laboratories could not confirm the role of PKR in NS5A-mediated IFN resistance. In fact, by transiently expressing full length NS5As or in the context of a subgenomic replicon system, both groups could not show any physical interaction between NS5A and PKR or a change in its kinase activity.46,51
N-terminally-deleted forms of NS5A also show transcriptional activity, but it remains to be established whether such truncated NS5A molecules do exist in vivo and how they might participate in NS5A biological properties. Importantly, a correlation between ISDR mutations and NS5A transcriptional activity has been reported.52
It has been also proposed that NS5A may partially inhibit the IFN-induced antiviral response transactivating the Interleukin-8 (IL-8) promoter and increasing IL-8 levels in HeLa cells.53 NS5A might bind the Grb-2 adaptor protein and modulates the ERK1/2-dependent signaling54 or induces the activation of NF-κB and STAT-3 via oxidative stress originated by intracellular calcium level disturbance.55
NS5A is not the only HCV protein implicated in the interference with IFN antiviral signaling; in fact, it has been proposed that the E2 envelope protein might also interact with PKR and inhibit its actions.56 Interestingly, recent studies of our group demonstrated, in vivo and in vitro, the physical interaction between PKR and HCV core proteins isolated from tumor tissue of HCV-related HCC patients. These natural mutants present in HCC tissue showed only few point mutations in comparison to HCV 1b reference sequence, but revealed different biological properties including the enhancement of PKR auto-phosphorylation and the phosphorylation of its physiological target, the translation initiation factor eIF2α.57 Along this line, it has been also shown that HCV core protein specifically activated the 2′-5′-oligoadenylate synthetase (2′-5′-OAS) gene promoter in a dose-dependent manner in different human hepatocyte cell lines.58
HCV interactions with cellular pathways regulating cell proliferation and viability
A large number of studies have shown that in most parts of the world, HCV constitutes a major risk factor for HCC. However, the figures remain sketchy in some regions. In Japan and Southern Europe, anti-HCV prevalence levels respectively of 80 and 60% have been found, in Northern Europe, recent data suggest that approximatively 30% of cases of HCCs are associated with chronic HCV infection.59,60 In Sub-Saharian Africa and South-East Asia, where HBV-associated HCC is common, the impact of HCV is much less marked (around 10%). Prospective studies have also emphasized the role of HCV in various geographical areas.60
Much data has accumulated over the past few years concerning the mechanisms which may be involved in HCV-related HCC.61 Clearly, chronic hepatitis, with its combination of chronic inflammation, liver cell necrosis and regeneration, and extensive fibrosis, constitutes a major step in this process.62 Some cases of HCV-related HCC, however, develop in the absence of associated cirrhosis and chronic hepatitis; interestingly, most of these tumors contain HCV genomes classified as 1b.63 Viral multiplication is sustained throughout the long-term course of chronic HCV infection and the development of cirrhosis and HCC. HCV genomes persist in tumor cells and can replicate, albeit at a lower rate than in non-tumoral livers.64,65 In contrast with HBV, the HCV genome does not integrate into cellular DNA and replication is thus necessary to its persistence. HCV proteins can be also detected in tumor cells, testifying to HCV genome expression in these cells (B Lebail et al. unpublished observations). A certain amount of evidence, indicates that, beside inducing chronic liver inflammation, some HCV proteins may modulate liver cell proliferation and viability by interfering directly with the major cellular transduction networks. Some studies have recently focused on core, NS3 and NS5A proteins.
HCV core (Figure 3)
In addition to its role in the packaging of viral RNA, the HCV core can indeed modulate cellular transduction pathways (reviewed in66). HCV core protein is mainly cytoplasmic, located on the endoplasmic reticulum membranes and around lipid vesicles.67,68 HCV core has been found to be associated with TNF-receptor I and TNF-related lymphotoxin-receptor, and this interaction modulates signaling by these cytokines.69,70 As discussed below, HCV core protein also associates with apolipoprotein AII. The viral core actually binds a number of cellular proteins, also including a cellular helicase71 and an heterogeneous nuclear ribonucleoprotein K.72 However, the relevance of these various interactions is still to be fully established. The HCV core transactivates a number of cellular promoters, including c-myc, and activates NF-κB, AP-1 and SRE elements, interacting with c-JNK, ERK and MAPK signaling.73 The interference exerted by HCV core on Raf/MAPK/Erk pathways has been also established by several groups in different cell types with different expression systems, which could partially justify some discrepancies in the obtained results.74,75,76,77,78
Expression of HCV core protein in Jurkat T-lymphocytic cell line activated Interleukin-2 promoter through the NF-AT pathway79 and suppressed the induction of Th-1 immunity by inhibition of Interleukin-12 and nitric oxide production from activated macrophages.80 In addition, it suppressed the p53 and p21 promoters.81,82,83
Whether or not a fraction of HCV core may also act in the nucleus during in vivo infection is still a matter for debate. C-terminally truncated core translocates to the nucleus and may exert distinct biological effects;67,68,84 it has not been, however, shown that such sequences are actually present during natural HCV infection. Some reports have nonetheless suggested that such truncated cores might be identified in tumor tissues from patients with HCC.85,86
In the last years, an increasing number of reports depicted HCV core as a pleiotropic modulator of cell growth and viability. In cooperation with v-Ha-ras, HCV core can induce the transformation of immortalized Rat1 fibroblasts and BALB/3T3 cells87,88 and, possibly, primary rat fibroblasts.89 It also interacts with and inactivates a bZIP nuclear transcription protein, LZIP, triggering cell transformation.90 It has been also shown that HCV core can bind to Retinoid-X-Receptor-α (RXR-α) and modulate RXR-α controled gene expression.91 HCV core can also modify cell cycle acting on different regulatory proteins such as pRb and cyclin E.92,93
Such in vitro results have been substantiated by in vivo studies in transgenic mice, which demonstrated the induction of HCC on expression of the core sequence only94 or of the full-length genome;95 as discussed in a previous section the development of HCC in these animal models is preceded by steatosis in the absence of liver inflammation. However, other reports on HCV core-expressing transgenic mice did not demonstrate such changes, and the reasons for these discrepancies remain unknown.96
Oxidative stress has been indicated as one of the possible mechanisms of HCV-induced hepatocarcinogenesis. In vivo data, obtained in HCV core expressing transgenic mice, provided evidence of elevated peroxide products in older animals, in which scavenger systems are less effective, potentially contributing to the development of HCC in these animals.97 Along this line, HCV core protein showed a mitochondrial localization and induced a marked increase in reactive oxygen species (ROS) levels in hepatoma cells. In addition, in mice expressing the structural HCV proteins, an augmented intrahepatic lipid peroxydation was found. Thus, the oxidative injury induced by HCV core protein may contribute to HCV pathogenesis.98
In line with these results, HCV core also modulates cell viability, although the results obtained are still under debate. It has been suggested that HCV core protein may inhibit or enhancing programmed cell death induced by different agents both in vitro and in vivo. In fact, HepG2 cells are sensitized to apoptosis induced by HCV structural proteins99 or by Fas100 and TNFα69 or in contrast, there is inhibition of the apoptosis induced in HepG2 and Jurkat cells by Fas, in HepG2, HEK-293 and MCF7 cells by TNF, and in HeLa, BRK and CHO cells by cisplatin and c-myc or has no effect on HepG2, Ramos, Wil2-NS and BL2 apoptosis.101,102,103,104,105,106 Recent in vivo data support the inhibitory effect of HCV structural proteins on Fas-mediated Cytochrome c release.107
Thus, HCV core appears to be a multifunctional, cytosolic protein, capable of modulating several major cellular transduction pathways, its eventual overall effect on cell proliferation and viability being dependent on its intracellular concentration, cell differentiation and the cellular environment.
