Abstract
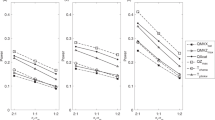
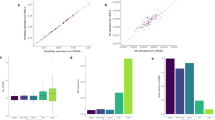
Errors in genotyping can substantially influence the power to detect linkage using affected sib-pairs, but it is not clear what effect such errors have on quantitative trait analyses. Here we use Monte Carlo simulation to examine the influence of genotyping error on multipoint vs two-point analysis, variable map density, locus effect size and allele frequency in quantitative trait linkage and association studies of sib-pairs. The analyses are conducted using variance components methods. We contrast the effects of error on quantitative trait analyses with those on the affected sib-pair design. The results indicate that genotyping error influences linkage studies of affected sib pairs more severely than studies of quantitative traits in unselected sibs. In situations of modest effect size, 5% genotyping error eliminates all supporting evidence for linkage to a true susceptibility locus in affected pairs, but may only result in a loss of 15% of linkage information in random pairs. Multipoint analysis does not suffer substantially more than two-point analysis; for moderate error rates (<5%), multipoint analysis with error is more powerful than two-point with no error. Map density does not appear to be an important factor for linkage analysis. QTL association analyses of common alleles are reasonably robust to genotyping error but power can be affected dramatically with rare alleles.
Similar content being viewed by others
Article PDF
References
1 Buetow KH: . Influence of aberrant observations on high-resolution linkage analysis outcomes. Am J Hum Genet 1991 49: 985–994.
Ott J: . Analysis of Human Genetic Linkage (Johns Hopkins University Press, Baltimore 1991
Lincoln SE, Lander ES: . Systematic detection of errors in genetic linkage data. Genomics 1992 14: 604–610.
Shields DC, Collins A, Buetow KH, Morton NE: . Error filtration, interference, and the human linkage map. Proc Natl Acad Sci USA 1991 88: 6501–6505.
Ott J: . Strategies for characterizing highly polymorphic markers in human gene mapping. Am J Hum Genet 1992 51: 283–290.
Douglas JA, Boehnke M, Lange K: . A multipoint method for detecting genotyping errors and mutations in sibling-pair linkage data. Am J Hum Genet 2000 66: 1287–1297.
Risch N, Merikangas K: . The future of genetic studies of complex human diseases. Science 1996 273: 1516–1517.
Risch NJ: . Searching for genetic determinants in the new millennium. Nature 2000 405: 847–856.
Brzustowicz LM, Merette C, Xie X et al. Molecular and statistical approaches to the detection and correction of errors in genotype databases. Am J Hum Genet 1993 53: 1137–1145.
Goring HH, Terwilliger JD: . Linkage analysis in the presence of errors II: marker-locus genotyping errors modeled with hypercomplex recombination fractions. Am J Hum Genet 2000 66: 1107–1118.
Roberts L: . Human genome research. SNP mappers confront reality and find it daunting. Science 2000 287: 1898–1899.
Abecasis GR, Cookson WO, Cardon LR: . Pedigree tests of transmission disequilibrium. Eur J Hum Genet 2000 8: 545–551.
Abecasis GR, Cardon LR, Cookson WO: . A general test of association for quantitative traits in nuclear families. Am J Hum Genet 2000 66: 279–292.
Risch N: . Linkage strategies for genetically complex traits. I. Multilocus models. Am J Hum Genet 1990 46: 222–228.
Kong A, Cox NJ: . Allele-sharing models: LOD scores and accurate linkage tests. Am J Hum Genet 1997 61: 1179–1188.
Lander ES, Green P: . Construction of multilocus genetic linkage maps in humans. Proc Natl Acad Sci, USA 1984 84: 2363–2367.
Fulker DW, Cherny SS, Sham PC, Hewitt JK: . Combined linkage and association sib-pair analysis for quantitative traits. Am J Hum Genet 1999 64: 259–267.
Risch N, Zhang H: . Extreme discordant sib pairs for mapping quantitative trait loci in humans. Science 1995 268: 1584–1589.
Acknowledgements
The authors are supported by the Wellcome Trust; GRA is a Wellcome Prize student. This work was supported in part by grant EY-12562 from the National Eye Institute, National Institutes of Health, USA. We wish to thank two anonymous reviewers for their helpful comments.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Abecasis, G., Cherny, S. & Cardon, L. The impact of genotyping error on family-based analysis of quantitative traits. Eur J Hum Genet 9, 130–134 (2001). https://doi.org/10.1038/sj.ejhg.5200594
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200594
Keywords
This article is cited by
-
Marker genotyping error effects on genomic predictions under different genetic architectures
Molecular Genetics and Genomics (2021)
-
Mapping quantitative resistance loci for bacterial leaf streak disease in hard red spring wheat using an identity by descent mapping approach
Euphytica (2015)
-
Simultaneous estimation of QTL effects and positions when using genotype data with errors
Journal of Genetics (2015)
-
Using Mendelian inheritance errors as quality control criteria in whole genome sequencing data set
BMC Proceedings (2014)
-
Influence of genotyping error in linkage mapping for complex traits – an analytic study
BMC Genetics (2008)