Abstract
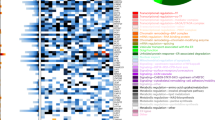
DNA microarrays are powerful tools for the analysis of gene expression on a genomic scale. The importance of individual regulatory events for the process under study can however not be deduced unequivocally without additional experiments. We devised a strategy to identify central regulators of cancer drug responses by combining the results of microarray experiments with efficient methods for phenotypic testing of candidate genes. We exposed murine FL5.12 pro-B cells to cisplatin, camptothecin, methotrexate or paclitaxel, respectively and analysed the patterns of gene expression with cDNA microarrays. Drug-specific regulatory events as well as intersections between different apoptotic pathways, including previously studied responses to staurosporine and interleukin-3 (IL-3) deprivation, were identified. Genes shared by at least three pathways were chosen for further analysis. Ectopic expression of three such genes, TEAP, GP49B, and Lipin1 was found to have an anti-proliferative effect on pro-B cells. Interestingly, we identified hemoglobin alpha as a strong pro-apoptotic regulator. While hemoglobin-expressing cells were growing normally in the presence of IL-3, they displayed accelerated apoptosis with similar kinetics as Bax overexpressing cells upon IL-3 removal. The pro-apoptotic effect of hemoglobin was suppressed by Bcl-2 and was characterized by enhanced stimulation of caspase activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Amundson SA, Bittner M, Chen Y, Trent J, Meltzer P, Fornace Jr AJ . 1999 Oncogene 18: 3666–3672
Arevalo-Rodriguez M, Cardenas ME, Wu X, Hanes SD, Heitman J . 2000 EMBO J. 19: 3739–3749
Beinert T, Munzing S, Possinger K, Krombach F . 2000 J. Leukoc. Biol. 67: 369–373
Blagosklonny MV, Fojo T . 1999 Int. J. Cancer 83: 151–156
Brachat A, Pierrat B, Brungger A, Heim J . 2000 Oncogene 19: 5073–5082
Brouard S, Otterbein LE, Anrather J, Tobiasch E, Bach FH, Choi AM, Soares MP . 2000 J. Exp. Med. 192: 1015–1026
Carrier A, Nguyen C, Victorero G, Granjeaud S, Rocha D, Bernard K, Miazek A, Ferrier P, Malissen M, Naquet P, Malissen B, Jordan BR . 1999 Immunogenetics 50: 255–270
Cohen SM, Lippard SJ . 2001 Prog. Nucleic Acid Res. Mol. Biol. 67: 93–130
Dunn SE, Hardman RA, Kari FW, Barrett JC . 1997 Cancer Res. 57: 2687–2693
Ferrante K, Winograd B, Canetta R . 1999 Cancer Chemother. Pharmacol. 43: (Suppl) 61–68
Gu Z, Flemington C, Chittenden T, Zambetti GP . 2000 Mol. Cell. Biol. 20: 233–241
Katz HR, Vivier E, Castells MC, McCormick MJ, Chambers JM, Austen KF . 1996 Proc. Natl. Acad. Sci. USA 93: 10809–10814
Lee SP, Hwang YS, Kim YJ, Kwon KS, Kim HJ, Kim K, Chae HZ . 2001 J. Biol. Chem. 276: 29826–29832
Lehar SM, Nacht M, Jacks T, Vater CA, Chittenden T, Guild BC . 1996 Oncogene 12: 1181–1187
Leist M, Jäättelä . 2001 Nat. Rev. Mol. Cell Biol. 2: 589–598
Matsuda S, Kawamura-Tsuzuku J, Ohsugi M, Yoshida M, Emi M, Nakamura Y, Onda M, Yoshida Y, Nishiyama A, Yamamoto T . 1996 Oncogene 12: 705–713
Matsumoto Y, Wang LL, Yokoyama WM, Aso T . 2001 J. Immunol. 166: 781–786
Meguro T, Chen B, Lancon J, Zhang JH . 2001 J. Neurochem. 77: 1128–1135
Nicholson DW . 2000 Nature 407: 810–816
Noma T, Fujisawa K, Yamashiro Y, Shinohara M, Nakazawa A, Gondo T, Ishihara T, Yoshinobu K . 2001 Biochem. J. 358: 225–232
Okamoto K, Prives C . 1999 Oncogene 18: 4606–4615
Olsen EA . 1991 J. Am. Acad. Dermatol. 25: 301–318
Peterfy M, Phan J, Xu P, Reue K . 2001 Nat. Genet. 27: 121–124
Rouault JP, Falette N, Guehenneux F, Guillot C, Rimokh R, Wang Q, Berthet C, Moyret-Lalle C, Savatier P, Pain B, Shaw P, Berger R, Samarut J, Magaud JP, Ozturk M, Samarut C, Puisieux A . 1996 Nat. Genet. 14: 482–486
Schena M, Shalon D, Davis RW, Brown PO . 1995 Science 270: 467–470
Schena M, Shalon D, Heller R, Chai A, Brown PO, Davis RW . 1996 Proc. Natl. Acad. Sci. USA 93: 10614–10619
Schneider E, Hsiang YH, Liu LF . 1990 Adv. Pharmacol. 21: 149–183
Swift S, Lorens J, Achacoso P, Nolan GP . 1999 Cur. Proto. Immunol. Unit 10 28: Suppl
Tomasini R, Azizi Samir A, Vaccaro MI, Pebusque MJ, Dagorn JC, Iovanna JL, Dusetti NJ . 2001 J. Biol. Chem. 276: 44185–44192
Voehringer DW, Hirschberg DL, Xiao J, Lu Q, Roederer M, Lock CB, Herzenberg LA, Steinman L, Herzenberg LA . 2000 Proc. Natl. Acad. Sci. USA 97: 2680–2685
Wang LL, Chu DT, Dokun AO, Yokoyama WM . 2000 J. Immunol. 164: 5215–5220
Yang WM, Yao YL, Seto E . 2001 EMBO J. 20: 4814–4825
Yoneda T, Sato M, Maeda M, Takagi H . 1998 Brain Res. Mol. Brain Res. 62: 187–195
Acknowledgements
We thank Marion Kamke and Brigitte Besenreuther for excellent technical assistance.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brachat, A., Pierrat, B., Xynos, A. et al. A microarray-based, integrated approach to identify novel regulators of cancer drug response and apoptosis. Oncogene 21, 8361–8371 (2002). https://doi.org/10.1038/sj.onc.1206016
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1206016
Keywords
This article is cited by
-
Proteomic evidences for microcystin-RR-induced toxicological alterations in mice liver
Scientific Reports (2018)
-
Selection for methotrexate resistance in mammalian cells bearing a Drosophila dihydrofolate reductase transgene
Cell Biology and Toxicology (2010)
-
In praise of arrays
Pediatric Nephrology (2009)
-
A role for Drosophila in understanding drug-induced cytotoxicity and teratogenesis
Cytotechnology (2008)
-
Upregulation of alpha globin promotes apoptotic cell death in the hematopoietic cell line FL5.12
Apoptosis (2005)