HCV NS3
The expression of NS3 may also be important, in addition to its major role in viral protein maturation and genome replication. In vitro, the stable expression in NIH3T3 cells of the N-terminal part of NS3, encoding for one of the two viral serine-proteases implicated in HCV polyprotein processing, can induce a transformed phenotype.108 A similar transforming activity, dependent on its serine protease enzymatic activity, has been obtained expressing a full-length NS3 in rat fibroblasts.109 It has been also suggested that the N- and C-terminal moieties of NS3 may regulate the catalytic activity of Protein kinase A as well as the localization of p53 (reviewed in110). NS3 is also responsible for a p53-dependent specific suppression of p21waf1 promoter activity in NIH3T3 fibroblasts.111 A recent report indicates that NS3 may induce reactive oxygen species (ROS) production in human monocytes via activation of the NADPH oxidase.112
HCV NS5A
This is another candidate protein which, besides its effects on the antiviral properties of interferon we have discussed in a previous section, might also regulate cell proliferation and viability. An inhibitory effect of NS5A expression on cell apoptosis as well as a transforming effect in NIH3T3 has been reported.49,113 This effect might depend on the NS5A/PKR interaction49 and transactivation and repression of cellular promoters.45,114 PKR indeed does not only show antiviral activity but also regulates cell proliferation and viability.115 As stated in the previous section, the NS5A/PKR interactions have not been, however, yet confirmed. Recently, the regulatory potential of NS5A of cell proliferation and viability has been further investigated; stable and transient expression of NS5A in hepatic and non hepatic cell lines induced perturbation of cell cycle by targeting Cdk1/2-cyclin complexes.116 It has been also shown that NS5A may repress p21 promoter activity binding to p53117 and modulate cellular transcription interacting with SRCAP transcription factor.118
Still, collectively, the interaction between HCV proteins and interferon signaling brings us back to the general and most important issue of the potential dual effect of interferon in HCV infection which is antiviral on the one hand and anti-proliferative on the other. This issue has major therapeutical implications since it would reinforce the concept raised from some clinical investigations of a protective effect of Interferon-α on HCC development, even in the absence of a detectable virological response to therapy.
The biological effects of HCV proteins on both antiviral and antiproliferative signaling also raise the hypothesis that the influence of some HCV types on Interferon response (a well documented observation) might not be dissociated from the potential, and highly debated, effect of some genotypes on the intensity of liver lesions and, possibly, liver carcinogenesis (reviewed in119). The impact of HCV genotypes in the risk of developing HCC is indeed an important issue to analyse. A number of recent studies point to the severity of HCV type 1 associated liver lesions, including HCC. Several, but not all, cross-sectional analysis indeed have shown higher relative prevalence of HCV 1 in patients with cirrhosis than in those with moderate CAH. These findings are however difficult to interprete since the molecular epidemiology of HCV is presently changing with the recent introduction, at least in areas such as France and Italy, of other types such as HCV 3 by intravenous drug users; in these conditions, duration of HCV infection will differ from one type to the other and markedly influence the risk of cirrhosis.1,120 There is still evidence for a more aggressive course of type 1 induced CAH. After liver transplantation in Europe, reinfection of the liver graft by HCV 1 induces a more rapidly progressive CAH than other HCV types and this is not related to epidemiological factors nor to the level of viremia.26,121 In some recent prospective studies on the risk of developing HCC in patients with HCV related cirrhosis, infection by genotype 1 has emerged as an independent risk factor;122,123 along the same line, HCV 1b was the most prevalent type among the patients we reported with HCV associated HCC in the absence of cirrhosis.63 Clearly these are still indirect evidences, the interpretation of which is complicated by intervening factors, several of them (such as alcohol and HBV coinfections) being underestimated in most studies; altogether, they are suggestive of a particular profile of HCV 1 infection. In vitro studies are now mandatory to analyze comparatively the different biological properties of various HCV isolates. In particular, as discussed for HBV, a major technical limitation to these studies is the analysis of isolated domains of the viral genomes; given the size of the HCV RNA, studies on full length clones are feasible but will require extensive efforts.
HCV infection and liver lipid metabolism
There are several observations, which point out the potential importance of such association. Circulating serum HCV particles show heterogeneous density, which reflect in part the binding of a fraction of the virions to very low density lipoproteins (VLDL) and low density lipoproteins (LDL).124,125 Moreover, the LDL receptor has been proposed as one of the potential viral receptors.126,127 A further important hallmark of chronic HCV, and not HBV, infection is its association to steatosis in at least 50% of infected subjects.24 The pathogenic importance of steatosis in the course of HCV infection is not established; it has been, however, recently suggested by several studies that, in some patients with steatosis from various etiologies, liver lipid overload might contribute in the development of fibrosis.128,129 Several groups showed that HCV core, (and more recently NS5A) is present on the surface of lipid droplets and may alter lipid metabolism in vitro interacting with cellular proteins involved in lipid accumulation and storage in liver cells.67,130,131 This specific feature of the core protein seems to be attributed to a proline-rich domain, absent in the core protein of flavi- and pestivirus, but present in GBV-B core and in other proteins responsible for lipid storage in plants.132 In addition, expression of HCV core in some transgenic mice induces steatosis;94 interestingly, these mice eventually develop HCC in the absence of detectable chronic inflammation. Consistent with this result, transgenic mice containing the complete HCV genome95 also show steatosis followed by HCC. In contrast, however, other core-expressing transgenic mice do not show liver lesions96 emphasizing the importance of the core expression levels, the mice genetic as well as likely environmental parameters.
