Abstract
One major limitation of pre-implantation genetic diagnosis (PGD) practice comes from the need to develop single cell PCR protocols. For a disease such as cystic fibrosis (CF), for which almost 1000 mutations have been identified, the development of a mutation based PGD protocol is impracticable. An elegant way to overcome this problem is to set up an indirect diagnosis using polymorphic markers allowing the identification of the pathogenic haplotype instead of the mutation. We present here a new PGD protocol for CF. Our strategy is based on a multiplex fluorescent PCR co-amplifying the ΔF508 mutation and two CFTR intragenic polymorphic microsatellites (IVS8CA and IVS17bCA). Such an approach is justified since in 91% of the cases at least one partner of the couple carries the ΔF508 mutation. The use of intragenic markers reduces the risk of misdiagnosis due to meiotic recombination. In 97% of the single lymphoblasts (151/155) tested a PCR signal was obtained. A complete haplotyping was achieved in 137/151 (91%) lymphoblasts and a 6% rate of allele drop out (ADO) was observed. Three cases were performed. Case one was at risk of transmitting mutations ΔF508 and R1162X, case 2 ΔF508 and R1066C and case 3 ΔF508 and 1341+1A. Considering these three cases and the re-analysis of the affected embryos, we have analysed 62 blastomeres from which we had PCR signal for 58 (94%) and a complete haplotype for 49 (84%). With the degree of polymorphism of the markers used in this work (48 and 39%) and the fact that we co-amplified the F508 locus our test should be suitable for nearly 80% of the couples requesting PGD for CF. This fluorescent multiplex PCR indirect diagnosis provides also a safer test since it allows the confirmation of the diagnosis, the detection of contamination and could give an indication on the ploidy of the embryos tested.
Similar content being viewed by others
Introduction
Pre-implantation genetic diagnosis (PGD) represents an alternative to prenatal diagnosis (PDN) avoiding the burden of a termination of pregnancy (TOP) if the embryo or foetus is diagnosed as affected. PGD requires the combined efforts of geneticists and workers in the field of reproductive medicine. It involves in vitro fertilisation (IVF), embryo biopsy to obtain one or two cells from eight-cell stage embryos and genetic analysis of the isolated cells by polymerase chain reaction (PCR) or fluorescent in situ hybridisation (FISH). PGD is available for an expanding range of genetic defects such as point mutations, deletions or insertions involved in monogenic diseases (recessive or dominant) or chromosomal abnormalities.1,2 The major difficulty of the PGD procedure relies on the genetic analysis of a single cell. When analysed by PCR this involves the successful amplification of a specific DNA copy of each chromosome. Although the efficiency and the accuracy of the diagnosis procedure has increased with the introduction of fluorescent PCR3,4 and multiplex PCR,5,6 a low risk of misdiagnosis persists.7 Another difficulty is the development of multiplex PCR from a single cell. The range of mutations responsible for a particular disease is another difficulty in the development of new PGD. One way to considerably improve the accuracy and efficiency, while limiting the number of assays to diagnose one specific disorder, would be to base the diagnosis on the characterisation of the pathogenic haplotype (using polymorphic markers) rather than on the identification of the mutation.
