Abstract
To understand the expansion of multicopy microRNA (miRNA) families in plants, we localized the reported miRNA genes from Arabidopsis and rice to their chromosomes, respectively, and observed that 37% of 117 miRNA genes from Arabidopsis and 35% of 173 miRNA genes from rice were segmental duplications in the genome. In order to characterize whether the expression diversification has occurred among plant multicopy miRNA family members, we designed PCR primers targeting 48 predicted miRNA precursors from 10 families in Arabidopsis and rice. Results from RT-PCR data suggest that the transcribed precursors of members within the same miRNA family were present at different expression levels. In addition, although miR160 and miR162 sequences were conserved in Arabidopsis and rice, we found that the expression patterns of these genes differed between the two species. These data suggested that expression diversification has occurred in multicopy miRNA families, increasing our understanding of the expression regulation of miRNAs in plants.
Similar content being viewed by others
Introduction
MicroRNA (miRNA) comprises an extensive family of important regulatory noncoding RNAs of approximately 21-22 nucleotides in length and is found both in animals and in plants 1, 2, 3, 4, 5, 6. These are generated by RNase III-like enzymes from long primary transcripts encoded in intergenic regions (IGRs) or introns of the mRNA primary transcripts 5, 7, 8, 9. Mature miRNAs can downregulate gene expression by pairing with messages of protein-coding genes to promote mRNA cleavage, or repression of productive translation upon incorporation into a ribonucleoprotein complex (miRNP) similar to the RNA-induced silencing complex (RISC) 10, 11, 12, 13, 14. Indeed, the vast majority of the predicted or verified miRNA targets encode members of large families of transcription factors that regulate development 15. Mutants that lack miRNA pathway proteins, such as DICER-LIKE 1 (DCL1), ARGONAUTE 1 (AGO1) and HUA ENHANCER 1 (HEN1), exhibit morphological defects in Arabidopsis and rice, suggesting an important role for miRNA in plant development 8, 14, 16, 17, 18, 19, 20. Recently, more and more plant miRNAs have been reported for their specialized developmental functions, such as miR159, miR-JAW, miR160, miR162, miR165/166, miR168, and miR172. These miRNAs are involved in flower anther development 21, rosette leaf curvature 22, organ polarity 23, 24, floral organ identity and flowering time 25, 26, auxin response 27, and feedback regulation of the miRNA pathway 19, 28.
In Arabidopsis and rice, exceeding 100 miRNAs have been cloned or predicted. Although some have a single copy in the genome, many miRNA sequences correspond to multiple loci of 2-19 copies (http://microrna.sanger.ac.uk/cgi-bin/sequences/browse.pl). The mature miRNA sequence in each locus is predicted to pair with a nearby genomic segment, forming a dsRNA stem-loop structure, which is contained in miRNA precursors (pri-miRNA and pre-miRNA) and is required for Dicer processing. The mature miRNA can then be processed from either the 5′ or the 3′ arm of this structure 9. This type of multiple loci phenomenon has been widely observed in plants and animals 9, 29, 30, 31. Although recent research has demonstrated that nearly all the miRNA genes of Arabidopsis are expressed 32, little is known, in plants, about whether the expression or function is different among members within the same miRNA family.
In this paper, we observed that the expansion of miRNAs to multiple loci might be partly resulted from genome duplication in Arabidopsis and rice, and RT-PCR was utilized to estimate the expression of miRNA members within gene families. We demonstrated there were different expression levels of the precursors of multiple members, within the same family. In Arabidopsis and rice, differences in the expression patterns of the conserved miRNA families in two species suggest that the regulation of miRNA expression in plants is highly complex. Our data help to further elucidate the expression regulatory mechanisms of miRNA genes in plants.
Materials and methods
Plant materials and growth condition
Wild-type Arabidopsis thaliana plants (Columbia) and transgenic plants were grown in long days (16 h light, 8 h dark) at 22 °C with cool white light. Rice (Oryza sativa, japonica cultivar 9522) was grown in a greenhouse at 30 °C during the day and 24 °C at night.
miRNA genes location and duplication analysis
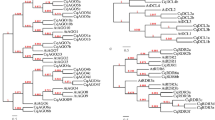
Duplicated block map of Arabidopsis, provided by TIGR (http://www.tigr.org/tdb/e2k1/ath1/arabGenomeDups.html), and rice predicted chromosome duplicated regions deduced from bioinformatics analysis of transcription factors 33 were used for analyzing the reported miRNA gene location of Arabidopsis and rice (http://microrna.sanger.ac.uk/cgi-bin/sequences/browse.pl) to deduce segmental duplications on the chromosomes. In these maps (Figure 1C and D), the boxes of the same color indicate the duplicated regions.
miRNA gene location and duplication analysis in Arabidopsis and rice genomes. (A) Statistical chart of the copy number of the identified miRNAs in Arabidopsis genome. (B) Statistical chart of the copy number of the identified miRNAs in rice genome. (C) and (D) Chromosome locations and duplication events for miRNAs indicated in Arabidopsis and rice genomes. Segmentally duplicated regions in the chromosomes are indicated by boxes of the same color. The scale of Arabidopsis (C) is adapted from the scale available on the NCBI database, and that of rice (D) is from TIGR, both are in megabases (Mb). Connecting lines mark the specific cases in which there is a strong correlation between duplicated genomic regions and the presence of miRNA genes in the same family.