In this context, we have provided direct in vitro evidence for the role of HCV core expression in steatosis induction.67 Moreover, we have shown the association of HCV core to Apolipoprotein AII (Apo-AII) and the impact of this binding in the in vitro secretion of the viral protein.130 We have recently investigated further this issue in vivo by using HCV core-expressing transgenic mice as well as mice expressing both HCV core and the human Apo-AII. Our data demonstrate that HCV core expression does not affect betaoxydation but, instead, directly inhibits VLDL assembly and secretion. Furthermore, ectopic in vivo ApoAII expression induces core secretion in serum, decreases its intrahepatic level, and thus abrogates its effects on VLDL secretion.133 Collectively, these data lead to propose a model whereby HCV/apolipoprotein AII interactions might regulate core accumulation, the viral protein in turn impairing liver triglyceride secretion (Figures 4 and 5).
A model for the interactions between HCV core and liver lipid metabolism. (a) The figure suggests that binding of HCV core to apolipoprotein AII (AII) might trigger core secretion by a presently unknown mechanism. Thus, upon translation of the core-expressing sequence, core protein might follow different pathways: (1) As described in the literature, it will be located on the cytosolic face of the endoplasmic reticulum (ER); (2) a proportion of core will be located around triglyceride (TG)-containing vesicles; and (3) binding of core with AII will, in case of high AII expression, drives core secretion. (b) Increased expression of ApoAII, such as that obtained by fibrate treatment or, in transgenic mice, ectopic ApoAII expression, would therefore lower intracellular core concentration (right panel). On the other hand, core inhibits VLDL and thus triglyceride secretion (left panel) and this effect is a major factor in steatosis induction; the consequences of such triglyceride accumulation remain to be explored
Recent data confirm the presence of non-enveloped nucleocapsid in the serum of HCV infected patients.134 The presence of HCV core molecules in bloodstream of infected subjects may contribute in the establishment and/or maintenance of virus persistence in the host by interacting with the complement receptor (gC1qR) and reducing T-lymphocyte responsiveness.135
Further studies will now aim to elucidate the molecular bases of these observations and their potential impact in liver lesions development; it will be also interesting to investigate whether or not, as suggested,136 HCV type 3 does specifically disturb liver lipid metabolism.
Conclusions
The large amount of information nowadays available concerning HCV protein biological effect has clearly shown a direct role of these viral proteins in the pathogenesis of HCV chronic infection. In fact, in addition to stimulating immune response and inflammation, they interact with several cellular signal transduction pathways regulating cell proliferation and apoptosis. These interactions contribute to viral persistence in the host and to the viral pathogenetic action driving to chronic hepatitis, cirrhosis and liver cancer.
Abbreviations
- HCC:
-
hepatocellular carcinoma
- CAH:
-
chronic active hepatitis
- STAT:
-
signal transducer and activator of transcription
- Grb-2:
-
growth factor receptor-bound protein 2
- ERK:
-
extracellular signal-regulated kinase
- NF-κB:
-
nuclear factor κB
- TNFα:
-
tumor necrosis factor α
- MAPK:
-
mitogen activated protein kinase
- JNK:
-
c-jun N-terminal kinase
- NF-AT:
-
nuclear factor of activated T cells
References
Bréchot C (1997) Hepatitis C virus 1b, cirrhosis, and hepatocellular carcinoma. Hepatology 25: 772–774.
Pawlotsky JM, Germanidis G, Frainais PO, Bouvier M, Soulier A, Pellerin M & Dhumeaux D (1999) Evolution of the hepatitis C virus second envelope protein hypervariable region in chronically infected patients receiving alpha interferon therapy. J. Virol. 73: 6490–6499.
Lohmann V, Korner F, Koch J, Herian U, Theilmann L & Bartenschlager R (1999) Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science 285: 110–113.
Blight KJ, Kolykhalov AA & Rice CM (2000) Efficient initiation of HCV RNA replication in cell culture. Science 290: 1972–1974.
Pietschmann T, Lohmann V, Rutter G, Kurpanek K & Bartenschlager R (2001) Characterization of cell lines carrying self-replicating hepatitis C virus RNAs. J. Virol. 75: 1252–1264.
Krieger N, Lohmann V & Bartenschlager R (2001) Enhancement of hepatitis C virus RNA replication by cell culture- adaptive mutations. J. Virol. 75: 4614–4624.
Mercer DF, Schiller DE, Elliott JF, Douglas DN, Hao C, Rinfret A, Addison WR, Fischer KP, Churchill TA, Lakey JR, Tyrrell DL & Kneteman NM (2001) Hepatitis C virus replication in mice with chimeric human livers. Nat. Med. 7: 927–933.
Dandri M, Burda MR, Torok E, Pollok JM, Iwanska A, Sommer G, Rogiers X, Rogler CE, Gupta S, Will H, Greten H & Petersen J (2001) Repopulation of mouse liver with human hepatocytes and in vivo infection with hepatitis B virus. Hepatology 334: 981–988.
Ohashi K, Marion PL, Nakai H, Meuse L, Cullen JM, Bordier BB, Schwall R, Greenberg HB, Glenn JS & Kay MA (2000) Sustained survival of human hepatocytes in mice: A model for in vivo infection with human hepatitis B and hepatitis delta viruses. Nat. Med. 6: 327–331.
Fend F, Kremer M & Quintanilla-Martinez L (2000) Laser capture microdissection: methodical aspects and applications with emphasis on immuno-laser capture microdissection. Pathobiology 68: 209–214.
Zignego AL, Macchia D, Monti M, Thiers V, Mazzeti M, Foschi M, Maggi E, Romagnani S, Gentilini P & Bréchot C (1992) Infection of peripheral mononuclear blood cells by hepatitis C virus. J. Hepatol. 15: 382–386.
Zignego AL, Ferri C, Monti M, LaCivita L, Giannini C, Careccia G, Giannelli F, Pasero G, Bombardieri S & Gentilini P (1995) Hepatitis C virus as a lymphotropic agent: evidence and pathogenetic implications. Clin. Exp. Rheumatol. 13 (Suppl 13): S33–S37.
Moldvay J, Deny P, Pol S, Bréchot C & Lamas E (1994) Detection of hepatitis C virus RNA in peripheral blood mononuclear cells of infected patients by in situ hybridization. Blood 83: 269–273.
Agnello V, Chung R & Kaplan L (1992) A role for hepaitits C virus infection in type II cryoglobulinemia. New Engl. J. Med. 327: 1490–1495.
Zignego A & Bréchot C (1999) Extrahepatic manifestations of HCV infection: facts and controversies. J. Hepatol. 31: 1152–1154.
Bronowicki JP, Loriot MA, Thiers V, Grignon Y, Zignego AL & Brechot C (1998) Hepatitis C virus persistence in human hematopoietic cells injected into SCID mice. Hepatology 28: 211–218.