Amongst monogenic disorders, cystic fibrosis (CF) represents the most common request for PGD.2 CF is the most common severe autosomal recessive disorder in the Caucasian population with a carrier frequency of one in 25, affecting one in 2500 newborn children. The major symptoms are obstruction and chronic infection of the respiratory tract and pancreatic insufficiency leading, in most of the cases, to an early death. A minor form of CF results in a congenital bilateral absence of the vas deferens (CBAVD).8,9 Since CBAVD does not affect spermatogenesis, an IVF with intracytoplasmic sperm injection (ICSI) with testicular sperm can be offered to these patients.10,11,12 If the female partner is also carrier of a mutation in the cystic fibrosis transmembrane conductance regulator (CFTR) gene, the ICSI procedure can be followed by a PGD. There have been almost 1000 mutations characterised in the CFTR gene (http://www.genet.sickkids.on.ca/cftr/). The mutation ΔF508, (deletion of the phenylalanine at codon 508) accounts for almost 70% of the mutations in the Caucasian population, all the other mutations have a frequency ranging from a few per cent to private mutations. Therefore different situations can be encountered: (i) couples where both partners carry the ΔF508 mutation (49%). In such cases, a simple ΔF508 detection test can be used;13,14,15 (ii) couples with one partner carrying the ΔF508 mutation and the other partner carrying another mutation (42%). In this case, a ΔF508 detection test can also be used to eliminate all embryos with the ΔF508 mutation. Such a strategy, even though proposed, is not satisfactory because it results in the rejection of healthy ΔF508 carrier embryos; (iii) couples where both partners carry a mutation different from ΔF508 (9%). In this last situation, either a specific test is developed for each new mutation, which has been done for the most common mutations16,17,18 or an indirect diagnosis is set up.17,19,20,21 Allele identification combines the following advantages: (i) availability of a single test applicable for a large number of couples as soon as they are informative for the polymorphic markers used; (ii) availability of an internal amplification and contamination control; (iii) the use of two or more polymorphic markers provides additional information, increasing the accuracy of the diagnosis; (iv) to provide an evaluation of the ploidy status of the blastomere tested.19
We present here a novel fluorescent multiplex PCR strategy based on the coamplification of the ΔF508 mutation and of two polymorphic intragenic microsatellites, IVS8CA and IVS17bCA. This strategy, albeit similar, differs from the one described by Dreesen et al;19 who used external markers only and did not simultaneously detect the ΔF508 mutation. Such an approach is warranted since in most cases (91%) at least one member of the couple carries the ΔF508 mutation. This strategy allows PGD for informative couples at risk of transmitting CF to their children whatever CF mutation is implied. Furthermore, we report three CF PGD cycles performed in our centre.
Material and methods
Single lymphoblast and blastomere testing
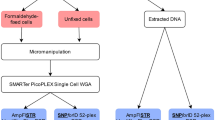
Three Epstein-Barr transformed lymphoblast cell lines were used to perform single cell analysis: one homozygous for the ΔF508 mutation (ΔF/ΔF; line GM04540A; Coriell Cell Repositories, Camden, USA), one heterozygous for this mutation (ΔF/F; line CF carrier, kindly provided by Dr K Sermon, Dutch-speaking Brussels free University, Belgium) and one homozygous normal for the F508 locus (F/F; line SMNt, kindly provided by Dr Judith Melki, Genopole Evry, France). For the three cell lines the marker profiles were first determined on genomic DNA. Tubing procedure was performed as previously described.15 Supernumerary IVF embryos not suitable for freezing and therefore scheduled for destruction were incubated for 10 min in an embryo biopsy medium, Ca and Mg free (EBM, Medicult-France, Lyon, France) prior to biopsy. They were then biopsied as previously described22 under an inverted microscope (Leica, Wetzlar, Germany) using Research Instrument micromanipulators and micropipettes (Research Instruments Ltd, Penryn, UK). Zona pellucida drilling was performed using a laser set up (Hamilton-Thorn, Beverly, USA) and blastomeres were gently aspirated and placed in G2.1 medium (Scandinavian Science, Gothenburg, Sweden). Each blastomere was washed three times in EBM, transferred into the reaction tube containing 2.5 μl of Lysis Buffer (LB: 200 mM KOH, 50 mM DTT) and frozen at −20°C until amplification by PCR.