The stem-loop regions of miRNA genes retained as duplicated pairs or nonduplicated pairs were selected for alignment using the Clustal X program (http://www-igbmc.u-strasbg.fr/BioInfo) 34. GendDoc (http://www.psc.edu/biomed/genedoc) 35 was used to highlight the conserved and similar nucleotide sequences (Figure 2).
Alignment of the predicted pre-miRNAs sequences within deduced duplicated pairs. The loop sequence of the pre-miRNA was in the box. (A) Alignment of At-MIR159a (chromosome 1: 184 bp from 27716894 to 27717077), At-MIR159b (chromosome 1: 196 bp from 6220639 to 6220834), and At-MIR159c (chromosome 2: 225 bp from 19001705 to 19001929), the At-MIR159a and At-MIR159b were predicted to be the duplicated pair. (B) and (C) Sequence alignment of duplicated pairs from Os-miR156 family. (B) Os-MIR156d (chromosome 2: 129 bp from 4512881 to 4513009) and Os-MIR156h (chromosome 6: 101 bp from 26510033 to 26510133). (C) Os-MIR156i (chromosome 2: 90 bp from 24065107 to 24065196) and Os-MIR156e (chromosome 4: 104 bp from 24650629 to 24650732). (D) Alignment of sequences from different duplicated pairs of Os-miR156 family, Os-MIR156d and Os-MIR156e.
Primer design
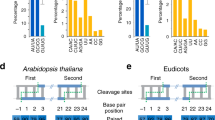
The hairpin structures of each miRNA were referenced from a miRNA database (http://microrna.sanger.ac.uk/cgi-bin/sequences/browse.pl). PCR primers were designed with Primer Express version 2.0 software (Applied Biosystems, Foster City, CA, USA). RT-PCR primers for detecting precursors were designed to target the nucleotide sequence within the predicted hairpin structure of these miRNA family members, so that the designed primer pairs could amplify both the long primary transcripts (pri-miRNA) and the short processed transcripts (pre-miRNA) (Figure 3). The term “miRNA precursors” refers to both pri-miRNA and pre-miRNA 36, and the criteria used to design sense and antisense primers were similar to what has been reported previously 36.
Primer design and miRNA processing in plants taking At-MIR168a as an example. (A) Plant miRNA primary transcripts (pri-miRNA) are proposed to be stepwise processed by the same enzyme DCL1 in nucleus that generated (B) pre-miRNA and (C) miRNA:miRNA* duplex, respectively. Both the pri-miRNA and pre-miRNA contain the stem-loop structure. The mature miRNA sequence is in red, and the hybridization sequence of sense and antisense primers are underlined. (D) RT-PCR detection of At-miR168 family, the PCR products for cDNA (lanes 1, 4), genomic DNA (lanes 2, 5), and the same reactions without RT (lanes 3, 6) were resolved on a 10% PAGE gel.
The designed antisense primers were used as gene-specific primers to promote the RT reaction. To ensure accurate amplification of each miRNA precursor, at least one primer sequence of the primer pair was designed to partly mismatch the mature miRNA or its complementary (miRNA*) sequence (Figure 3). The amplicon size of miRNA precursors ranged from 63 to 179 bp based on the various lengths of the predicted miRNA hairpin structures in plants. To minimize differences in RT efficiency, we utilized the same gene-specific primers between different members from the same family as possible. In the case of At-miR159, At-miR162, At-miR165/166, Os-miR162, Os-miR166, and Os-miR172 families, the same antisense primer (complementary to mature miRNA sequence) was designed for each member of the same family. Since the mature miRNA sequence was located on the 5′ arm of the hairpin structure of the members of At-miR157, At-miR160, At-miR168, and Os-miR160 families, and the complementary (miRNA*) sequence on the 3′ arm was partly variable among those members, making it was difficult to design one antisense primer for reverse transcription. For this reason, different antisense primers (At-MIR157aRTA, At-MIR157bRTA, At-MIR157cRTA, At-MIR157dRTA, At-MIR160aRTA, At-MIR160bRTA, At-MIR160cRTA, At-MIR168aRTA, At-MIR168bRTA, Os-MIR160a/b/c/dRTA, Os-MIR160eRTA, and Os-MIR160fRTA) were designed for each member respectively. The difference between the annealing temperatures of these antisense primers was within 2 °C and the GC content difference was less than 10%. In order to analyze At-MIR162a and At-MIR162b precursors, one sense primer At-MIR162RTS was designed on the At-miR162* location (sequence complementary to At-miR162). Since the At-miR162* sequence had high similarity between the At-MIR162a and At-MIR162b, the sense primer At-MIR162RTS and antisense primer At-MIR162RTA could be used to monitor both precursors of At-MIR162a and At-MIR162a. In order to specifically detect the At-MIR162a precursor, another sense primer, At-MIR162aRTS, that mismatched the At-miR162* sequence, was designed on the At-MIR162a stem-loop region; thus the primer pair At-MIR162aRTS and At-MIR162RTA could be used specifically for amplifying the At-MIR162a precursor. The Tm value of all primer pairs ranged from 54 to 59 °C, and all primers were confirmed to be capable of amplifying the target DNA fragments by using genomic DNA as a template.