Lerat H, Rumin S, Habersetzer F, Berby F, Trabaud MA, Trepo C & Inchauspe G (1998) In vivo tropism of hepatitis C virus genomic sequences in hematopoietic cells: influence of viral load, viral genotype, and cell phenotype. Blood 91: 3841–3849.
Shimizu YK, Igarashi H, Kanematu T, Fujiwara K, Wong DC, Purcell RH & Yoshikura H (1997) Sequence analysis of the Hepatitis C virus genome recovered from serum, liver, and peripheral blood mononuclear cells of infected chimpanzees. J. Virol. 71: 5769–5773.
Morsica G, Tambussi G, Sitia G, Novati R, Lazzarin A, Lopalco L & Mukenge S (1999) Replication of hepatitis C virus in B lymphocytes (CD19+). Blood 94: 1138–1139.
Maggi F, Fornai C, Vatteroni ML, Giorgi M, Morrica A, Pistello M, Cammarota G, Marchi S, Ciccorossi P, Bionda A & Bendinelli M (1997) Differences in Hepatitis C virus quasispecies composition between liver, peripheral blood mononuclear cells and plasma. J. Gen. Virol. 78: 1521–1525.
Afonso AM, Jiang J, Penin F, Tareau C, Samuel D, Petit MA, Bismuth H, Dussaix E & Feray C (1999) Nonrandom distribution of hepatitis C virus quasispecies in plasma and peripheral blood mononuclear cell subsets. J. Virol. 73: 9213–9221.
Loriot MA, Bronowicki JP, Lagorce D, Lakehal F, Barba G, Merget M, Vons C, Franco D, Belghiti J, Housset C & Bréchot C (1999) Permissivity of human biliary epithelial cells to infection by hepatitis C virus. Hepatology 29: 1587–1595.
Bichr S, Rende-Fournier R, Vona G, Yamamoto AM, Depla E, Maertens G & Brechot C (2002) Detection of neutralizing antibodies to hepatitis C virus using a biliary cell infection model. J. Gen. Virol. 83: 1673–1678.
Goodman ZD & Ishak KG (1995) Histopathology of Hepatitis C virus infection. Sem. Liver Dis. 15: 70–81.
Feray C, Gigou M, Samuel D, Paradis V, Wilber J, David MF, Urdea M, Reynes M, Brechot C & Bismuth H (1994) The course of hepatitis C virus infection after liver transplantation. Hepatology 20: 1137–1143.
Feray C, Gigou M, Samuel D, Paradis V, Mishiro S, David MF, Reynes M, Okamoto H, Bismuth H & Brechot C (1995) Influence of the genotypes of hepatitis C virus on the severity of recurrent liver disease after liver transplantation. Gastroenterology 108: 1088–1096.
Zignego AL, Giannelli F, Marrocchi ME, Mazzocca A, Ferri C, Giannini C, Monti M, Caini P, Villa GL, Laffi G & Gentilini P (2000) T(14;18) translocation in chronic hepatitis C virus infection. Hepatology 31: 474–479.
Zignego A, Ferri C, Giannelli F, Giannini C, Marrocchi M, Monti M, Caini P, La Villa G, Laffi G & Gentilini P (2002) Prevalence of bcl-2 rearrangement in hepatitis C virus-related mixed cryoglobulinemia with or without complicating B cell lymphoma. Ann. Inter. Med. 137: 571–580.
Zuckerman E, Zuckerman T, Sahar D, Streichman S, Attias D, Sabo E, Yeshurun D & Rowe JM (2001) The effect of antiviral therapy on t(14;18) translocation and immunoglobulin gene rearrangement in patients with chronic hepatitis C virus infection. Blood 97: 1555–1559.
Heim MH, Moradpour D & Blum HE (1999) Expression of hepatitis C virus proteins inhibits signal transduction through the Jak-STAT pathway. J. Virol. 73: 8469–8475.
Enomoto N, Sakuma I, Asahina Y, Kurosaki M, Murakami T, Yamamoto C, Izumi N, Marumo F & Sato C (1995) Comparison of full-length sequences of interferon-sensitive and resistant hepatitis C virus 1b. Sensitivity to interferon is conferred by amino acid substitutions in the NS5A region. J. Clin. Invest. 96: 224–230.
Enomoto N, Sakuma I, Asahina Y, Kurosaki M, Murakami T, Yamamoto C, Ogura Y, Izumi N, Marumo F & Sato C (1996) Mutations in the nonstructural protein 5A gene and response to interferon in patients with chronic hepatitis C virus 1b infection. New Engl. J. Med. 334: 77–81.
Gale MJ, Korth MJ, Tang NM, Tan SL, Hopkins DA, Dever TE, Polyak SJ, Gretch DR & Katze MG (1997) Evidence that hepatitis C virus resistance to interferon is mediated through repression of the PKR protein kinase by the nonstructural 5A protein. Virology 230: 217–227.
Gale, Jr M, Blakely CM, Kwieciszewski B, Tan SL, Dossett M, Tang NM, Korth MJ, Polyak SJ, Gretch DR & Katze MG (1998) Control of PKR protein kinase by hepatitis C virus nonstructural 5A protein: molecular mechanisms of kinase regulation. Mol. Cell. Biol. 18: 5208–5218.
Chayama K, Tsubota A, Kobayashi M, Okamoto K, Hashimoto M, Miyano Y, Koike H, Kobayashi M, Koida I, Arase Y, Saitoh S, Suzuki Y, Murashima N, Ikeda K & Kumada H (1997) Pretreatment virus load and multiple amino acid substitutions in the interferon sensitivity-determining region predict the outcome of interferon treatment in patients with chronic genotype 1b hepatitis C virus infection. Hepatology 25: 745–749.
Kurosaki M, Enomoto N, Murakami T, Sakuma I, Asahina Y, Yamamoto C, Ikeda T, Tozuka S, Izumi N, Marumo F & Sato C (1997) Analysis of genotypes and amino acid residues 2209 to 2248 of the NS5A region of hepatitis C virus in relation to the response to interferon-beta therapy. Hepatology 25: 750–753.
Komatsu H, Fujisawa T, Inui A, Miyagawa Y & Onoue M (1997) Mutations in the nonstructural protein 5A gene and response to interferon therapy in young patients with chronic hepatitisc virus 1b infection. J. Med. Virol. 53: 361–365.
Squadrito G, Leone F, Sartori M, Nalpas B, Berthelot P, Raimondo G, Pol S & Brechot C (1997) Mutations in the nonstructural 5A region of hepatitis C virus and response of chronic hepatitis C to interferon α. Gastroenterology 113: 567–572.
Rispeter K, Lu M, Zibert A, Wiese M, Mendes de Oliveira J & Roggendorf M (1999) A suggested extension of the HCV ISDR does not alter our former conclusions on its predictive value for IFN response. J. Hepatol. 30: 1163–1164.