PCR reactions
IVS8CA and IVS17bCA are CFTR intragenic markers with heterozygous frequencies of 48% and 39% respectively.23,24 Primers for IVS8CA and IVS17bCA amplification were designed to have a similar annealing temperature so that the DNA fragments could be co-amplified with the F508 locus in a single-round of fluorescent multiplex PCR (Table 1). For each primer pair, one was fluorescently labelled with 6-fam (F508) or tet (IVS8CA) or hex (IVS17bCA) fluorochrome (Perkin Elmer Applied Biosystems, Warrington, UK). Aliquots of 22.5 μl PCR master mix containing neutralising buffer (NB: 900 mM TrisHCl, 300 mM KCl and 200 mM HCl), 1.5 mM MgCl2, 200 μM each dNTP, 0.2 μM each primer for F508 locus, 0.4 μM each primer for IVS8CA and IVS17bCA loci and 1.5 U HotStar Taq DNA Polymerase (Qiagen, Courtaboeuf, France) were added to the reaction tubes. PCR reactions were performed using a Mastercycler Gradient PCR machine (Eppendorf, Hamburg, Germany). The program used was 15 min of initial denaturation at 95°C followed by 30 s of denaturation at 96°C for the first 10 cycles and 94°C for the remaining 35 cycles. Annealing and elongation cycles were performed for 30 s at 61°C and 30 s at 72°C, respectively. Reactions were terminated by 10 min at 72°C. PCR products were analysed on either an ABI310 or an ABI377 automated sequencer (Applied Biosystems, Warrington, UK). PCR efficiency was calculated from samples for which a PCR signal for at least one tested locus was obtained.
Informativity testing for couples requesting PGD
Genomic DNA from each couple requesting PGD and from their affected foetus/child was extracted from whole blood samples or from an amniocentesis sample using the standard saline protocol.25 Haplotype analysis for IVS8CA and IVS17bCA microsatellites and segregation analysis with regard to the locus F508 of the CFTR gene were performed using the triplex PCR procedure described above, except that NB was replaced by buffer II (BII: 500 mM KCL, 100 mM Tris HCl pH 8.3) and 30 cycles were sufficient for amplification.
Clinical PGDs
Three couples requested PGD for CF.
Case 1: a 30-year-old woman and her 26-year-old husband had previously had a TOP after prenatal diagnosis of a foetus diagnosed because of an hyperechogenic bowel detected during ultrasonography. This foetus was a compound heterozygote for mutations ΔF508 (paternally inherited) and R1162X (maternally inherited).
Case 2: a 33-year-old woman and her 34-year-old husband had an affected girl with a severe form of CF. She is a compound heterozygote for mutations ΔF508 (maternally inherited) and R1066C (paternally inherited).
Case 3: a 29-year-old woman and her 33-year-old husband with an affected girl with a severe form of CF had experienced a TOP. The daughter and the affected foetus were compound heterozygotes for mutations ΔF508 (maternally inherited) and 1341+1A (paternally inherited).
For the three couples, ovarian stimulation, oocyte recovery and ICSI procedures were carried out using standard protocols.26,27 Three days after ICSI, embryos were biopsied and blastomeres were collected–one blastomere from embryos with less than six cells and two from embryos with six or more cells–and analysed. For each biopsied cell, a blank control was prepared from the final EBM wash drop. PCR controls containing single lymphoblast from control cell lines (ΔF/F and ΔF/ΔF) were added.
From untransferred affected embryos at least two blastomeres were individually reanalysed with the exception of embryo 10 from case 2 which was reanalysed as a whole.
Results
Lymphoblast testing
Table 2 shows the results of triplex PCRs performed on single lymphoblasts from three different cell lines. All three lines were informative for both markers tested. We performed 155 single cell analysis for which an amplification of at least one marker was obtained in 151 (97%) single lymphoblasts. Genotypes were reliably determined in 144 out of 151 (95%), in 143 out of 151 (95%) and in 145 out of 151 (96%) of cells for loci F508, IVS8CA and IVS17bCA, respectively. Complete haplotyping was achieved in 137 out of 151 (91%) lymphoblasts. A global allele drop out (ADO) rate of 6% was observed whatever locus and whatever cell line was considered. Eight blanks out of 58 (14%) presented a PCR signal for at least one of the loci tested. Such a contamination rate is quite high, but it is worth noticing that in one experiment five blanks out of 22 presented a PCR signal, accounting for most of the contaminations detected in these experiments. For all the other experiments the contamination rate was acceptable.