RT-PCR
Total cellular RNA was isolated by guanidine thiocyanate/phenol-chloroform extraction as described 37, and treated with DNaseI (Promega, Shanghai, China). The RNA concentration was quantified using absorbance at 260 nm. First strand cDNA synthesis was performed with 3 μg RNA using RevertAid First Strand cDNA Synthesis Kit (Fermentas, Vilnius, Lithuania) according to the manufacturer's recommendations with either oligodT(18) primer or the gene-specific primers.
RT reaction was heated (70 °C for 5 min) to denature the RNA, incubated (37 °C for 5 min) to anneal the primers, and then cooled on ice. The remaining reagents (5 × reaction buffer, dNTPs, RNase inhibitor, M-MuLV reverse transcriptase) were then added and mixture was incubated (42 °C for 60 min). The reaction was heated (80 °C for 10 min) to inactivate reverse transcriptase and the RT products were purified by phenol/chloroform (1:1) and resolved in 100 μl of water.
PCR was performed using sense (RTS) and antisense (RTA) primers of each miRNA precursor. Reaction volumes (30 μl) contained 1×PCR Buffer, 200 μM of dATP, dGTP, dCTP, and dTTP (TaKaRa, Dalian, China), 1 mM of each primer, 2.5 μl diluted RT template in water, and 2.5 units of Taq DNA polymerase (TaKaRa). Amplifications were performed utilizing a PTC-100 thermocycler (MJ Research, Waltham, MA, USA) as follows: one step of 5 min at 94 °C, 34 cycles of 94 °C for 30 s, 54 – 58 °C (based on the Tm values of the primers) for 30 s, and 72 °C for 30 s. PCR amplicons were resolved in either 2.2% agarose or 10% polyacrylamide electrophoresis in 0.5 × TBE buffer and stained with ethidium bromide. Gel bands were evaluated by direct observation on a UV transilluminator, and then recorded using a computer imaging system. Each PCR pattern was verified in triplicate; negative controls (no template and no reverse transcription) were also included for each gene. The PCR products amplified with cDNA or genomic DNA were subcloned into pMD18-T vector (TaKaRa) and were confirmed by sequencing.
5′ RACE PCR and computational identification of conserved motifs in promoter region
5′ RACE PCR was performed with 5′ Full RACE Core set (TaKaRa) following the manufacturer's instructions (Figure 4A). 5′ RACE PCR primers were also designed according to the manufacturer's instructions. RT primers were located less than 60 bp downstream of the 3′ end of the predicted stem-loop sequence (Figure 4C and 4D) and were phosphorylated at the 5′ end, facilitating cDNA self-ligation. Based on the manufacturer's suggestion, the length of RT primers was designed to be less than 20 nt, with Tm values of approximately 45 °C. First round PCR primers were located in the predicted stem-loop structure of each member, and second round PCR primers were located with a maximal extension of 40 bp over the 5′ or 3′ end of the predicted stem-loop structure. The range of Tm values between the nested PCR primer pairs was within 4 °C.
Expression of some miRNA family members tested in Arabidopsis inflorescences, stems, cauline leaves, and rosette leaves. Three micrograms total RNA from inflorescences, stems, cauline leaves, and rosette leaves was analyzed by RT-PCR. At-MIR166a, At-MIR166b, At-MIR166e, and At-MIR168a could be detectable in all the tested organs, whereas the amplicon of At-MIR165a and At-MIR166f could only be visible in some organs. Other tested miRNA variants could not be detected in all the organs.
Total RNA was extracted as described above; cDNA was generated with 3 mg RNA using gene-specific primers, e.g. At-165aRTP, At-165bRTP, At-168aRTP, At-168bRTP, Os-162aRTP, and Os-162bRTP, and self-ligated by T4 RNA ligase. The first round PCR amplification using first PCR primer pairs was performed with RT products, respectively. To obtain the specific products, the nested PCR was performed using the first round PCR dilution with the second PCR primer pairs. Amplified products were gel purified and cloned into pMD18-T vector for sequencing.
Conserved sequences in the promoter regions were identified using a Bioprospector program as described previously 32.