Duverlie G, Khorsi H, Castelain S, Jaillon O, Izopet J, Lunel F, Eb F, Penin F & Wychowski C (1998) Sequence analysis of the NS5A protein of European hepatitis C virus 1b isolates and relation to interferon sensitivity. J. Gen. Virol. 79: 1373–1381.
Polyak SJ, Mac Ardle S, Liu SL, Sullivan DG, Chung M, Hofgartner WT, Carithers RL, Mac Mahon B, Mullins JI, Corey L & Gretch DR (1998) Evolution of hepatitis C virus quasispecies in hypervariable region 1 and the putative interferon sensitivity-determining region during interferon therapy and natural infection. J. Virol. 72: 4288–4296.
Pawlotsky JM, Germanidis G, Neumann AU, Pellerin M, Frainais PO & Dhumeaux D (1998) Interferon resistance of hepatitis C virus genotype 1b: Relationship to nonstructural 5A gene quasispecies mutations. J. Virol. 72: 2795–2805.
Nakano I, Fukuda Y, Katano Y, Nakano S, Kumada T & Hayakawa T (1999) Why is the interferon sensitivity-determining region (ISDR) system useful in Japan?. J. Hepatol. 30: 1014–1022.
Paterson M, Laxton CD, Thomas HC, Ackrill AM & Foster GR (1999) Hepatitis C virus NS5A protein inhibits interferon antiviral activity, but the effects do not correlate with the clinical response. Gastroenterology 117: 1187–1197.
Song J, Fujii M, Wang F, Itoh M & Hotta H (1999) The NS5A protein of hepatitis C virus partially inhibits the antiviral activity of interferon. J. Gen. Virol. 80: 879–886.
Podevin P, Sabile A, Gajardo R, Delhem N, Abadie A, Lozach PY, Beretta L & Brechot C (2001) Expression of hepatitis C virus NS5A natural mutants in a hepatocytic cell line inhibits the antiviral effect of interferon in a PKR-independent manner. Hepatology 33: 1503–1511.
Ide Y, Tanimoto A, Sasaguri Y & Padmanabhan R (1997) Hepatitis C virus NS5A protein is phosphorylated in vitro by a stably bound protein kinase from HeLa cells and by cAMP-dependent protein kinase A-α catalytic subunit. Gene 201: 151–158.
Koch J & Bartenschlager R (1999) Modulation of hepatitis C virus NS5A hyperphosphorylation by nonstructural proteins NS3, NS4A, NS4B. J. Virol. 73: 7138–7146.
Gale, Jr M, Kwieciszewski B, Dossett M, Nakao H & Katze MG (1999) Antiapoptotic and Oncogenic Potentials of Hepatitis C Virus Are Linked to Interferon Resistance by Viral Repression of the PKR Protein Kinase. J. Virol. 73: 6506–6516.
Pflugheber J, Fredericksen B, Sumpter, Jr R, Wang C, Ware F, Sodora DL & Gale, Jr M (2002) Regulation of PKR and IRF-1 during hepatitis C virus RNA replication. Proc. Natl. Acad. Sci. USA 99: 4650–4655.
Francois C, Duverlie G, Rebouillat D, Khorsi H, Castelain S, Blum HE, Gatignol A, Wychowski C, Moradpour D & Meurs EF (2000) Expression of hepatitis C virus proteins interferes with the antiviral action of interferon independently of PKR-mediated control of protein synthesis. J. Virol. 74: 5587–5596.
Fukuma T, Enomoto N, Marumo F & Sato C (1998) Mutations in the interferon-sensitivity determining region of hepatitis C virus and transcriptionnal activity of the nonstructural region 5A protein. Hepatology 28: 1147–1153.
Polyak SJ, Khabar KS, Paschal DM, Ezelle HJ, Duverlie G, Barber GN, Levy DE, Mukaida N & Gretch DR (2001) Hepatitis C virus nonstructural 5A protein induces interleukin-8, leading to partial inhibition of the interferon-induced antiviral response. J. Virol. 75: 6095–6106.
Tan SL, Nakao H, He Y, Vijaysri S, Neddermann P, Jacobs BL, Mayer BJ & Katze MG (1999) NS5A, a nonstructural protein of hepatitis C virus, binds growth factor receptor-bound protein 2 adaptor protein in a Src homology 3 domain/ligand-dependent manner and perturbs mitogenic signaling. Proc. Natl. Acad. Sci. USA 96: 5533–5538.
Gong G, Waris G, Tanveer R & Siddiqui A (2001) Human hepatitis C virus NS5A protein alters intracellular calcium levels, induces oxidative stress, and activates STAT-3 and NF-kappa B. Proc. Natl. Acad. Sci. USA 98: 9599–9604.
Taylor DR, Shi ST, Romano PR, Barber GN & Lai MM (1999) Inhibition of the interferon-inducible protein kinase PKR by HCV E2 protein. Science 285: 107–110.
Delhem N, Sabile A, Gajardo R, Podevin P, Abadie A, Blaton M, Kremsdorf D, Beretta L & Brechot C (2001) Activation of the interferon-inducible protein kinase PKR by Hepatocellular carcinoma derived-Hepatitis C virus core protein. Oncogene 20: 5836–5845.
Naganuma A, Nozaki A, Tanaka T, Sugiyama K, Takagi H, Mori M, Shimotohno K & Kato N (2000) Activation of the interferon-inducible 2′-5′-oligoadenylate synthetase gene by hepatitis C virus core protein. J. Virol. 74: 8744–8750.
Brechot C, Jaffredo F, Lagorce D, Gerken G, Meyer zum Buschenfelde K, Papakonstontinou A, Hadziyannis S, Romeo R, Colombo M, Rodes J, Bruix J, Williams R & Naoumov N (1998) Impact of HBV, HCV and GBV-C/HGV on hepatocellular carcinomas in Europe: results of a European concerted action. J. Hepatol. 29: 173–183.
Colombo M & Romeo R (1994) Hepatitis C virus infection and hepatocellular carcinoma. In Primary liver cancer: Etiological and progression factors Bréchot C, ed BocaRaton, Fl, USA: CRC Press.
Bréchot C (1996) Hepatitis C virus: molecular biology and clinics. InAcute and chronic liver diseases. Molecular biology and clinics Vol 9: Schmid R, Bianchi L, Blum HE, Gerok W, Maier KP and Stalder GA, ed Basel: Kluwer Academic pp.7–27.