Blastomere testing
Thirteen blastomeres from surplus untransferable and unfreezable embryos were tested. Positive amplification was observed in 12 out of 13 (92%) samples. Ten (83%) of them gave a positive PCR signal for all three loci. The accuracy of the diagnosis was impossible to evaluate since we did not know the parental haplotype, which also prevented the evaluation of the ADO rate. Five out of 13 blank controls showed amplification (38%). Such a high rate of contamination is probably due to the poor sample quality since some blastomeres were lysed during biopsy and DNA might be present in the wash drop as previously noticed.4,28 It is worth noticing that our contamination rate during the last ten PGD performed in our centre was 6.7% over 89 blanks tested.
We thus concluded that our triplex PCR is efficient and reliable and felt confident for clinical application.
Informativity testing for couples requesting PGD
The results of haplotype analysis are shown in Figure 1. Genotyping was performed in any case on both parents and for case 1 on an amniocentesis sample from an affected foetus (only marker IVS8CA was informative); for case 2 on their affected daughter (only marker IVS8CA was informative); for case 3 on their affected daughter (both markers could be used for the PGD). In all cases a triplex PCR was performed since the non-informative marker can still be used to detect DNA contamination.
Clinical PGDs
The results of PGD cycles are summarized in Table 3. An electrophoregram is shown in Figure 2.
Case 1: Nine cumulus-oocyte complexes were retrieved from which seven were inseminated by ICSI and four 2PN stage embryos were obtained. On day 3, four embryos had reached the eight cells stage and two blastomeres per embryo were biopsied. Three embryos were diagnosed as affected and one (embryo 2) presented a discordant result between the two blastomeres analysed, no embryo was transferred. Reanalysis of at least two blastomeres from the untransferred embryos confirmed our previous diagnoses even for embryo 2, for which a discordant result was again obtained. (Table 3).
Case 2: 20 cumulus-oocyte complexes were retrieved from which 18 were inseminated by ICSI, 14 of which exhibited two pronuclei after ICSI. Three days after fertilisation, 11 embryos were suitable for biopsy and one or two blastomeres were successfully removed and analysed. Nine embryos gave a clear result: one was unaffected, three were carriers and five were affected. A complete amplification failure was obtained for embryo 17. This embryo was not transferred and the re-analysis of two blastomeres diagnosed it as affected. For embryo 18, we obtained an ambiguous result because of a discordant result between the two blastomeres analysed.
Two embryos were transferred but no pregnancy ensued. Reanalysis of the affected embryos confirmed the initial results, for embryo 18, the genotype remained ambiguous (Table 3).
Case 3: six cumulus-oocyte complexes were retrieved and inseminated by ICSI, all of them exhibited two pronuclei. Three days later two embryos were suitable for biopsy and two blastomeres were successfully removed and analysed. A full diagnosis was obtained on both cells for both embryos resulting in the transfer of one unaffected embryo free of any mutation, no pregnancy ensued. The re-analysis of three blastomeres of the affected embryo confirmed our previous diagnosis (Table 3).
In total, we have analysed 31 blastomeres for which we obtained a PCR signal for 28 (90%). From these 26 (93%), 25 (89%) and 25 (89%) were correctly diagnosed at the loci F508, IVS8CA and IVS17bCA respectively. A full genotype was possible in 22 cases (79%). A complete amplification failure from three blastomeres (10%), a partial amplification failure from two blastomeres (7%) and two ambiguous results (7%) were observed. If we combine these results with the data obtained from the re-analysis, we had 58 out of 62 (94%) blastomeres that presented a PCR signal from which 55 out of 58 (95%), 54 out of 58 (93%) and 55 out of 58 (95%) showed a signal at the F508, IVS8CA and IVS17bCA loci, respectively. A complete haplotype was obtained in 49 out of 58 (84%). A complete amplification failure was observed four times out of 62 (6%). The ADO rate is difficult to estimate because of the two ambiguous results obtained which can be attributed either to ADO or to a chaotic constitution of the embryos (as often observed in IVF embryos29) and because of the absence of informativity of the markers.