Transgenic vector and plant transformation
To test whether the mature miR168 could be generated from both predicted stem-loop structures of At-MIR168a and At-MIR168b, a Col-0 genomic DNA fragment with 276 bp containing the predicted At-MIR168a hairpin precursor was cloned using the sense primer MIR168aS (5′ ttt tgg tac cCC CAT TAC AAA ACC TAG AAC 3′ from 162619 to 162638bp, Genbank Accession. AL161550) 81 bp upstream of the 5′ end of the hairpin structure and the antisense primer MIR168aA (5′ ttt gaa ttc TAA AGA AGG AGA AGC GTA GA 3′, from 162894 to 162875 bp, Genbank Accession. A L161550) 57 bp downstream of the 3′ end of the hairpin structure, containing the KpnI and EcoRI restriction sites, respectively. A 305 bp genomic DNA fragment of At-MIR168b with 81 bp upstream from the 5′ end of hairpin structure and 100 bp downstream from the 3′ end of hairpin precursor was amplified with genomic DNA from Col-0 plants using the sense primer MIR168bS (5′ ttt ggt acc TGC CGG ATT TGG TAA CTG TTT TCT CC 3′ from 40450 to 40425 bp, Genbank Accession. AB020744) and the antisense primer MIR168bA (5′ ttt gaa ttc CAA TAC CGA ATC AAT CAA ATC ACA CAG 3′ from 40146 to 40172 bp, Genbank Accession. AB020744), respectively; kpnI and EcoRI restriction enzyme sites were also introduced into this genomic DNA fragment.
The two PCR products were gel purified and then subcloned into the pMD18-T vector for sequencing. The resulting clones were digested with Kpn I and EcoR I. Recovered DNA fragments were ligated into a pMON530 Vector (Monsanto, St. Louis, MO, USA) digested with the same set of enzymes. The fragments were inserted between the cauliflower mosaic virus (CaMV) 35S promoter and the nos terminator.
Wild-type Arabidopsis plants were transformed with Agrobacterium strain GV3101 carrying the constructed plasmid utilizing the flower-dipping method 38.
Northern blot analysis of mature miR168
Total RNAs were isolated from rosettes of wild-type and transgenic plants. Total RNA (10 μg) was resolved in a 19% polyacrylamide gel and were blotted to a Nylon membrane (Schleicher&Schuell, Dassel, Germany). DNA oligonucleotides (5′ TTC CCG ACC TGC ACC AAG CGA 3′, from 21 to 1 bp, Genbank Accession. AJ493652) complementary to mature miR168 were end-labeled with [α-32p] ATP and T4 polynucleotide kinase (New England Biolabs, Beverly, MA, USA) to generate high-specific activity probes. Hybridization was carried out as described 29 and blots were rehybridized with a designed probe (5′ GAG GGA TGC AAC ACG AGG ACT T 3′, from 296 to 317 bp, Genbank Accession. AY130753) complementary to 5S rRNA.
Results
Chromosome location and duplication analysis of multicopy miRNA genes in Arabidopsis and rice genomes
On the basis of location information provided by the Sanger Institution (http://microrna.sanger.ac.uk/cgi-bin/sequences/browse.pl), we realized that approximately 50% of the identified miRNAs corresponded to multiple loci in Arabidopsis (24/45) (Figure 1A) and in rice (21/41) (Figure 1B) genome. According to this location information, we localized a total of 117 deduced miRNA genes from Arabidopsis and 173 identified miRNA genes from rice on five Arabidopsis chromosomes and 12 rice chromosomes, respectively. Furthermore, we investigated the relationship between the designated genome-wide intra and interchromosomally duplicated areas and the location of miRNA genes belonging to the same family. Our data suggest that many miRNA genes were located within the proposed genomic segmental duplicated regions (Figure 1C and 1D). Among the 173 miRNA genes in rice, 65% (113/173) of the miRNA genes resided on the deduced duplicated genomic segments 33, whereas 35% (60/173) were retained as duplicated pairs (Figure 1D). Similarly, the ratios in Arabidopsis were 82% (96/117) (http://www.tigr.org/tdb/e2k1/ath1/arabGenomeDups.html) and 37% (43/117) (Figure 1C). In addition, we observed clusters of related miRNA genes (one cluster of MIR169 and two clusters of MIR395) in both Arabidopsis and rice genomes, which may indicate the occurrence of tandem gene duplication events (Figure 1C and 1D).