Paterlini P & Bréchot C (1994) Hepatitis B virus and primary liver cancer in hepatitis B surface antigen-positive and negative patients.. In Primary liver cancer: Etiological and progression factors Bréchot C, ed(Paris: CRC Press)pp.167–190
De Mitri S, Poussin K, Baccarini P, Pontisso P, D'Errico A, Simon N, Grigioni W, Alberti A, Beaugrand M, Pisi E, Brechot C & Paterlini P (1995) HCV-associated liver cancer without cirrhosis. Lancet 345: 413–415.
Paterlini P, Driss F, Pisi E, Franco D, Berthelot P & Bréchot C (1993) Persistence of hepatitis B and hepatitis C viral genomes in primary liver cancers from HBsAg negative patients: a study of a low endemic area. Hepatology 17: 20–29.
Tsuboi SO, Nagamori S, Miyazaki M, Mihara K, Fukaya K, Teruya K, Kosaka T, Tsuji T & Namba M (1996) Persistence of hepatitis c virus RNA in established human hepatocellular carcinoma cell lines. J. Med. Virol. 48: 133–140.
Lai MM & Ware CF (2000) Hepatitis C virus core protein: possible roles in viral pathogenesis. Curr. Top. Microbiol. Immunol. 242: 117–134.
Barba G, Harper F, Harada T, Kohara M, Goulinet S, Matsuura Y, Eder G, Schaff Z, Chapman M, Miyamura T & Bréchot C (1997) Hepatitis C virus core protein shows a cytoplasmic localization and associates to cellular lipid storage droplets. Proc. Natl. Acad. Sci. USA 94: 1200–1205.
Yasui K, Wakita T, Tsukiyama-Kohara K, Funahashi SI, Ichikawa M, Kajita T, Moradpour D, Wands JR & Kohara M (1998) The native form and maturation process of hepatitis C virus core protein. J. Virol. 72: 6048–6055.
Zhu N, Khoshnan A, Schneider R, Matsumoto M, Dennert G, Ware C & Lai MM (1998) Hepatitis C virus core protein binds to the cytoplasmic domain of tumor necrosis factor (TNF) receptor 1 and enhances TNF-induced apoptosis. J. Virol. 72: 3691–3697.
You LR, Chen CM & Lee YHW (1999) Hepatitis C virus core protein enhances NF-kappaB signal pathway triggering by lymphotoxin-beta receptor ligand and tumor necrosis factor alpha. J. Virol. 73: 1672–1681.
Owsianka AM & Patel AH (1999) Hepatitis C virus core protein interacts with a human DEAD box protein DDX3. Virology 257: 330–340.
Hsieh TY, Matsumoto M, Chou HC, Schneider R, Hwang SB, Lee AS & Lai MM (1998) Hepatitis C virus core protein interacts with heterogeneous nuclear ribonucleoprotein K. J. Biol. Chem. 273: 17651–17659.
Shrivastava A, Manna SK, Ray R & Aggarwal BB (1998) Ectopic expression of hepatitis C virus core protein differentially regulates nuclear transcription factors. J. Virol. 72: 9722–9728.
Giambartolomei S, Covone F, Levrero M & Balsano C (2001) Sustained activation of the Raf/MEK/Erk pathway in response to EGF in stable cell lines expressing the Hepatitis C Virus (HCV) core protein. Oncogene 20: 2606–2610.
Fukuda K, Tsuchihara K, Hijikata M, Nishiguchi S, Kuroki T & Shimotohno K (2001) Hepatitis C virus core protein enhances the activation of the transcription factor, Elk1, in response to mitogenic stimuli. Hepatology 33: 159–165.
Aoki H, Hayashi J, Moriyama M, Arakawa Y & Hino O (2000) Hepatitis C virus core protein interacts with 14-3-3 protein and activates the kinase Raf-1. J. Virol. 74: 1736–1741.
Hayashi J, Aoki H, Kajino K, Moriyama M, Arakawa Y & Hino O (2000) Hepatitis C virus core protein activates the MAPK/ERK cascade synergistically with tumor promoter TPA, but not with epidermal growth factor or transforming growth factor alpha. Hepatology 32: 958–961.
Erhardt A, Hassan M, Heintges T & Haussinger D (2002) Hepatitis C Virus Core Protein Induces Cell Proliferation and Activates ERK, JNK, and p38 MAP Kinases Together with the MAP Kinase Phosphatase MKP-1 in a HepG2 Tet-Off Cell Line. Virology 292: 272–284.
Bergqvist A & Rice CM (2001) Transcriptional activation of the interleukin-2 promoter by hepatitis C virus core protein. J. Virol. 75: 772–781.
Lee CH, Choi YH, Yang SH, Lee CW, Ha SJ & Sung YC (2001) Hepatitis C virus core protein inhibits interleukin 12 and nitric oxide production from activated macrophages. Virology 279: 271–279.
Ray R, Steele R & Meyer Kea (1998) Hepatitis C virus core protein represses p21 WAF1/Cip1/Sid1 promoter activity. Gene 208: 331–336.
Ray RB, Steele R, Meyer K & Ray R (1997) Transcriptional repression of p53 promoter by hepatitis C virus core protein. J. Biol. Chem. 272: 10983–10986.
Jung EY, Lee MN, Yang HY, Yu D & Jang KL (2001) The repressive activity of hepatitis C virus core protein on the transcription of p21(waf1) is regulated by protein kinase A-mediated phosphorylation. Virus Res. 79: 109–115.
Suzuki R, Matsuura Y, Suzuki T, Ando A, Chiba J, Harada S, Saito I & Miyamura T (1995) Nuclear localisation of the truncated hepatitis C virus core protein with its hydrophobic C-terminus deleted. J. Gen. Virol. 76: 53–61.
Rüster B, Zeuzem S & Roth WK (1996) Hepatitis C virus sequences encoding truncated core proteins detected in a hepatocellular carcinoma. Bioch. Bioph. Res. Comm. 219: 911–915.
Horie C, Iwahana H, Horie T, Shimizu I, Yoshimoto K, Ishikawa T, Nagai Y & Kakumu S (1996) Detection of different quasispecies of hepatitis C virus core region in cancerous and non-cancerous lesions. Bioch. Bioph. Res. Comm. 218: 674–681.
Chang S, Hu T & Hsieh TS (1998) Analysis of a core domain in Drosophila DNA topoisomerase II. Targeting of an antitumor agent ICRF-159. J. Biol. Chem. 273: 19822–19828.
Tsuchihara K, Hijikata M, Fukuda K, Kuroki T, Yamamoto N & Shimotohno K (1999) Hepatitis C virus core protein regulates cell growth and signal transduction pathway transmitting growth stimuli. Virology 258: 100–107.
Ray RB, Lagging LM, Meyer K & Ray R (1996) Hepatitis C virus core protein cooperates with ras and transforms primary rat embryo fibroblasts to tumorigenic phenotype. J. Virol. 70: 4438–4443.