Discussion
Since the development of new PGD tests is highly time consuming (3 to 6 months for the development of a reliable protocol for a new mutation) and considering that most genetic disorders are due to multiple defects in one gene, it is not reasonable to develop a PGD for each mutation. This is especially true for CF since almost 1000 different mutations have been described. A general solution to this problem is to base the diagnosis on an indirect approach by the characterisation of the morbide allele(s), using linked (intra or extragenic) polymorphic markers, instead of detecting the mutation(s) itself. The advantages of such an approach are: (i) having a single diagnosis procedure for most of the couples requesting PGD whatever mutations are involved, as soon as the couple is informative for the polymorphic markers used; (ii) considerably improving the reliability of the tests since it can give multiple information from a single cell. In the most favourable situation each locus tested can be considered as an independent diagnosis; (iii) possibly detecting amplification problems (ADO or amplification failure) or exogenous DNA contamination; (iv) evaluating the ploidy status of the blastomere tested.
As shown by Lewis et al, the misdiagnosis rate is considerably reduced when using a linked maker in addition to the detection of the mutation. This is true for recessive as well as for dominant diseases. The error decreased from 5.8% to 0.44% and from 10.9% to 0.1% respectively when the test is based on the detection of the mutation and a linked marker instead of just the detection of the mutation.7 The misdiagnosis rate and the reliability of the test will correlate with the number of markers and their informativity. Therefore, the use of linked markers allows PGD for a variety of mutations for the same disorder but also provides safer tests.
The use of polymorphic markers in CF PGD has already been described either in simplex PCR protocols using the markers IVS17bTA or M470V or IVS8CA20 or in a multiplex PCR protocol using four highly polymorphic makers (D7S523; D7S486; D7S480; D7S490) flanking the CFTR gene19 or in different duplex combinations involving either markers IVS8CA and IVS17bTA or ΔF508 and IVS17bTA.21 We describe here a new protocol based on the co-amplification of the ΔF508 and two intragenic polymorphic markers (IVS8CA and IVS17bCA). We decided to coamplify the ΔF508 mutation because, if we consider that its frequency is 70% of the CF mutations, for 91% of the couples requesting PGD for CF at least one of the members of the couple will be a carrier of the ΔF508 mutation. Furthermore the use of intragenic markers will avoid the problem of recombination since no crossing-over has been reported so far in the CFTR gene. The main advantage of the protocol developed by Dreesen et al, resides in the use of highly polymorphic markers.19 The disavantage is the risk of meiotic recombination since the markers used are located between 0.2 and 5 cM from the CFTR gene. Hence it is necessary to perform an analysis on at least four members, both parents, one affected child and one unaffected sibling and to have at least one informative marker on each side of the CFTR gene for the PGD application.19 Considering the degree of polymorphism of the markers and the necessity for having at least two informative markers they can, theoretically, propose their protocol to 87% of the couples.19 The degree of polymorphism of the markers used in this work (48 and 39%) represents the major limitation of the proposed approach. The absence of intragenic meiotic recombination in the CFTR gene considerably simplifies the family analysis compared to the strategy of Dreesen et al. However, if we consider that 91% of the couples present at least the ΔF508 mutation, and if we take into account the fact that the ΔF508 locus is co-amplified with the markers IVS8CA and IVS17bCA, our test should be suitable for nearly 80% of the couples requesting PGD for CF. For the couples for which these two markers will not be informative we still have the possibility of offering them a test based on the detection of the ΔF508 mutation only and the elimination of all the embryos carrying this mutation. This represents 42% of the 20% non-informative couples for which only 50% of the embryos will be available for transfer, meaning that a third of the unaffected embryos will be discarded.
Our first experience using this test shows its merits for the PGD of CF since a complete haplotype was obtained in 89% of the blastomeres analysed. Our strategy of coamplifying F508, IVS8CA and IVS17bCA is now used for all our informative couples whatever the mutation is, since it considerably increases the accuracy of the test even if both members of the couple are carriers of the ΔF508 mutation.