Moreover, the similarity of the miRNA family genes distributed in the duplicated pairs was observed by aligning the predicted stem-loop sequence of these miRNA genes. For example, alignment of the At-MIR159 family (Figure 2A) revealed that the nucleotide sequence similarity was high (76.1%) between At-MIR159a and At-MIR159b, both of which had previously been proposed to be involved in a duplicated event. Whereas, less similarity (49.3%) was observed for those not proposed to be a duplicated pair, such as At-MIR159a and At-MIR159c. Similar results were observed in rice Os-MIR156 family members (Figure 2B and 2C), in which the sequence similarity between the stem-loop sequences of the same duplicated pairs was above 70% (81.7% between Os-MIR156i and Os-MIR156e and 72.9% between Os-MIR156d and Os-MIR156lh), but less than 40% similarity was noted between the sequences from different duplicated pairs (Figure 2D). Even though the loop region sequences of miRNA genes were proposed to be divergent for undergoing no functional constraint 40, we also observed the conservation among members of a duplicated pair (Figure 2A-2C). These data serve to validate the duplication events, especially when considering that the loop region is usually less conserved in miRNA genes.
Monitoring the expression of miRNA family members with multicopy
Through above results, it was observed that different evolution might occur among those miRNA genes in the same family. Therefore, we proposed that the expression or functional diversification of miRNA family members with multicopy existed. To prove this point, we chose to detect the expression level of miRNA precursors transcribed from each miRNA gene. Firstly, a sensitive detection system, RT-PCR, was set up and employed to amplify and detect miRNA precursors in plants (Figure 3). The PCR sensitivity was tested and proved to be sufficient for detecting at least 100 copies of the target fragment using genome DNA as template (data not shown). RT reactions were carried out with either OligodT(18) primer or gene-specific primers, respectively. We found that only three transcribed miRNA precursors, e.g. At-MIR166b, Os-MIR162a, and Os-MIR162b, could be amplified with the use of OligodT(18) primed RT products (data not shown), whereas a greater number of miRNA precursors could be amplified by employing gene-specific primers for the RT reaction (Table 1). This result may be explained by the fact that many miRNA primary transcripts in plants could have been processed rapidly and that the 3′ end structures of the transcribed products disappeared quickly 41. All the expected RT-PCR products were confirmed by sequencing. These results indicated that the established PCR system can be used for monitoring miRNA precursors in plants.
Since there are difficulties in designing primer pairs of the hairpin structures to discriminate the miRNA family members, finally 6 miRNA families with 23 predicted members from Arabidopsis and 4 miRNA families with 25 predicted members from rice were selected for detecting the expression level. These miRNAs were predicted to correspond to 2, 3 , 4, 6 , 7, and 13 copies in the genome. The results showed that the expression levels of miRNA members within the same family were different in plants (Table 1), which suggested that the expression diversification might occur among them. Interestingly, the precursors of At-MIR159a and At-MIR159b, both of which had been proposed to be a duplicated pair, could be detected, whereas, the precursor of At-MIR159c could not be detected, implying similar expression regulatory mechanisms may be performed on miRNA genes in the same duplication pair.
To further investigate the detailed expression pattern of miRNA family variants, At-miR165/166 family and At-miR168 family were selected to monitor the corresponding transcribed precursors in cauline leaves, rosette leaves, stem, and inflorescences (Figure 4). The miRNA precursors of At-MIR166a, At-MIR166b, At-MIR166e, and At-MIR168a could be monitored in all the tested organs, suggesting that these might play roles in all these organs. The amplicon of one miRNA precursor, At-MIR166f, which could not be detected in the sample of mixed tissues, was visible in inflorescences, suggesting that At-MIR166f might function specifically during reproductive stage. Additionally, we interestingly observed that the tested miRNA variants were expressed highly in inflorescences (Figure 4), reflecting their possible important roles in reproductive organs.
To date, approximately 30 miRNA families have been found to be conserved in Arabidopsis and rice (http://microrna.sanger.ac.uk/cgi-bin/sequences/browse.pl). Interestingly, although the precursors of all the At-miR160 and Os-miR162 family members could be detected, some of their homologous members, e.g. At-MIR162a, Os-MIR160b, Os-MIR160e and Os-MIR160f, were not detected in our experiment (Table 1), implying there might exist differences in miRNA expression regulation between monocot and dicot plants.
The 5′-terminal structure of miRNA primary transcripts in Arabidopsis and rice
In order to further confirm the RT-PCR results, we utilized a 5′ RACE PCR method to monitor the expression of some miRNA genes including At-miR165 and At-miR168 family members. Our data demonstrate that only the specific amplified DNA fragments, corresponding to the transcripts of At-MIR165a and At-MIR168a, were obtained after the two rounds of PCR amplification, but no product was generated to the At-MIR165b and At-MIR168b transcripts (Figure 5B). This result indicated that the transcript levels of At-MIR165a and At-MIR168a were high enough to be detected, whereas the levels of At-MIR165b and At-MIR168b were not high enough after the two rounds of PCR amplification. These data were also consistent with the RT-PCR analysis results of the At-miR165 and At-miR168 families. The same method was performed on Os-miR162 family members and both members were detected, similar to RT-PCR result.