Jin DY, Wang HL, Zhou Y, Chun AC, Kibler KV, Hou YD, Kung H & Jeang KT (2000) Hepatitis C virus core protein-induced loss of LZIP function correlates with cellular transformation. EMBO J. 19: 729–740.
Tsutsumi T, Suzuki T, Shimoike T, Suzuki R, Moriya K, Shintani Y, Fujie H, Matsuura Y, Koike K & Miyamura T (2002) Interaction of hepatitis C virus core protein with retinoid X receptor alpha modulates its transcriptional activity. Hepatology 35: 937–946.
Cho J, Baek W, Yang S, Chang J, Sung YC & Suh M (2001) HCV core protein modulates Rb pathway through pRb down-regulation and E2F-1 up-regulation. Biochim. Biophys. Acta 1538: 59–66.
Cho JW, Baek WK, Suh SI, Yang SH, Chang J, Sung YC & Suh MH (2001) Hepatitis C virus core protein promotes cell proliferation through the upregulation of cyclin E expression levels. Liver 21: 137–142.
Moriya K, Fujie H, Shintani Y, Yotsuyanagi H, Tsutsumi T, Ishibashi K, Matsuura Y, Kimura S, Miyamura T & Koike K (1998) The core protein of hepatitis C virus induces hepatocellular carcinoma in transgenic mice. Nat. Med. 4: 1065–1070.
Lerat H, Honda M, Beard MR, Loesch K, Sun J, Yang Y, Okuda M, Gosert R, Xiao SY, Weinman SA & Lemon SM (2002) Steatosis and liver cancer in transgenic mice expressing the structural and nonstructural proteins of hepatitis C virus. Gastroenterology 122: 352–365.
Pasquinelli C, Shoenberger JM, Chung J, Chang KM, Guidotti LG, Selby M, Berger K, Lesniewski R, Houghton M & Chisari FV (1997) Hepatitis C virus core and E2 protein expression in transgenic mice. Hepatology 25: 719–727.
Moriya K, Todoroki T, Tsutsumi T, Fujie H, Shintani Y, Miyoshi H, Ishibashi K, Takayama T, Makuuchi M, Watanabe K, Miyamura T, Kimura S & Koike K (2001) Increase in the concentration of carbon 18 monounsaturated fatty acids in the liver with hepatitis C: analysis in transgenic mice and humans. Biochem. Biophys. Res. Commun. 281: 1207–1212.
Okuda M, Li K, Beard MR, Showalter LA, Scholle F, Lemon SM & Weinman SA (2002) Mitochondrial injury, oxidative stress, and antioxidant gene expression are induced by hepatitis C virus core protein. Gastroenterology 122: 366–375.
Kalkeri G, Khalap N, Garry RF, Fermin CD & Dash S (2001) Hepatitis C virus protein expression induces apoptosis in HepG2 cells. Virology 282: 26–37.
Ruggieri A, Harada T, Matsuura Y & Miyamura T (1997) Sensitization to Fas-mediated apoptosis by hepatitis C virus core protein. Virology 229: 68–76.
Ray R, Ghosh A, Meyer K & Ray R (1999) Functional analysis of a transrepressor domain in the hepatitis C virus core protein. Virus Res. 59: 211–217.
Ray R, Meyer K, Steele R, Shrivastava A, Aggarwal B & Ray R (1998) Inhibition of tumor necrosis factor (TNF-a)-mediated apoptosis by hepatitis C virus core protein. J. Biol. Chem. 273: 2256–2259.
Marusawa H, Hijikata M, Chiba T & Shimotohno K (1999) Hepatitis C virus core protein inhibits Fas- and tumor necrosis factor alpha-mediated apoptosis via NF-kappaB activation. J. Virol. 73: 4713–4720.
Zhu N, Ware CF & Lai MM (2001) Hepatitis C virus core protein enhances FADD-mediated apoptosis and suppresses TRADD signaling of tumor necrosis factor receptor. Virology 283: 178–187.
Dumoulin FL, vsn dem Bussche A, Sohne J, Sauerbruch T & Spengler U (1999) Hepatitis C virus core protein does not inhibit apoptosis in human hepatoma cells. European. J. Clin. Invest. 29: 940–946.
Giannini C, Caini P, Giannelli F, Fontana F, Kremsdorf D, Brechot C & Zignego AL (2002) Hepatitis C virus core protein does not significantly modify main proliferative and apoptosis pathways. J. Gen. Virol. 83: 1665–1671.
Machida K, Tsukiyama-Kohara K, Seike E, Tone S, Shibasaki F, Shimizu M, Takahashi H, Hayashi Y, Funata N, Taya C, Yonekawa H & Kohara M (2001) Inhibition of cytochrome c release in Fas-mediated signaling pathway in transgenic mice induced to express hepatitis C viral proteins. J. Biol. Chem. 276: 12140–12146.
Sakamuro D, Furukawa T & Takegami T (1995) Hepatitis C virus nonstructural protein NS3 transforms NIH 3T3 cells. J. Virol. 69: 3893–3896.
Zemel R, Gerechet S, Greif H, Bachmatove L, Birk Y, Golan-Goldhirsh A, Kunin M, Berdichevsky Y, Benhar I & Tur-Kaspa R (2001) Cell transformation induced by hepatitis C virus NS3 serine protease. J. Viral. Hepat. 8: 96–102.
Borowski P, Heiland M, Feucht H & Laufs R (1999) Characterisation of non-structural protein 3 of hepatitis C virus as modulator of protein phosphorylation mediated by PKA and PKC: evidences for action on the level of substrate and enzyme. Arch. Virol. 144: 687–701.
Kwun HJ, Jung EY, Ahn JY, Lee MN & Jang KL (2001) p53-dependent transcriptional repression of p21(waf1) by hepatitis C virus NS3. J. Gen. Virol. 82: 2235–2241.
Bureau C, Bernad J, Chaouche N, Orfila C, Beraud M, Gonindard C, Alric L, Vinel JP & Pipy B (2001) Nonstructural 3 protein of hepatitis C virus triggers an oxidative burst in human monocytes via activation of NADPH oxidase. J. Biol. Chem. 276: 23077–23083.
Ghosh AK, Steele R, Meyer K, Ray R & Ray RB (1999) Hepatitis C virus NS5A protein modulates cell cycle regulatory genes and promotes cell growth. J. Gen. Virol. 80: 1179–1183.
Polyak SJ, Paschal DM, McArdle S, Gale, Jr MJ, Moradpour D & Gretch DR (1999) Characterization of the effects of hepatitis C virus nonstructural 5A protein expression in human cell lines and on interferon-sensitive virus replication. Hepatology 29: 1262–1271.