References
Geraedts J, Handyside A, Harper J et al. ESHRE Preimplantation Genetic Diagnosis (PGD) Consortium: preliminary assessment of data from January 1997 to September 1998. ESHRE PGD Consortium Steering Committee Hum Reprod 1999 14: 3138–3148
Geraedts J, Handyside A, Harper J et al. ESHRE preimplantation genetic diagnosis (PGD) consortium: data collection II (May 2000) Hum Reprod 2000 15: 2673–2683
Findlay I, Quirke P, Hall J, Rutherford A . Fluorescent PCR: a new technique for PGD of sex and single-gene defects J Assist Reprod Genet 1996 13: 96–103
Sermon K, DeVos A, VandeVelde H et al. Fluorescent PCR and automated fragment analysis for the clinical application of preimplantation genetic diagnosis of myotonic dystrophy (Steinert's disease) Mol Hum Reprod 1998 4: 791–796
Findlay I, Matthews P, Quirke P . Multiple genetic diagnosis from single cells using multiplex PCR: reliability and allele dopout Prenat Diagn 1998 18: 1413–1421
Findlay I, Urquhart A, Quirke P, Sullivan K, Rutherford AJ, Lilford RJ . Simultaneous DNA ‘fingerprinting’, diagnosis of sex and single-gene defect status from single cells Hum Reprod 1995 10: 1005–1013
Lewis CM, Pinel T, Whittaker JC, Handyside AH . Controlling misdiagnosis errors in preimplantation genetic diagnosis: a comprehensive model encompassing extrinsic and intrinsic sources of error Hum Reprod 2001 16: 43–50
Chillon M, Casals T, Mercier B et al. Mutations in the cystic fibrosis gene in patients with congenital absence of the vas deferens N Engl J Med 1995 332: 1475–1480
Claustres M, Guittard C, Bozon D et al. Spectrum of CFTR mutations in cystic fibrosis and in congenital absence of the vas deferens in France Hum Mutat 2000 16: 143–156
Josserand RN, Bey-Omar F, Rollet J et al. Cystic fibrosis phenotype evaluation and paternity outcome in 50 males with congenital bilateral absence of vas deferens Hum Reprod 2001 16: 2093–2097
Lissens W, Liebaers I . The genetics of male infertility in relation to cystic fibrosis Baillieres Clin Obstet Gynaecol 1997 11: 797–817
Viville S, Warter S, Meyer JM et al. Histological and genetic analysis and risk assessment for chromosomal aberration after ICSI for patients presenting with CBAVD Hum Reprod 2000 15: 1613–1618
Ray PF, Ao A, Taylor DM, Winston RM, Handyside AH . Assessment of the reliability of single blastomere analysis for preimplantation diagnosis of the delta F508 deletion causing cystic fibrosis in clinical practice Prenat Diagn 1998 18: 1402–1412
Findlay I, Ray P, Quirke P, Rutherford A, Lilford R . Allelic drop-out and preferential amplification in single cells and human blastomeres: implications for preimplantation diagnosis of sex and cystic fibrosis Hum Reprod 1995 10: 1609–1618
Moutou C, Viville S . Improvement of preimplantation genetic diagnosis (PGD) for the cystic fibrosis mutation delta F508 by fluorescent polymerase chain reaction Prenat Diagn 1999 19: 1248–1250
Avner R, Laufer N, Safran A, Kerem BS, Friedmann A, Mitrani-Rosenbaum S . Preimplantation diagnosis of cystic fibrosis by simultaneous detection of the W1282X and delta F508 mutations Hum Reprod 1994 9: 1676–1680
Goossens V, Sermon K, Lissens W et al. Clinical application of preimplantation genetic diagnosis for cystic fibrosis Prenat Diagn 2000 20: 571–581
Scobie G, Woodroffe B, Fishel S, Kalsheker N . Identification of the five most common cystic fibrosis mutations in single cells using a rapid and specific differential amplification system Mol Hum Reprod 1996 2: 203–207