Scheme of 5′ RACE PCR analysis. (A) Illustration of 5′ RACE PCR method used in this study. RT-PCR products were self-ligated, followed by two rounds of PCR amplification. (B) Image of agarose gel showing the 5′ RACE PCR products of At-MIR165a (216bp), At-MIR165b, At-MIR168a (206bp), and At-MIR168b. (C) and (D) Sequences around the transcription start sites of Os-MIR162a and Os-MIR162b. The primers used for reverse transcription and PCR are indicated. The filled arrow indicates the transcription start sites and distance from 5′ end of mature miRNA. The sequence of mature miRNA is in red; TATA box-like sequence was identified upstream from the transcribed region.
Based on the nucleotide sequence analysis of the 5′ RACE PCR amplified products, we observed two additional G bases in the 5′-terminal of tested transcripts, which were proposed to be the G cap structure. Moreover, TATA box-like sequences were observed approximately 10-30 bp upstream from the transcription start sites of Os-MIR162a and Os-MIR162b (Figure 5C and 5D). The 5′-terminal characters of Os-MIR162a and Os-MIR162b were similar to those tested miRNA genes in Arabidopsis 32, demonstrating that both in monocots and in dicots, miRNA genes may be transcribed by RNA polymerases II 32, 42.
Overexpressed precursors of both At-MIR168a and At-MIR168b can result in higher accumulation of miR168
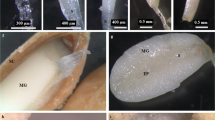
In the current study, we also investigated whether functional diversification occurred between miRNA members in the same family. Two genomic DNA fragments of approximately 300 bp containing the stem-loop structure of At-MIR168a and At-MIR168b were cloned under the control of the CaMV 35 s promoter. Both constructs were subsequently introduced into wild-type plants. As reported previously, miR168 can negatively regulate AGO1 expression 19; therefore, after selection with kanamycin, we observed that some transgenic plants displayed developmental defects similar to the AGO1 mutants in both 35S::MIR168a and 35S::MIR168b transformants. Total RNA was extracted from rosette leaves of wild type, ago1 mutant like 35S::MIR168a, and 35S::MIR168b, and the levels of miR168 were determined. Higher accumulation levels of precursors and mature miR168 were noted in 35S::MIR168a and 35S::MIR168b plants (Figure 6D and 6E). This suggested that even though the predicted precursor of At-MIR168b was beyond detection, the stem-loop structure of At-MIR168b still contain the sequence information responsible for mature miR168 generation. In addition, similar phenotypes caused by overexpressing both At-MIR168a and At-MIR168b also indicated that no obvious functional diversification occurred between At-MIR168a and At-MIR168b (Figure 6A-6C).
Development defects caused by overexpressing of At-MIR168a and At-MIR168b, and expression levels of miR168 precursors and mature miR168 in 35S::MIR168a and 35S::MIR168b transformants. (A)-(C) Similar ago1 mutant phenotypes were observed in both 35S::MIR168a and 35S::MIR168b transformants. (D) MiR168 precursors accumulation detected using RT-PCR in transgenic plants. The At-MIR168b precursor could be detected in 35S::MIR168b transformants. (E) Mature miR168 accumulation in both 35S::MIR168a and 35S::MIR168b transformants.
Discussion
The discovery of miRNAs in both plants and animals expands our knowledge of gene modulation. In plants, miRNA genes are mostly discrete, independent transcription units, and about 50% of miRNAs are predicted to be multiple loci in the genome, similar to that seen with protein-coding genes (recent rice sequencing data showed that 29% of protein-coding genes appear in gene families) 43. The evidence here suggesting that the expansion of miRNA genes in plants resulted from genome segmental and tandem duplication events during evolution is similar to that of many protein-coding gene families, especially transcription factor families 33, 44. Thus, these kinds of duplication events are proposed to have contributed to the expansion of both protein-coding gene families and miRNA gene families. The same expression status of At-MIR159a and At-MIR159b in a duplication pair increases this possibility too.
Recently, the transcripts of most miRNA genes in Arabidopsis have been detected by 5′ RACE and 3′ RACE approaches 32. However, it is difficult to demonstrate expression differences of miRNA genes utilizing this method as multiple (2-4) rounds of PCR amplification are not able to estimate differences in the initial cDNA templates. Detection of miRNA precursors by gel-based methods is also difficult, given the rapid processing of miRNA precursors in plants. In the current study, we have developed a method to better detect expression differences of miRNA family members in plants utilizing RT-PCR approach, and demonstrate that it can successfully identify different transcript levels within same miRNA family members. Even though our system could not detect very low level precursor molecule, these tested results provide direct evidence that the expression levels of miRNA members within the same family were different in plants, from which we infer that complex mechanisms may regulate the expression of these members.