Kaufman RJ (1999) Double-stranded RNA-activated protein kinase mediates virus-induced apoptosis: a new role for an old actor [comment]. Proc. Natl. Acad. Sci. USA 96: 11693–11695.
Arima N, Kao CY, Licht T, Padmanabhan R & Sasaguri Y (2001) Modulation of cell growth by the hepatitis C virus nonstructural protein NS5A. J. Biol. Chem. 276: 12675–12684.
Majumder M, Ghosh AK, Steele R, Ray R & Ray RB (2001) Hepatitis C virus NS5A physically associates with p53 and regulates p21/waf1 gene expression in a p53-dependent manner. J. Virol. 75: 1401–1407.
Ghosh AK, Majumder M, Steele R, Yaciuk P, Chrivia J, Ray R & Ray RB (2000) Hepatitis C virus NS5A protein modulates transcription through a novel cellular transcription factor SRCAP. J. Biol. Chem. 275: 7184–7188.
Bréchot C (1998) Molecular mechanisms of hepatitis B and C viruses related to liver carcinogenesis. Hepato-gastroenterology 45: 1189–1196.
Nousbaum J, Pol S, Nalpas B, Landais P, Berthelot P & Bréchot C (1995) Hepatitis C virus type 1b (II) infection in France and Italy. Ann. Inter. Med. 122: 161–168.
Gane EJ, Naoumov NV, Qian KP, Mondelli MU, Maertens G, Portmann BC, Lau JYN & Williams R (1996) A longitudinal analysis of hepatitis C virus replication following liver transplantation. Gastroenterology 110: 167–177.
Bruno S, Silini S, Crosignani A, Borzio F, Leandro G, Bono F, Asti M, Rossi S, Largui A, Cerino A, Podda M & Mondelli M (1997) Hepatitis C Virus genotypes and risk of hepatocellular carcinoma in cirrhosis: a prospective study. Hepatology 25: 754–758.
Yamauchi M, Nakahara M, Hisato N, Kazuhiko S, Junnichi H & Toda G (1994) Different prevalence of hepatocellular carcinoma between patients with liver cirrhosis due to genotype II and III of hepatitis C virus. Inter. Hepatol. Comm. 2: 328–332.
Thomssen R, Bonk S & Thiele A (1993) Density heterogeneities of hepatitis C virus in human sera due to the binding of beta-lipoproteins and immunoglobulins. Med. Microbiol. Immunol. (Berl) 182: 329–334.
Thomssen R, Bonk S, Propfe C, Heermann KH, Kochel HG & Uy A (1992) Association of hepatitis C virus in human sera with beta-lipoprotein. Med. Microbiol. Immunol. 181: 293–300.
Monazahian M, Bohme I, Bonk S, Koch A, Scholz C, Grethe S & Thomssen R (1999) Low density lipoprotein receptor as a candidate receptor for hepatitis C virus. J. Med. Virol. 57: 223–229.
Agnello V, Abel G, Elfahal M, Knight GB & Zhang QX (1999) Hepatitis C virus and other flaviviridae viruses enter cells via low density lipoprotein receptor. Proc. Natl. Acad. Sci. USA 96: 12766–12771.
Czaja AJ, Carpenter HA, Santrach PJ & Moore SB (1998) Host- and disease-specific factors affecting steatosis in chronic hepatitis C. J. Hepatol. 29: 198–206.
Lettéron P, Fromenty B, Terris B, Degott C & Pessayre D (1996) Acute and chronic hepatic steatosis lead to in vivo lipid peroxidation in mice. J. Hepatol. 24: 200–208.
Sabile A, Perlemuter G, Bonbo F, Kohara K, Demaugre F, Matsuura Y, Bréchot C & Barba G (1999) Hepatitis C virus core protein binds to apolipoprotein A11 ands its secretion is modulated by fibrates. Hepatology 4: 1064–1076.
Shi ST, Polyak SJ, Tu H, Taylor DR, Gretch DR & Lai MM (2002) Hepatitis C Virus NS5A Colocalizes with the Core Protein on Lipid Droplets and Interacts with Apolipoproteins. Virology 292: 198–210.
Hope RG, Murphy DJ & McLauchlan J (2002) The domains required to direct core proteins of hepatitis C virus and GB virus-B to lipid droplets share common features with plant oleosin proteins. J. Biol. Chem. 277: 4261–4270.
Perlemuter G, Sabile A, Letteron P, Vona G, Topilco A, Chretien Y, Koike K, Pessayre D, Chapman J, Barba G & Brechot C (2002) Hepatitis C virus core protein inhibits microsomal triglyceride transfer protein activity and very low density lipoprotein secretion: a model of viral-related steatosis. FASEB J. 16: 185–194.
Maillard P, Krawczynski K, Nitkiewicz J, Bronnert C, Sidorkiewicz M, Gounon P, Dubuisson J, Faure G, Crainic R & Budkowska A (2001) Nonenveloped nucleocapsids of hepatitis c virus in the serum of infected patients. J. Virol. 75: 8240–8250.
Kittlesen DJ, Chianese-Bullock KA, Yao ZQ, Braciale TJ & Hahn YS (2000) Interaction between complement receptor gC1qR and hepatitis C virus core protein inhibits T-lymphocyte proliferation. J. Clin. Invest. 106: 1239–1249.
Rubbia-Brandt L, Giostra E, Quadri R, Malé P, Mentha G, Sparhr J, Zarski J, Borisch B, Hadengue A & Negro F (1999) Evidence that liver steatosis is virally-mediated in a subset of patients infected by hepatitis C virus (HCV) genotype 3. J. Hepatol. 30 (Suppl 1): 133
Author information
Authors and Affiliations
Corresponding author
Additional information
Edited by M Piacentini
Rights and permissions
About this article
Cite this article
Giannini, C., Bréchot, C. Hepatitis C virus biology. Cell Death Differ 10 (Suppl 1), S27–S38 (2003). https://doi.org/10.1038/sj.cdd.4401121
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cdd.4401121
Keywords
This article is cited by
-
Hepatitis C virus core protein interacts with Snail and histone deacetylases to promote the metastasis of hepatocellular carcinoma
Oncogene (2016)
-
ADP-ribosylation Factor-related Protein 1 Interacts with NS5A and Regulates Hepatitis C Virus Propagation
Scientific Reports (2016)
-
Estimating the Size of the HCV Infection Prevalence: A Modeling Approach Using the Incidence of Cases Reported to an Official Notification System
Bulletin of Mathematical Biology (2016)
-
Inhibitory effect of kaolin minerals compound against hepatitis C virus in Huh-7 cell lines
BMC Research Notes (2014)
-
Role of different regions of the hepatitis C virus genome in the therapeutic response to interferon-based treatment
Archives of Virology (2014)