Dreesen JC, Jacobs LJ, Bras M et al. Multiplex PCR of polymorphic markers flanking the CFTR gene; a general approach for preimplantation genetic diagnosis of cystic fibrosis Mol Hum Reprod 2000 6: 391–396
Eftedal I, Schwartz M, Bendtsen H, Andersen AN, Ziebe S . Single intragenic microsatellite preimplantation genetic diagnosis for cystic fibrosis provides positive allele identification of all CFTR genotypes for informative couples Mol Hum Reprod 2001 7: 307–312
Goossens V, Sermon K, Lissens W et al. Duplex-PCR of polymorphic markers and CF-mutations in the CFTR-gene: application in preimplantation genetic diagnosis 17th Annual Meeting of ESHRE, Lausanne Hum Reprod 2001 16: 53
Hardy K, Handyside AH . Biopsy of cleavage stage human embryos and diagnosis of single gene defects by DNA amplification Arch Pathol Lab Med 1992 116: 388–392 Issn: 0003–9985
Morral N, Estivill X . Multiplex PCR amplification of three microsatellites within the CFTR gene Genomics 1992 13: 1362–1364
Morral N, Girbau E, Zielenski J et al. Dinucleotide (CA/GT) repeat polymorphism in intron 17B of the cystic fibrosis transmembrane conductance regulator (CFTR) gene Hum Genet 1992 88: 356
Rousseau F, Heitz D, Biancalana V, Oberle I, Mandel JL . On some technical aspects of direct DNA diagnosis of the fragile X syndrome Am J Med Genet 1992 43: 197–207
Van Steirteghem AC, Nagy Z, Joris H et al. High fertilization and implantation rates after intracytoplasmic sperm injection Hum Reprod 1993 8: 1061–1066
Wittemer C, Ohl J, Bettahar-Lebugle K, Viville S, Nisand I . A quantitative and morphological analysis of oocytes collected during 438 IVF cycles J Assist Reprod Genet 2000 17: 44–50
Moutou C, Gardes N, Rongieres C et al. Allele-specific amplification for preimplantation genetic diagnosis (PGD) of spinal muscular atrophy Prenat Diagn 2001 21: 498–503
Handyside AH, Delhanty JD . Preimplantation genetic diagnosis: strategies and surprises Trends Genet 1997 13: 270–275
Acknowledgements
We would like to thank Serge Vicaire for his help with the ABI377, the IVF team at the SIHCUS-CMCO and Prof Ingeborg Liebaers, Dr Uwe Strahle, Dr Irwin Davidson and Prof Michel Koenig for their helpful comments on this manuscript. This work was supported by funds from the Association Française contre les Myopathies (AFM), the Association Française de Lutte contre la Mucoviscidose (AFLM-VLM), the Fond pour la recherche Organon (FARO), from the Centre National de la Recherche Scientifique (CNRS), the Institut National de la Santé et de la Recherche Médicale (INSERM) and the Hôpital Universitaire de Strasbourg.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Moutou, C., Gardes, N. & Viville, S. Multiplex PCR combining deltaF508 mutation and intragenic microsatellites of the CFTR gene for pre-implantation genetic diagnosis (PGD) of cystic fibrosis. Eur J Hum Genet 10, 231–238 (2002). https://doi.org/10.1038/sj.ejhg.5200794
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.ejhg.5200794
Keywords
This article is cited by
-
Universal strategy for preimplantation genetic testing for cystic fibrosis based on next generation sequencing
Journal of Assisted Reproduction and Genetics (2020)
-
Improving preimplantation genetic diagnosis for Fragile X syndrome: two new powerful single-round multiplex indirect and direct tests
European Journal of Human Genetics (2016)
-
High-efficiency derivation of human embryonic stem cell lines following pre-implantation genetic diagnosis
In Vitro Cellular & Developmental Biology - Animal (2010)
-
New tools for preimplantation genetic diagnosis of Huntington's disease and their clinical applications
European Journal of Human Genetics (2004)