The ability to detect some precursors may result from the fact that the corresponding loci may have been transcribed at relatively high levels in tested plant samples. On the other hand, undetectable precursors may be due to their low expression levels in tested samples or due to the fact that some loci may only be expressed in specific cells or during specific developmental stages, such as At-MIR166f, the transcript of which could only be detected in inflorescence. Additionally, some may be pseudogenes. Since the expression of nearly all miRNA genes in plants is proposed to be independently regulated by their own promoters, many miRNAs may have developmental stage-specific or cell (tissue, organ)-specific expression patterns determined by their promoter region. In animals, transcription studies of let-7 in C. elegans demonstrated that the temporal regulatory element (TRE) located at about 1200 nt upstream from the mature let-7 coding sequence was required for temporal upregulation of let-7 45. Recently, studies of the At-miR164 family showed that At-MIR164a and At-MIR164b participate in lateral root development 46, whereas At-MIR164c was demonstrated to control petal number in early flowers 47, indicating the nonredundant expression of the three members.
For protein-coding gene families, both the expression and functional diversification may occur among family members. Similar events may also occur with miRNA family members. However, here, molecular data have demonstrated that both the hairpin structures of At-MIR168a and At-MIR168b could contribute to mature miR168 processing. In addition, with phenotype analysis data, it was suggested that there is no obvious functional diversity between the At-MIR168a and At-MIR168b.
In order to gain further insight into the expression and functional diversity of multicopy miRNA families in plants, additional research is needed, like investigating the expression patterns of different miRNA members within the same families in details. Analysis of the miRNA promoter regions in plants may also reveal conserved elements required for regulating miRNA expression patterns.
References
Zamore PD . Ancient pathways programmed by small RNAs. Science 2002; 296:1265–1269.
Ambros V . MicroRNA pathways in flies and worms: growth, death, fat, stress, and timing. Cell 2003; 113:673–676.
Carrington JC, Ambros V . Role of microRNAs in plant and animal development. Science 2003; 301:336–338.
Bartel B, Bartel DP . MicroRNAs: at the root of plant development? Plant Physiol 2003; 132:709–717.
Bartel DP . MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004; 116:281–297.
Mallory AC, Vaucheret H . MicroRNAs: something important between the genes. Curr Opin Plant Biol 2004; 7:120–125.
Grishok A, Pasquinelli AE, Conte D, et al. Genes and mechanisms related to RNA interference regulate expression of the small temporal RNAs that control C. elegans developmental timing. Cell 2001; 106:23–34.
Park W, Li J, Song R, Messing J, Chen X . CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr Biol 2002; 12:1484–1495.
Reinhart BJ, Weinstein EG, Rhoades MW, Bartel B, Bartel DP . MicroRNAs in plants. Genes Dev 2002; 16:1616–1626.
Lee RC, Feinbaum RL, Ambros V . The C.elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993; 75:843–854.
Hutvágner G, Zamore PD . A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002; 297:2056–2060.
Llave C, Xie Z, Kasschau KD, Carrington JC . Cleavage of Scarecrow-like mRNA targets directed by a class of Arabidopsis miRNA. Science 2002; 297:2053–2056.
Kasschau KD, Xie Z, Allen E, et al. P1/HC-Pro, a viral suppressor of RNA silencing, interferes with Arabidopsis development and miRNA function. Dev Cell 2003; 4:205–217.
Tang G, Reinhart BJ, Bartel DP, Zamore, PD . A biochemical framework for RNA silencing in plants. Genes Dev 2003; 17:49–63.
Rhoades MW, Reinhart BJ, Lim LP, et al. Prediction of plant microRNA targets. Cell 2002; 110:513–520.
Jacobsen SE, Running MP, Meyerowitz EM . Disruption of an RNA helicase/RNAse III gene in Arabidopsis causes unregulated cell division in floral meristems. Development 1999; 126:5231–5243.
Schauer SE, Jacobsen SE, Meinke, DW, Ray A . DICER-LIKE1: blindmen and elephants in Arabidopsis development. Trends Plant Sci 2002; 7:487–491.
Boutet S, Vazquez F, Liu J, et al. Arabidopsis HEN1: A genetic link between endogenous miRNA controlling development and siRNA controlling transgene silencing and virus resistance. Curr Biol 2003; 13:843–848.
Vaucheret H, Vazquez F, Crété P, Bartel DP . The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Genes Dev 2004; 18:1187–1197.
Liu B, Li PC, Li X, et al. Loss of function of OsDCL1 affects microRNA accumulation and causes developmental defects in rice. Plant Physiol 2005; 139:296–305.
Millar AA, Gubler F . The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant cell 2005; 17:705–721.
Palatnik JF, Allen E, Wu X, et al. Control of leaf morphogenesis by microRNAs. Nature 2003; 425:257–263.
Emery JF, Floyd SK, Alvarez J, et al. Radial patterning of Arabidopsis shoots by class III HD-ZIP and KANADI genes. Curr Biol 2003; 13:1768–1774.
Kinder CA, Martienssen RA . Spatially restricted microRNA directs leaf polarity through ARGONAUTE1. Nature 2004; 428:81–84.
Aukerman MJ, Sakai H . Regulation of flowering time and floral organ identity by a microRNA and its APETALA2-Like target genes. Plant Cell 2003; 15:2730–2741.
Chen, X . A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004; 303:2022–2025.
Mallory AC, Bartel DP, Bartel B . microRNA-directed regulation of Arabidopsis AUXIN RESPONSE FACTOR17 is essential for proper development and modulates expression of early auxin response genes. Plant cell 2005; 17:1360–1375.
Xie Z, Kasschau KD, Carrington JC . Nagative feedback regulation of Dicer-Like1 in Arabidopsis by microRNA-guided mRNA. Curr Biol 2003; 13:784–789.
Wang JF, Zhou H, Chen YQ, Luo QJ, Qu LH . Identification of 20 microRNAs from Oryza sativa. Nucleic Acids Res 2004; 32:1688–1695.
Li Y, Li W, Jin YX . Computational identification of novel family members of microRNA genes in Arabidopsis thaliana and Oryza sativa. Acta Bioch Bioph Sin 2005; 37:75–87.
Sunkar R, Girke T, Jain PK, Zhu JK . Cloning and characterization of microRNAs from Rice. Plant cell 2005; 17:1397–1411.
Xie Z, Allen E, Fahlgren N, et al. Expression of Arabidopsis MIRNA genes. Plant Physiol 2005; 138:2145–2154.
Xiong Y, Liu T, Tian C, et al. Transcription factors in rice: a genome-wide comparative analysis between monocots and eudicots. Plant Mol Biol 2005; 59:191–203.
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG . The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 1997; 25:4876–4882.
Nicholas KB, Nicholas HB Jr, Deerfield DWII . Genedoc: analysis and visualization of genetic variation. Embnew News 1997; 4:14.
Schmittgen TD, Jiang JM, Liu Q, Yang LQ . A high-throughput method to monitor the expression of microRNA precursors. Nucleic Acids Res 2004; 32:e43.
Chomczynski P, Sacchi N . Single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction. AnalBiochem 1987; 162:732–735.
Clough SJ, Bent AF . Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 1998; 16:735–743.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2-DDCT method. Methods 2001; 25:402–408.
Berezikov E, Guryev V, van de Belt J, et al. Phylogenetic shadowing and computational identification of human microRNA genes. Cell 2005; 120:21–24.
Millar AA, Waterhouse PM . Plant and animal microRNAs: similarities and differences. Funct Integr Genomics 2005; 5:129–135.
Lee YT, Kim MJ, Han JJ, et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J 2004; 23:4051–4060.
International Rice genome sequencing project. The map-based sequence of rice genome. Nature 2005; 436:793–800.
Toledo-Qrtiz G, Huq E, Quail PH . The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell 2003; 15:1749–1770.
Johnson SM, Lin SY, Slack FJ . The time of appearance of the C. elegans let-7 microRNA is transcriptionally controlled utilizing a temporal regulatory element in its promoter. Dev Biol 2003; 259:364–379.
Guo SH, Xie Q, Fei JF, Chua NH . MicroRNA164 directs NAC1 mRNA cleavage to downregulate auxin signals for lateral root development. Plant cell 2005; 17:1376–1386.
Baker CC, Sieber P, Wellmer F, Meyerowitz EM . The early extra petals1 mutant uncovers a role for microRNA miR164c in regulating petal number in Arabidopsis. Curr Biol 2005; 15:303–315.
Acknowledgements
We acknowledge Prof Hervá Vaucheret for kindly supplying the ago1-27 mutant and Dr Xuemei Chen for helpful suggestions and reviewing this manuscript. This work was supported by the Fund of National Key Basic Research Developments Program of the Ministry of Science and Technology China (2001CB109002), National Natural Science Foundation of China (30370893) and Shanghai Municipal Committee of Science and Technology (03JC14061), the Program for New Century Excellent Talents in University (NCET-04-0403), and the ShuGuang Scholarship (04SG15).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jiang, D., Yin, C., Yu, A. et al. Duplication and expression analysis of multicopy miRNA gene family members in Arabidopsis and rice. Cell Res 16, 507–518 (2006). https://doi.org/10.1038/sj.cr.7310062
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.cr.7310062
Keywords
This article is cited by
-
Molecular characterization of miRNA genes and their expression in Dimocarpus longan Lour
Planta (2021)
-
MIR2111-5 locus and shoot-accumulated mature miR2111 systemically enhance nodulation depending on HAR1 in Lotus japonicus
Nature Communications (2020)
-
Regulation of PD-1/PD-L1 Pathway in Cancer by Noncoding RNAs
Pathology & Oncology Research (2020)
-
Endogenous target mimics, microRNA167, and its targets ARF6 and ARF8 during somatic embryo development in Dimocarpus longan Lour.
Molecular Breeding (2015)
-
Genome-wide analysis of aberrantly expressed circulating miRNAs in patients with coal workers’ pneumoconiosis
Molecular Biology Reports (2013)